Marinka Zitnik
@marinkazitnik
Followers
6,363
Following
226
Media
138
Statuses
2,741
Assistant Professor at Harvard | Faculty @Harvard @KempnerInst AI | Faculty @broadinstitute @harvard_data | Cofounder @ProjectTDC | @AI_for_Science
Harvard
Joined July 2009
Don't wanna be here?
Send us removal request.
Explore trending content on Musk Viewer
#ขวัญฤทัยตอนจบ
• 574087 Tweets
オーロラ
• 477961 Tweets
Joost
• 327556 Tweets
jeonghan
• 127459 Tweets
Fulham
• 79631 Tweets
#SixTONESANN
• 76439 Tweets
Man City
• 74328 Tweets
#الهلال_الحزم
• 61542 Tweets
自分これ
• 61279 Tweets
HeavenlyVoice WithMrC
• 54936 Tweets
Gvardiol
• 51763 Tweets
Ali Koç
• 44051 Tweets
カクレンジャー
• 38465 Tweets
太陽フレアのせい
• 38155 Tweets
キンスパ
• 37413 Tweets
Dremo
• 36451 Tweets
マリノス
• 29498 Tweets
Burnley
• 27714 Tweets
SUPER READY FOR SUPERNOVA
• 25151 Tweets
SEE YOU NEXT MONTH JIN
• 24910 Tweets
Luton
• 19603 Tweets
Haley
• 18797 Tweets
Dursun Özbek
• 17255 Tweets
#AdayOlAzizYıldırım
• 16137 Tweets
やまとなでしこ
• 13543 Tweets
Last Seen Profiles
Excited to teach a course on
#biomedical
#AI
this semester
@Harvard
@harvardmed
Follow our course materials, reading lists and paper highlights which we will release throughout the semester
Thankful for star TA team
@YEktefaie
@richardjchen
Y Huang
13
153
797
Excited to share our
@Nature
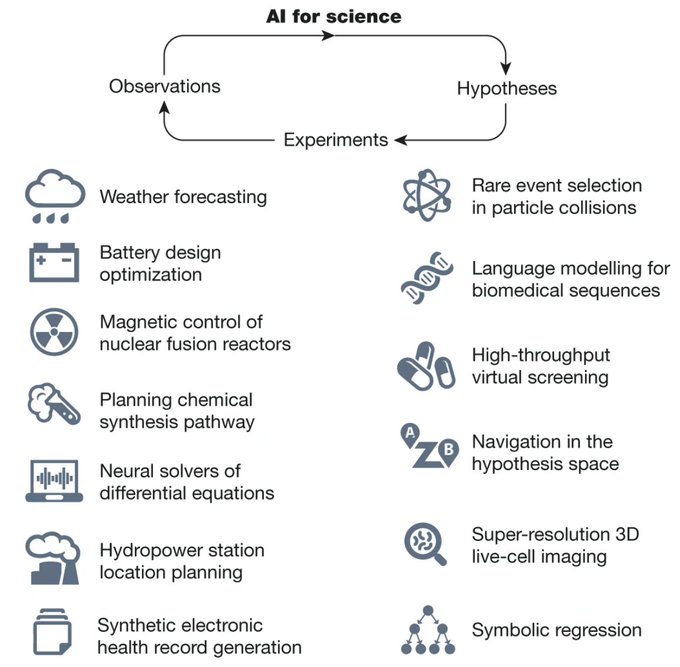
paper on the role of AI in scientific discovery 🌟🔬
#AI4Science
AI is transforming discovery across sciences 🤖🔍 From hypothesis generation to data interpretation, it is reshaping all stages of research in ways we could not imagine using…
11
200
659
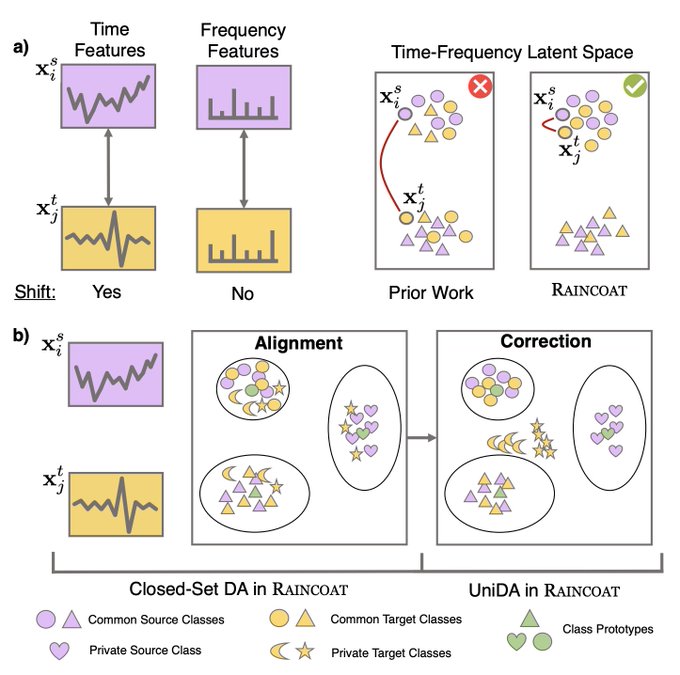
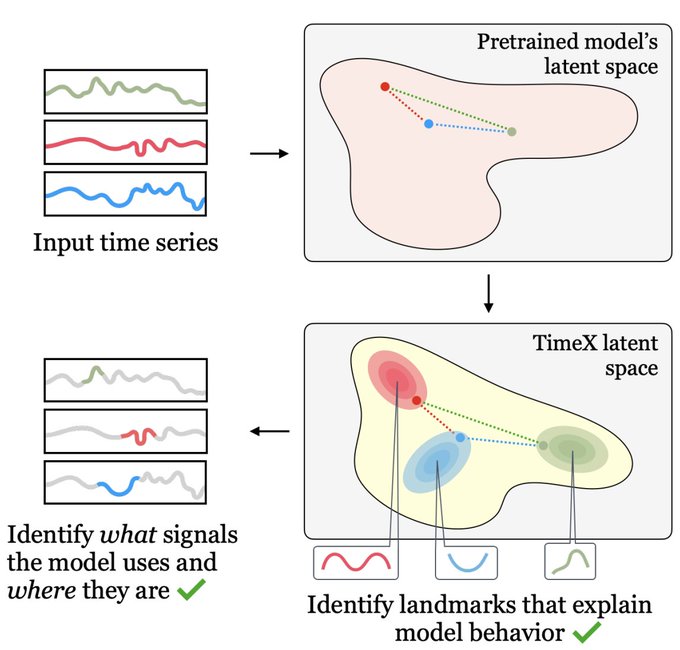
Can we infuse
#structure
into a time series (
#TS
) model from a diverse dataset so as to greatly improve
#generalization
on new TS coming from different datasets?
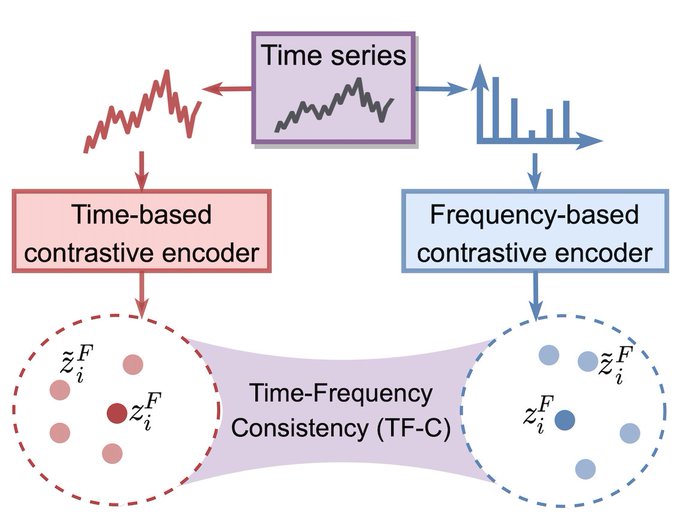
Yes, via a new principle called
#Representational
Time-Frequency Consistency (TF-C)
1/3
3
103
518
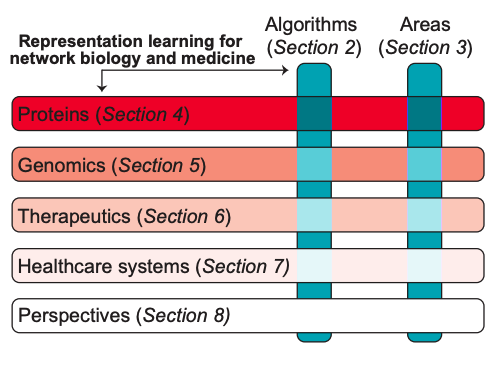
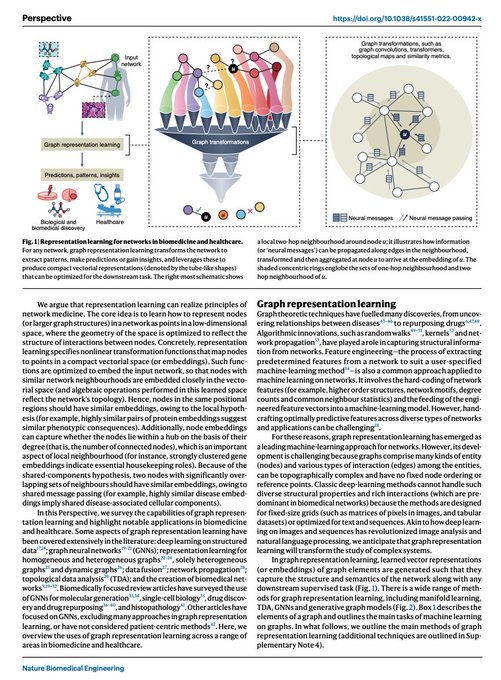
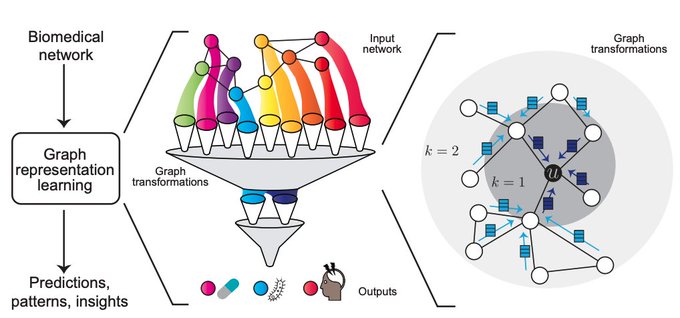
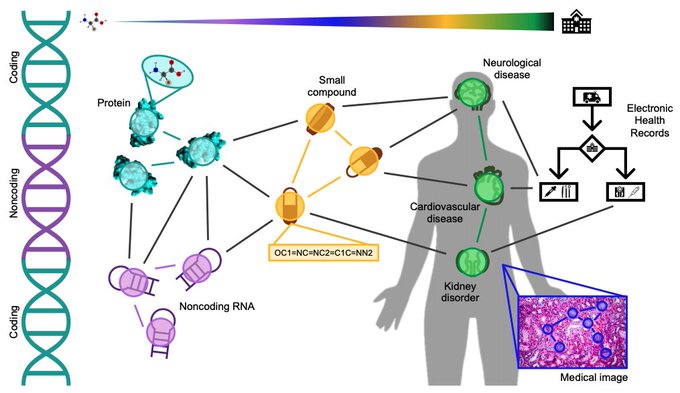
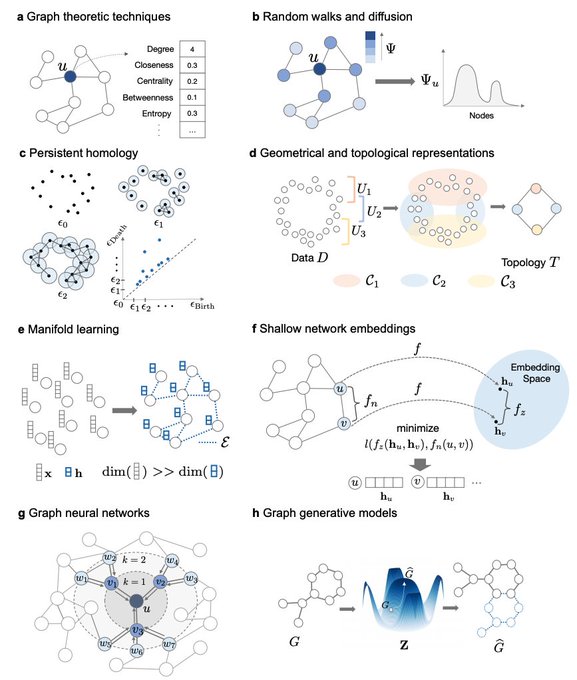
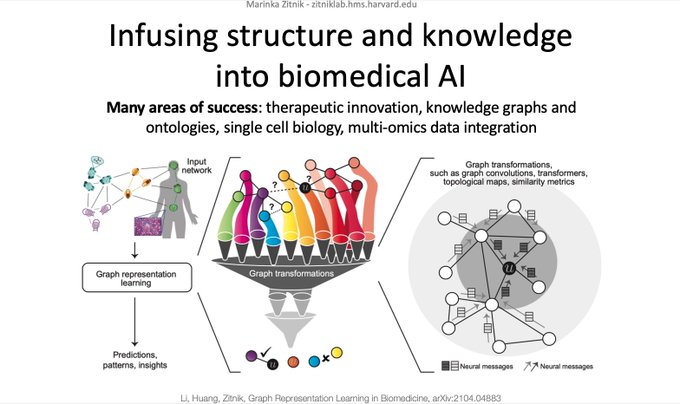
Survey on Representation Learning for Networks in Biology and Medicine
Long-standing principles of biomed nets (often unspoken in ML) provide grounding for representation learning, explain successes & limitations
@_michellemli
@KexinHuang5
#netbio
#GNN
#ML
3
110
449
With
@payal_chandak
and
@KexinHuang5
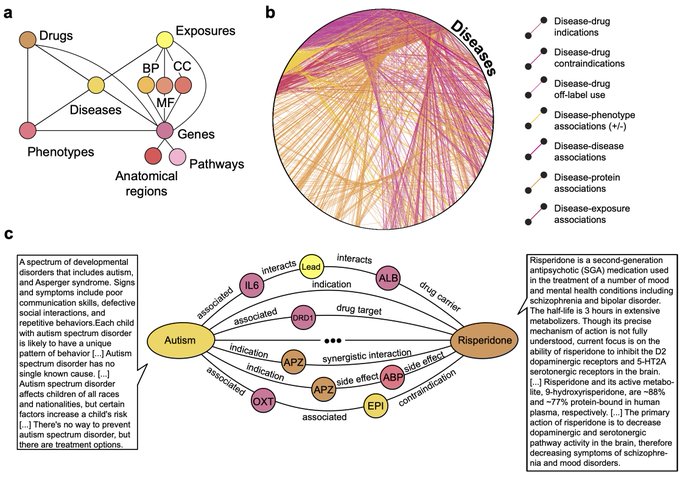
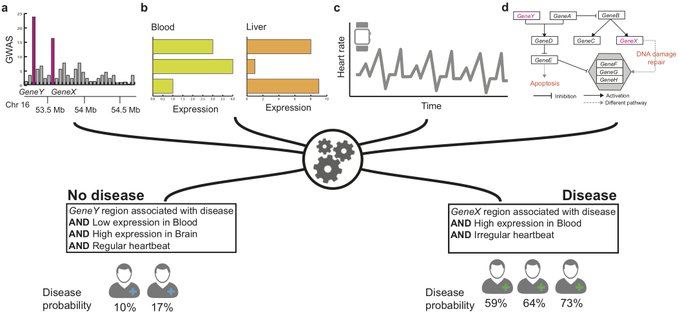
, we are excited to share PrimeKG, a precision medicine-oriented knowledge graph providing holistic and multimodal view of human disease (1/8)
4
49
227
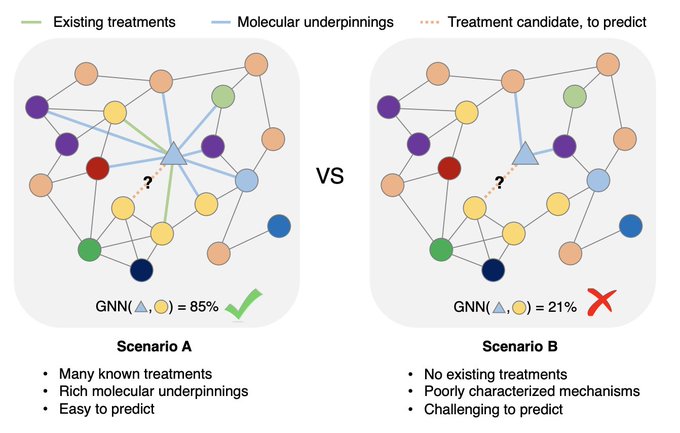
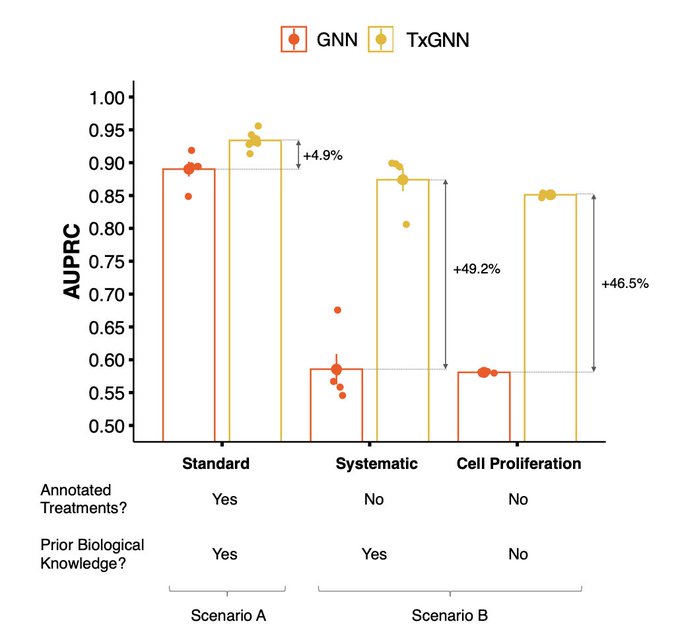
Meet TxGNN, a model that utilizes geometric deep learning and human-centered AI to make zero-shot predictions of therapeutic use across a vast range of 17,080 diseases
@HarvardDBMI
@IcahnMountSinai
@Stanford
@harvard_data
@harvardmed
@MIT_CSAIL
1/9
2
42
222
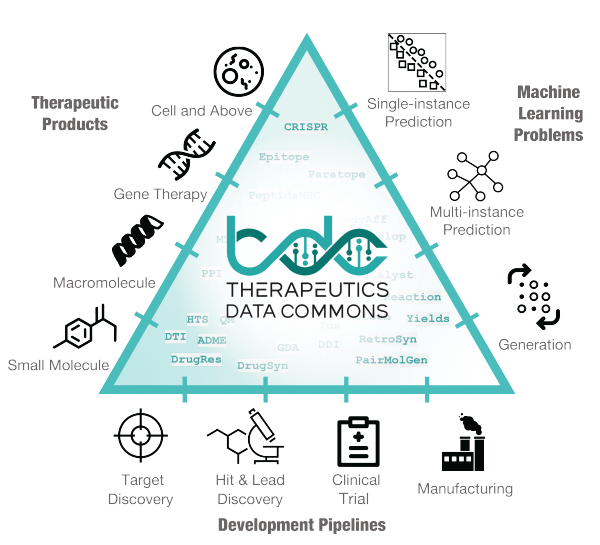
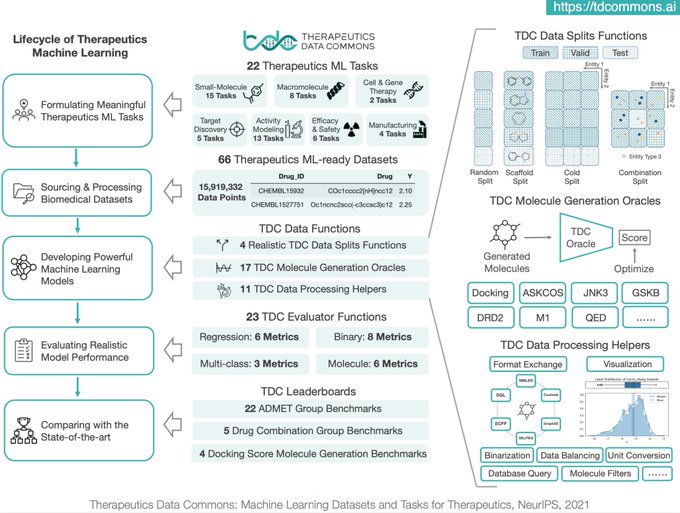
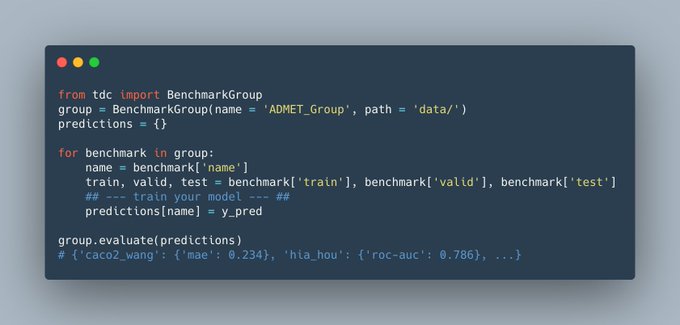
Excited to share preprint on Therapeutics Data Commons!
Paper:
Website:
TDC is a unifying framework across the entire range of
#therapeutics
#ML
. Ecosystem of tools, leaderboards & community resr
66 ML-ready datasets
22 ML tasks
1
62
220
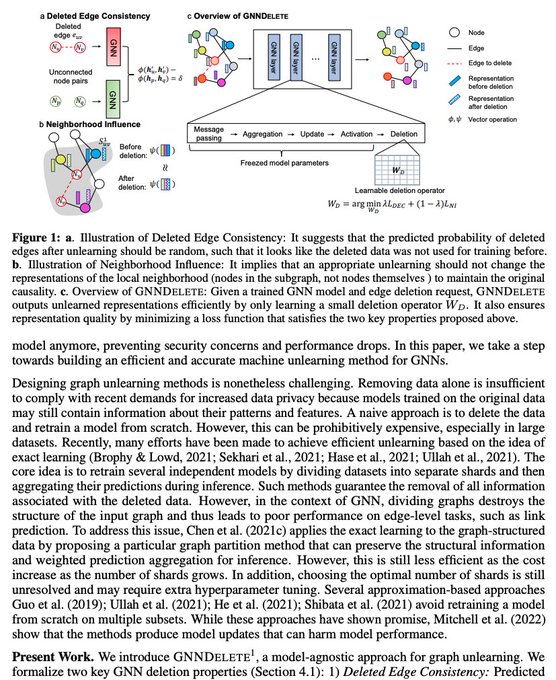
How can we delete data from a big model without training the model from scratch and sacrificing performance?
Having models unlearn is notoriously difficult
Our new
#ICLR2023
paper introduces a general strategy for unlearning on graphs
Exciting news 🥳: Our new research paper on Graph Unlearning is accepted at ICLR 2023
@iclr_conf
! Thanks to my coauthors Jiali Cheng, Huan He,
@_cagarwal
, and
@marinkazitnik
for the great collaboration!!
More info about our work will be out soon!
#ICLR2023
#ML
#GNN
#graphs
1
6
78
1
33
213
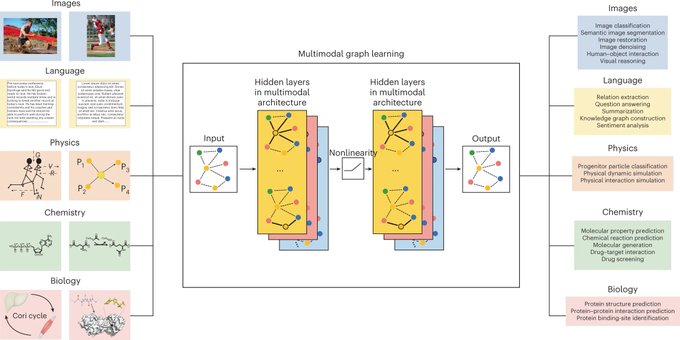
Excited to share the blueprint for multimodal graph learning in
@NatMachIntell
We envision graph AI playing a key role in future image-intensive, knowledge-grounded, and language-intensive systems 1/5
1
49
209
So excited about this new PhD program in Artificial Intelligence in Medicine
@Harvard
🎓🎓🎓
Join our info session on October 17th. Apply by December 1st
@HarvardDBMI
#ML4H
#AI4Medicine
#health
#research
2
32
186
Our
#ICLR2022
paper introduces Raindrop,
#graph
neural network learning on irregularly sampled and multivariate
#time
series
Check it out:
website:
code:
@xiangzhang1015
@MIT
@MITLL
@iclr_conf
#GNN
(1/7)
4
41
184
Thrilled that our lab has 4/4 papers accepted at
#NeurIPS2020
! Not bad for a lab just 5 months old at submission deadline. Congrats to fantastic students and collaborators
@_michellemli
@xiangzhang1015
@KexinHuang5
@IAmSamFin
@Emily_Alsentzer
@Harvard
@HarvardDBMI
@harvard_data
11
7
182
I was very happy to receive
@harvardmed
Young Mentor Award today.
Feeling fortunate to be working together with so many outstanding students
@HarvardDBMI
and am excited to see the bright future they are creating for themselves and our world
5
6
169
Excited to be elected to
@ELLISforEurope
as ELLIS Scholar in Geometric Deep Learning. Can't wait for new opportunities to further graph ML research on both sides of the Atlantic.
Many thanks to the ELLIS community for the support, esp
@mmbronstein
@maxwelling
@tacocohen
7
4
145
Super excited about winning
@amazon
Amazon Faculty Research Award on "Actionable Graph Learning for Finding Cures for Emerging Diseases" Thank you
@AmazonScience
for this recognition! More details soon
@HarvardDBMI
@harvardmed
@harvard_data
4
8
133
Excited to share our
@NeurIPSConf
papers accepted as spotlights!
@ZaixiZhang
@oq_35
#NeurIPS2023
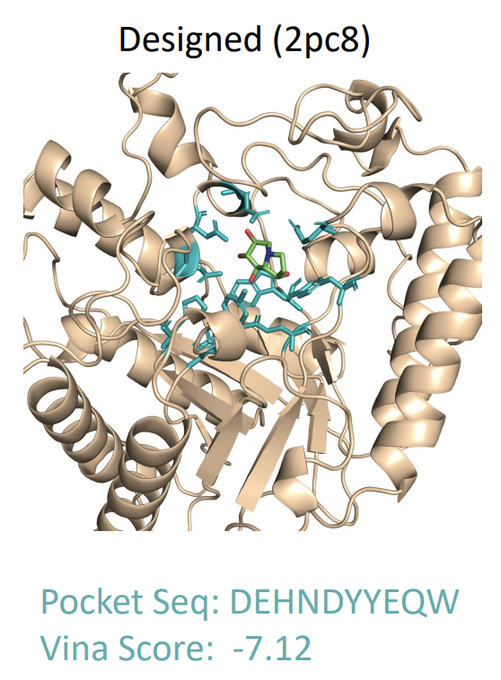
Full-Atom Protein Pocket Design via Iterative Refinement
🧬 Designing functional proteins for specific ligand binding is key for therapy & bio-engineering. A���
0
22
120
🚨 JOB ALERT 🚨
We are building the next generation of Therapeutics Commons
@ProjectTDC
!
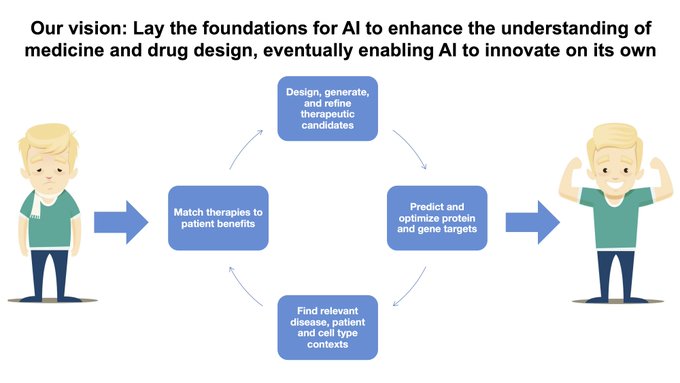
Our vision is to lay the foundations for AI & therapeutics, eventually enabling AI to learn on its own and acquire knowledge through continual refinement
We are seeking postdoctoral…
0
26
119
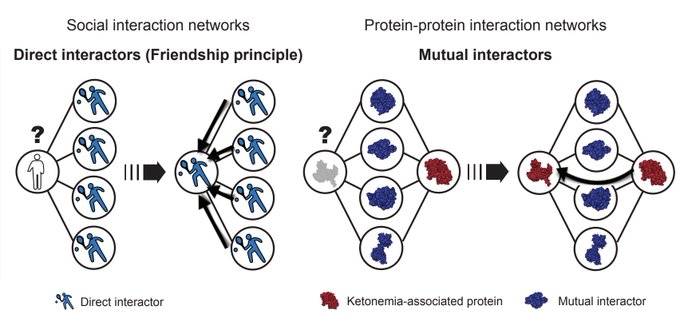
Today we share a paper introducing graph neural network
#GNN
powered mutual interactors to predict
#molecular
#phenotypes
Excited about this paper because it shows how thoughtful probing of AI model behavior can pave the way to improvements on many downstream tasks [1/6]
1
29
111
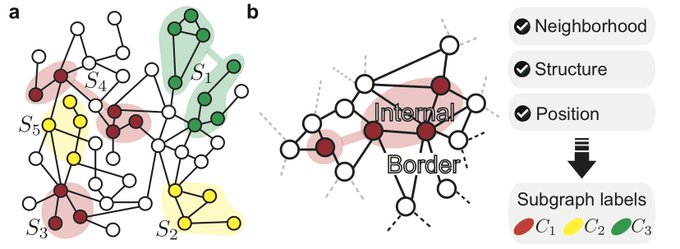
Enjoy Subgraph Neural Networks! Sub-
#GNNs
learn powerful
#representations
for
#subgraph
prediction. Going beyond predictions on nodes, edges and entire
#graphs
by
@Emily_Alsentzer
@IAmSamFin
@_michellemli
@HarvardDBMI
@harvard_data
@harvardmed
@MIT_CSAIL
3
20
102
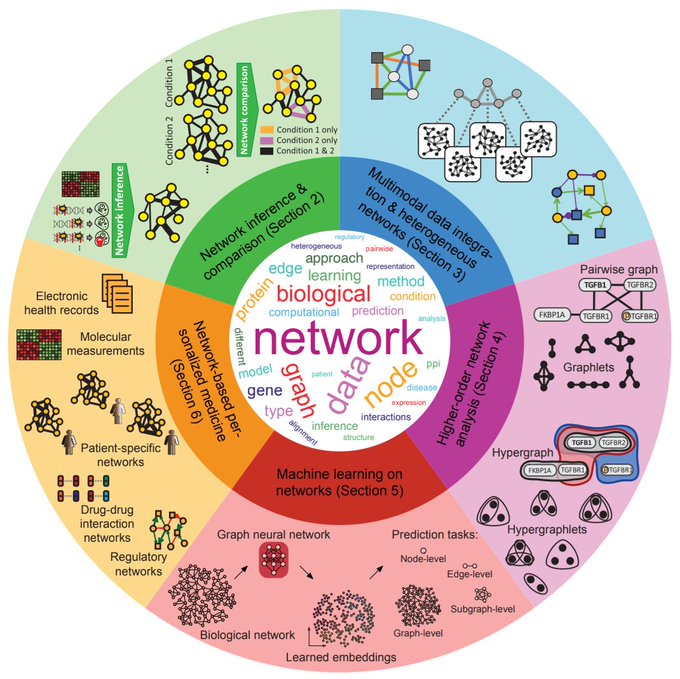
Envisioning a bright path ahead for networks, graphs, molecules, relational structures, and knowledge systems in biology and medicine 🧬🔬💊🩺
Our review+perspective paper on "Current and future directions in network biology" is online (). This has been a Herculean effort by many co-authors, thank you all! Special thanks to
@marinkazitnik
,
@_michellemli
,
@aydin_wells
, and all section coordinators.
1
132
407
0
10
99
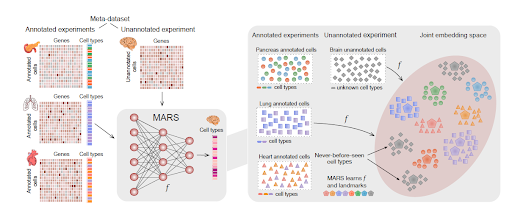
Hot off the press! A meta-learning algorithm for finding and naming novel cell types in single-cell datasets.
MARS uses a meta-learning strategy for annotating known cell types and identifying novel ones across single-cell RNA-seq datasets.
@jure
,
@mariabrbic
,
@marinkazitnik
,
@Rbaltman
0
62
164
4
13
94
We are thrilled to announce the National Symposium on Drugs for Future Pandemics (Nov 17-18)! Tune in for two days of visionary talks by stellar speakers. Registration is free and open
#futuretx20
3
28
90
Whoa, the current
@TheEconomist
edition reports on how AI can turbocharge scientific progress and lead to a golden age of discovery, echoing key insights from our recent
@Nature
paper,
0
21
84
Preparing for
#NeurIPS2023
! Nine from our group will attend, presenting papers, giving talks, and organizing workshops
@michellemli
,
@AdaFang
,
@YEktefaie
,
@ZaixiZhang
,
@oq_35
,
@TianlongChen4
,
@GDasoulas
,
@YepHuang
Find me giving four invited talks, I'm grateful for having a…
1
12
82
Many congratulations to co-authors
@WangQianwenToo
,
@KexinHuang5
@payal_chandak
@ngehlenborg
for the best paper award at
@icmlconf
Interpretable ML in Healthcare. So happy to work with amazing students and collaborators
@HarvardDBMI
#ICML2021
#XAI
#GNN
3
8
81
Thanks for organizing this,
@guadagonzalezzp
@logml2021
Live now: “Graph Representation Learning for Biomedical Discovery” by
@marinkazitnik
at
@logml2021
. Tune it to hear from an expert in the field how geometric deep learning is powering breakthroughs in biomedical discovery. Stream:
#MachineLearning
0
26
82
1
12
69
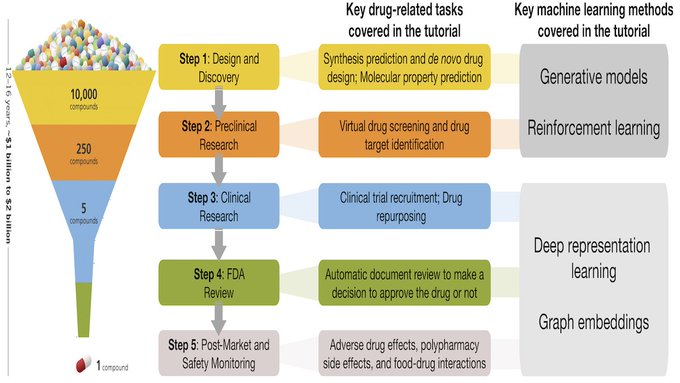
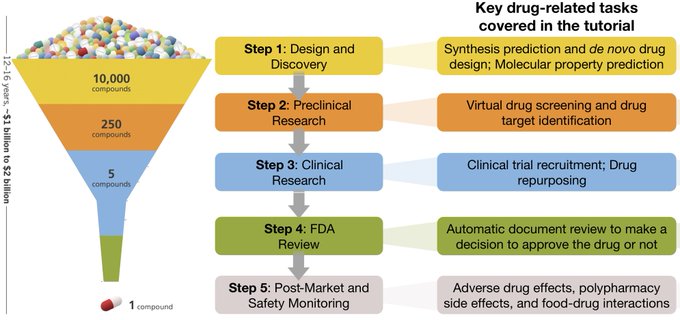
We will present a tutorial on Machine Learning for Drug Development at
#IJCAI2020
! Materials to follow on our website:
@IJCAIconf
#drugs
#networks
#AI
1
26
67
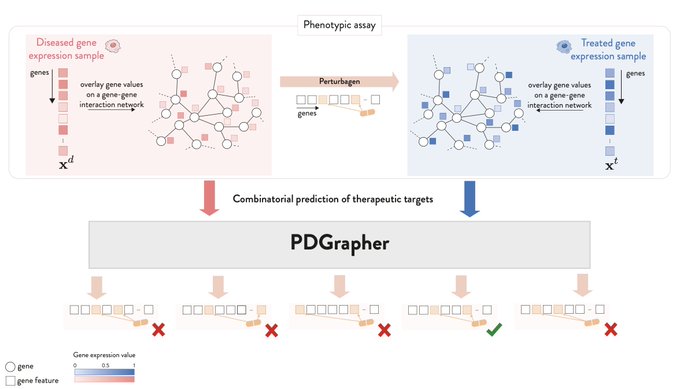
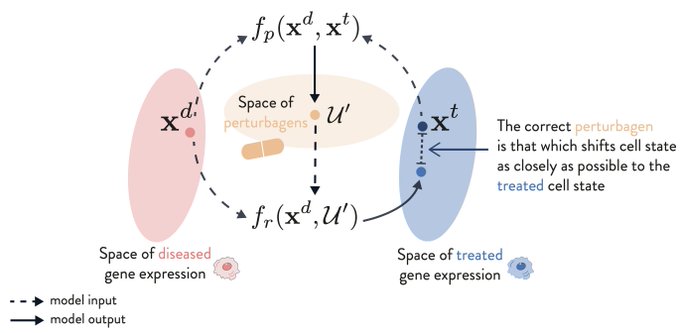
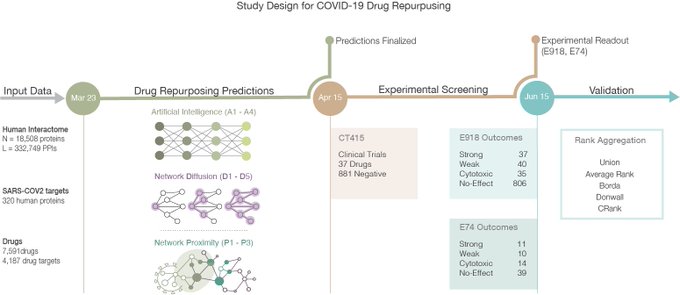
Very excited to share this research where we use
#graphML
and
#network
#medicine
to identify novel
#therapeutic
opportunities
0
13
66
Grateful for this collaboration with the brilliant minds
@PetarV_93
and
@pushmeet
@GoogleDeepMind
on how AI is reshaping the realm of science
#AI4Science
What could science at digital speed look like? 🚀
AI is poised to supercharge scientific discovery as we know it, by:
🔮 Exploring theories
🧪 Designing experiments
🔍 Analysing data
Find out how in
@Nature
. ⬇️
13
106
477
1
7
63
Mark your calendars for Nov 17-18! Join the discussion with leading experts in CS, bio, med, automation, and regulation about ways to innovate rapid therapeutics for future pandemics. Registration is free & open
#futuretx20
2
11
60
Excited for
#NeurIPS2022
-- We'll present two papers, organize two workshops and give some invited talks and panel discussions
#AI4Science
#biomedicalAI
[1/6]
1
4
59
Submit your latest research to Graph Representation Learning Workshop at
#ICML2020
Deadline: May 29
We encourage submissions on
#graph
representation learning, geometric deep learning, interdisciplinary applications, benchmarks, and research aiming to mitigate
#COVID19
Excited about Graph Representation Learning and its application across the sciences? Our Research Scientists
@PetarV_93
and
@jhamrick
are co-organising an
#icml2020
workshop on the topic -- consider submitting your latest research by 29 May! More information below:
1
35
116
0
12
58
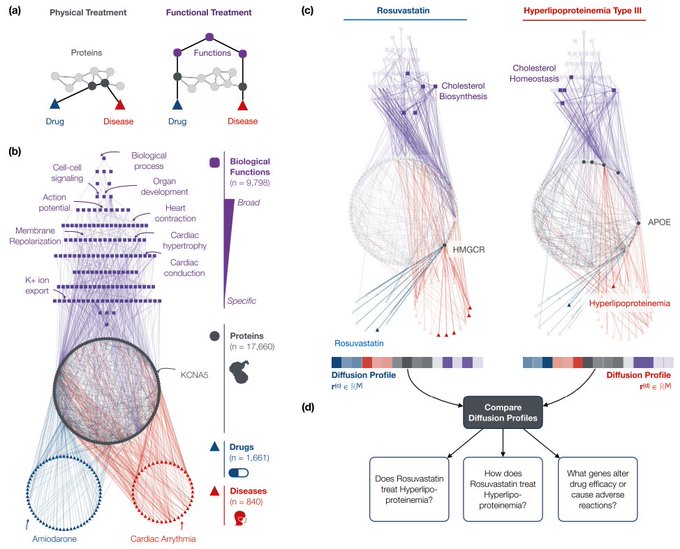
Great work,
@_camiloruiz
! We develop a multiscale interactome approach to explain disease treatments. It predicts drug-disease treatments, identifies proteins and biological functions related to treatment, and identifies genes that alter treatment's efficacy & adverse reactions.
.
@_camiloruiz
,
@marinkazitnik
and
@jure
develop the multiscale interactome, a powerful approach to explain disease treatment.
@StanfordBioE
@StanfordEng
@StanfordMed
@czbiohub
@HarvardDBMI
0
5
18
1
10
58
Congrats to
@payal_chandak
and
@KexinHuang5
on publishing PrimeKG, multimodal knowledge graph for precision medicine in
@ScientificData
🔬 🎉
Stay ahead of the curve with PrimeKG as it's continually updated with the latest data
With
@payal_chandak
and
@KexinHuang5
, we are excited to share PrimeKG, a precision medicine-oriented knowledge graph providing holistic and multimodal view of human disease (1/8)
4
49
227
1
19
56
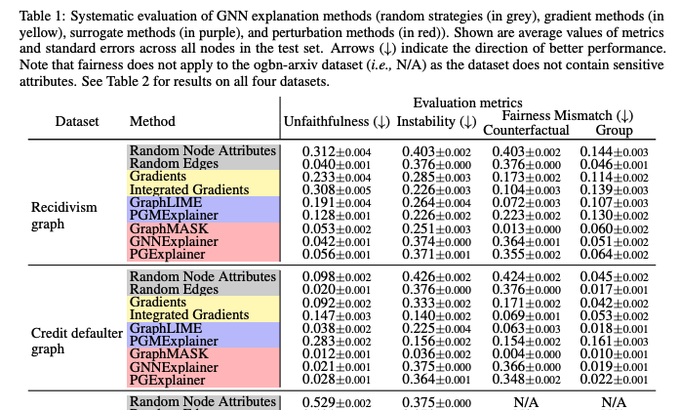
Our new
#AISTATS2022
paper introduces the first axiomatic framework for theoretically analyzing, evaluating, and comparing
#GNN
explainers.
Check it out:
w/
@hima_lakkaraju
@_cagarwal
(1/5)
@aistats_conf
3
13
56
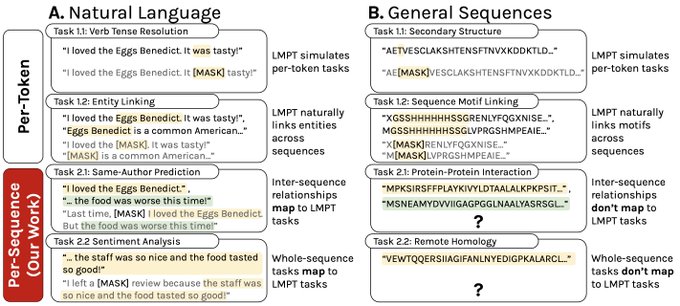
📢 1/7: Excited to share
@NatMachIntell
paper on language model pre-training and relational structure outside the realm of natural language
Thankful for a star team
@MattBMcDermott
, B Yap, P Szolovits
@HarvardDBMI
@harvard_data
@MIT
1
10
54
Excited for the future of AI in medicine that
@NEJM_AI
will help create
Dr.
@marinkazitnik
is an Assistant Professor of Biomedical Informatics
@harvardmed
with additional appointments
@harvard_data
and
@broadinstitute
of Harvard and MIT. Dr. Zitnik works on infusing knowledge, structure, and geometry into machine learning models.
2
9
38
1
9
54
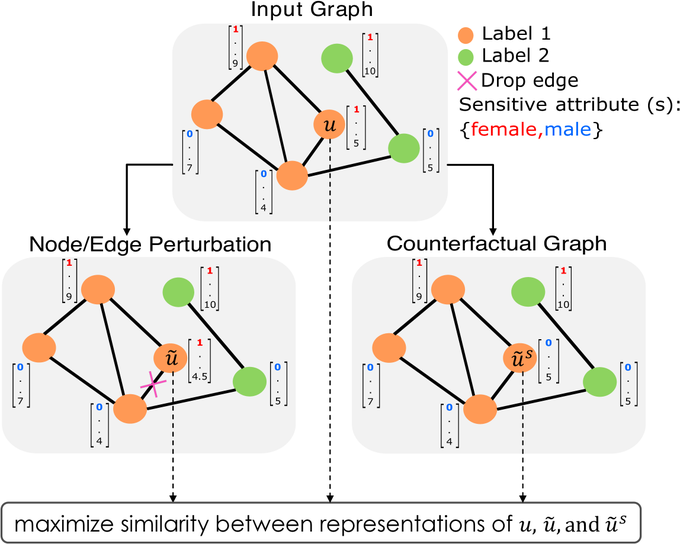
Very excited about this project! We need to start opening graph neural nets to high-stakes domains
#biomedicalAI
Our approach can be used with any
#GNN
to learn provably fair and stable embeddings
#graphML
#trustworthyML
@_cagarwal
@hima_lakkaraju
0
7
51
First talk in the session is David Kelley
@drklly
on sequence‐based modeling of single‐cell ATAC‐seq using convolutional neural networks
#biodata22
3
6
51
ML track
@iscb
#ISMB2022
starts on Wed at 10:30am CDT
Speakers
@DrAnneCarpenter
@suinleelab
@s_brunak
N. Beerenwinkel
@cbg_ethz
@GreeneScientist
@markowetzlab
@FertigLab
G. Rustici
@AstraZeneca
and many more
Excited to chair this track w/
@CataVallejosM
0
12
51
Excited about the future of
#multimodal
#AI
using
#graphs
to bring diverse modalities together
Check our preprint on graph-centric multimodal representation learning 👇
Led by fantastic students
@YEktefaie
@GDasoulas
@ayushnoori
collab w
@MahaFarhat
@HarvardDBMI
@harvard_data
0
14
48
It's an exciting double conference week,
#ISMBECCB2023
and
#ICML2023
. If you're attending, these talks and presentations are not to be missed!
At
@iscb
,
@_michellemli
will talk about her work with
@Emily_Alsentzer
on SHEPHERD, which is our latest zero-shot deep learning model…
1
6
50
Excited to share a new preprint on predicting molecular
#interactions
with skip-graph networks
Skip-
#GNN
uses 2nd-order interactions, which proved incredibly useful in
#bionets
over the last decade
Led by a fantastic student
@KexinHuang5
@harvard_data
0
7
49
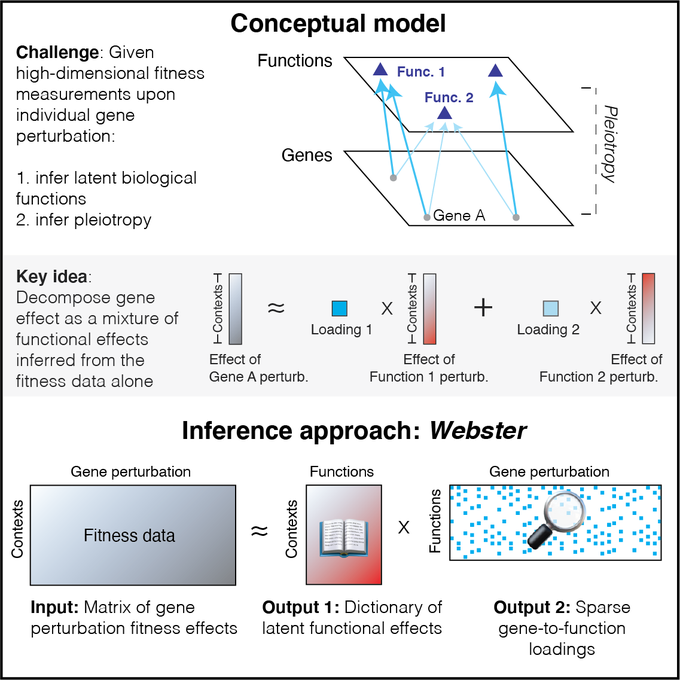
Genes can participate in multiple independent biological functions, a foundational genetic principle known as pleiotropy. We show how sparse approximation & learning can decipher pleiotropy in high-dim gene perturb datasets. Great work,
@joshbiology
!
#MLCB2021
Ever felt your favorite gene has *more than one function* after perturbing it in many cell contexts? Today at
#MLCB2021
, I'll discuss automatic genotype-phenotype inference from high-dimensional gene perturbation data using sparse approximation: [a 🧵]
6
82
357
0
9
48
Join us tomorrow
@iscb
#ISMB2022
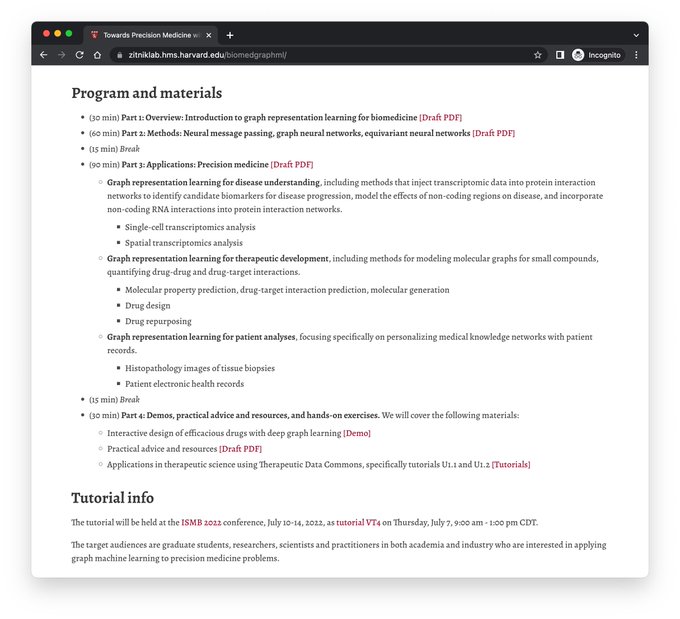
! How
#graph
#AI
can help advance precision medicine?
Advances in methods and key areas in
#disease
#therapeutics
&
#patients
, and demos, practical tips, and hands-on exercises
With star student
@_michellemli
@HarvardDBMI
0
12
48
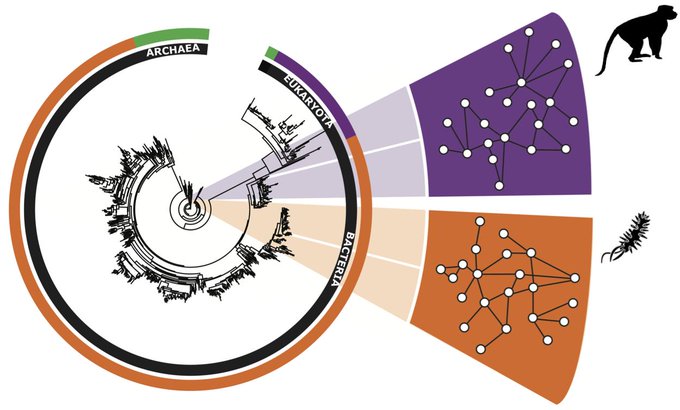
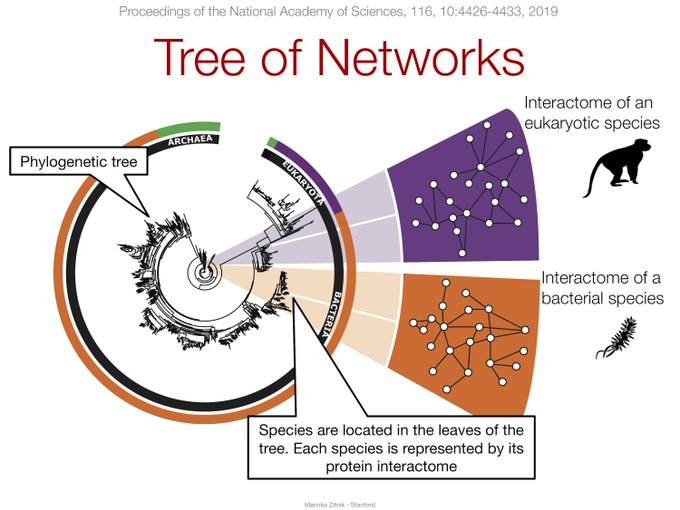
How do biological
#networks
change with
#evolution
? Our study just published in
@PNASNews
shows that evolution leads to resilient protein interactomes, which, in turn, are beneficial for organisms w/
@jure
, M.W. Feldman et al.
1
16
47
📢 New
@Harvard
Seminar Series on Efficient ML! 👇
Thanks
@schwarzjn_
for spearheading this effort!
@HarvardDBMI
@KempnerInst
@harvard_data
@harvardmed
@zakkohane
@ShamKakade6
🎓 Check out a phenomenal speaker lineup with an opening by LLM superstars
@sarahookr
@cohere
and…
🚨 Excited to announce our new
@Harvard
seminar series on Efficient ML! Please join LLM superstars
@sarahookr
(
@cohere
) &
@valentina__py
(
@allen_ai
) for our kick-off meeting next Monday!
Open to everyone! 📅
🔗
@harvard_data
@harvardmed
@KempnerInst
4
35
155
0
7
48
So excited about this new venue! Submit your finest work in graph representation learning, geometric deep learning, graph AI
0
7
47
New preprint: Meta-learning for identifying and naming cell types, even cell types that have never been seen before and do not exist in the training data
#MetaLearning
#TabulaMuris
#TabulaMurisSenis
#aging
1
7
44
Excited about this
#NeurIPS2022
paper on broadly generalizable self-supervised learning
Congrats to fantastic students
@xiangzhang1015
Z. Zhao
@HarvardDBMI
@harvard_data
and collab w T. Tsiligkaridis
@MITLL
project page:
code:
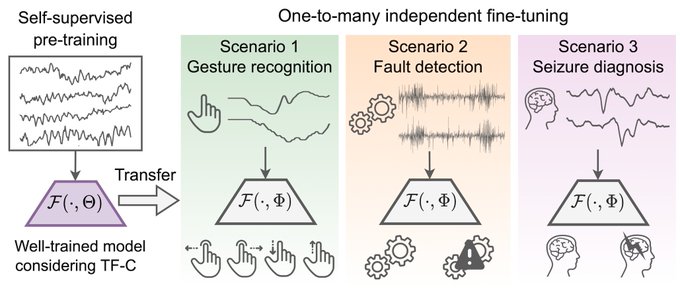
Can we infuse
#structure
into a time series (
#TS
) model from a diverse dataset so as to greatly improve
#generalization
on new TS coming from different datasets?
Yes, via a new principle called
#Representational
Time-Frequency Consistency (TF-C)
1/3
3
103
518
1
8
45
🔍 Join Professor
@marinkazitnik
's lab as a Postdoc
#Research
Fellow in
#AI
for Cancer Drug Discovery at
@Harvard
! Apply to lead the design, development, and implementation of novel AI methods for the analysis of clinical and biomarker
#data
in oncology:
0
2
5
0
9
45
So excited and honored that our research is recognized by the
@Bayer
Early Excellence in Science Award
Congratulations to Ruth Ley for obtaining the Otto Bayer Award 2020! Congratulations to Julia Mahamid, Josep Cornella, Nikolai Franzmeier and Marinka Zitnik for obtaining the Early Excellence in Science Awards 2020!
@CornellaLab
@nfranzme
@marinkazitnik
2
1
17
5
2
45
Excited about our new work accepted to
#ICLR2020
as full paper with spotlight!
@iclr_conf
TL;DR: We develop strategies for pre-training Graph Neural Networks and study their effectiveness on multiple datasets, GNN architectures, and diverse downstream tasks
#molecules
#proteins
(1) Strategies for Pre-training GNNs (spotlight) joint work with
@liubowen16
@marinkazitnik
@vijaypande
@percyliang
@jure
2
3
41
0
4
43
Thank you
@BiswasFamilyFdn
for your vision and generosity
Learn more about our project CURE-Bench to build and evaluate all-disease foundation models for identifying clinically relevant drug repurposing hits
Thankful to
@BiswasFamilyFdn
for supporting…
4
10
41
We are guest-editing special issue on Graph Embeddings and Deep Learning for Network Biology
@ComputerSociety
@TheOfficialACM
IEEE/ACM TCBB
Call:
Submit your finest work by July 30, 2020!
Spread the word!
#machinelearning
#graphs
#compbio
#medicine
0
12
38
Excited to see this published: We examined 10,443,476
#adverse
#drug
event reports, spanning 19,193 adverse events and 3,624 drugs, to extract insights into safe
#medication
use & how adverse events vary across
#patient
groups
Detailed tweetorial to follow shortly
#data
#AI
In a recent Article,
@xiangzhang1015
,
@marissa_sumathi
and
@marinkazitnik
analyze large-scale patient safety data to reveal demographic disparities of drug safety and identify at-risk patients during a pandemic.
1
3
25
1
8
40
We are thrilled to release the Open Graph Benchmark!
OGB contains numerous biomedical datasets, including protein interaction nets, cross-species graphs, drug-drug interaction nets, and biomedical knowledge graphs
#ML
#graphs
#networks
0
13
39
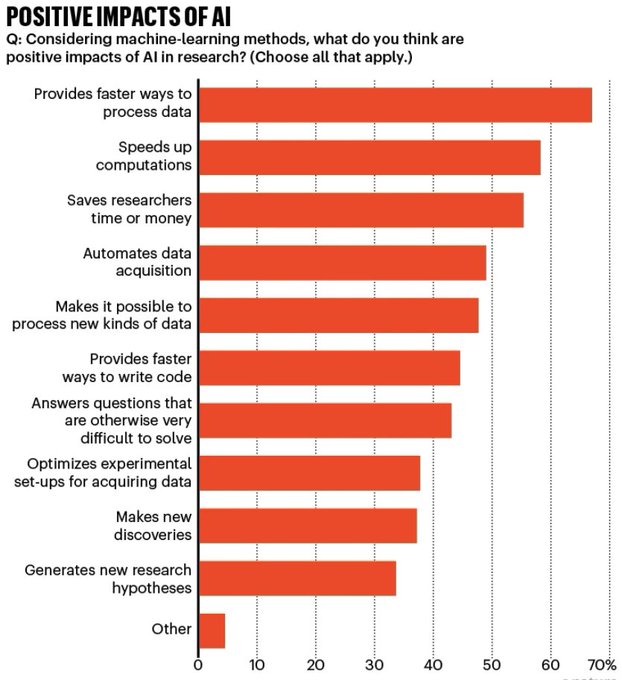
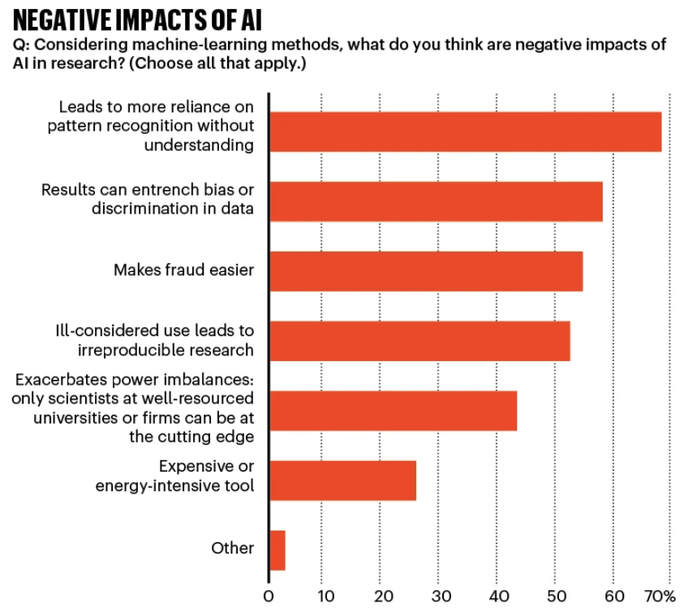
AI and science:
@Richvn
@Nature
polled 1,600 researchers around the world about their views on the rise of AI in science, including machine learning and generative AI
#AI4Science
@AI_for_Science
Among those who used AI in their research, more than 25% felt that AI tools would…
1
8
38
Excited to present a tutorial on
#ML
for drug development and discovery at
@IJCAIconf
.
Jan 6, 7-10pm EST / Jan 7, 9am-12pm JST w/
@jimeng
and Danica Xiao
@IQVIA_global
0
7
40
So glad to see this! We started to tackle this challenge through Therapeutics Data Commons, an ecosystem of AI/ML-ready datasets for therapeutics, together with tools, leaderboards and community resources like meaningful data splits
ICYMI: if you have interest in dataset generation for
#AI
, check out the new
@NIH_CommonFund
Bridge2AI program. Details here: …
@NLMdirector
@NHGRI_Director
@NCCIH_Director
@NIBIBgov
@aao_ophth
@aaopt
@AMIAinformatics
@ARVOinfo
@SfNtweets
0
13
22
0
8
37
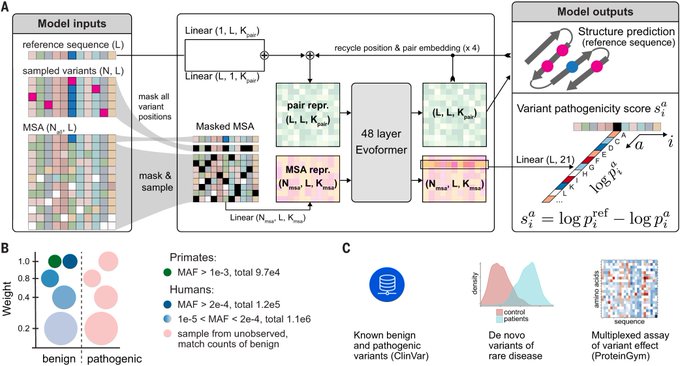
Such an excellent thread on AlphaMissense by
@simocristea
The human genome is gradually unravelling its secrets 🎁
AlphaMissense model
@ScienceMagazine
: one more path lit up by deep learning in exploring the code of life 🧬
We now know with high confidence if 89% of ALL missense variants are benign or pathogenic
Key contributions🧵🧵
11
177
747
2
6
38
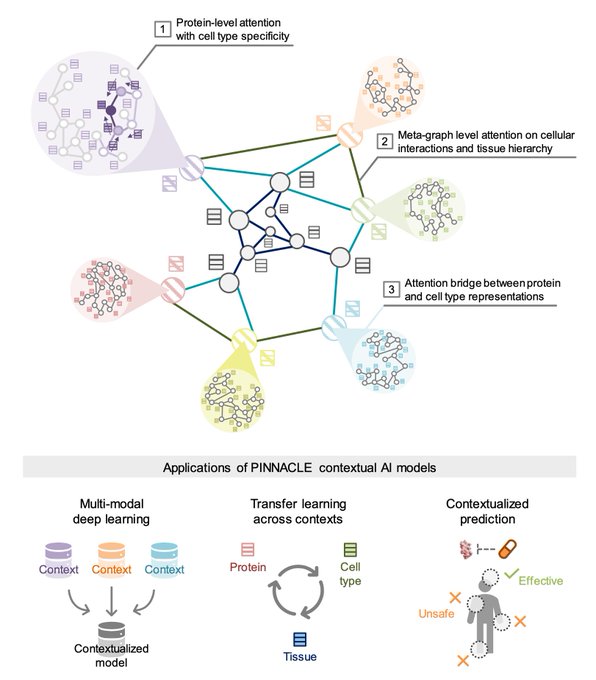
Phenomenal resource with ready-to-use embeddings and models through
@cziscience
Cell Census 🚀
Congrats to
@JCoolScience
@Paedugar
and the
#CELLxGENE
team 👏
Thanks for showcasing PINNACLE, our contextual AI model for single-cell protein biology, and scCIPHER, our multimodal AI…
🚨1/ New to CZ
#CELLxGENE
: models & embeddings that integrate up to 36M cells in the Census corpus.
Use embeddings to explore the corpus directly, or download the models to run your own data through them to enable direct comparisons to the reference. 🧵
4
48
190
0
8
38
Submit your papers to our Graph Learning Benchmarks
@GLB_Workshop
@TheWebConf
#TheWebConf
Call for contributions to establish novel ML tasks and novel graph-structured data
@Jiaqi_Ma_
@jiong971
@epsiloncorrect
@danaikoutra
@meiqzh
0
13
38
Excited for a day at the
#AACR2022
! Will be speaking on deep learning for interactomes, targeting disease-perturbed networks, and
@ProjectTDC
#AACR2022
#graphAI
#drugdiscovery
0
3
36
Exciting new resources to power openly available AI models of human cells and catalyze medical research
1
7
37
Excited about our
@NeurIPSConf
2021 workshop on AI for Science: Mind the Gaps
Introducing AI for Science — a
@NeurIPSConf
2021 Workshop! Our workshop focuses on bridging the gaps between machine learning and science. We have a stellar lineup of speakers! Submit your work and sign up for mentorship program now:
2
52
189
0
4
37
Grateful for the chance to discuss AI's 2024 prospects with
@NatMachIntell
. We covered LLM progress, multimodal AI, multi-task agents, and the crucial issue of bridging the digital divide across communities and world regions
Appreciating the insightful exchange,
@LCVenema
!
0
2
37
We are looking forward to continuing our research on actionable artificial intelligence for finding cures for emerging diseases
Congratulations to the 101 recipients of the 2020
#AmazonResearchAwards
, who represent 59 universities in 13 countries. Each award supports the work of one to two graduate or postdoctoral students for one year, under the supervision of a faculty member.
0
14
81
3
3
35
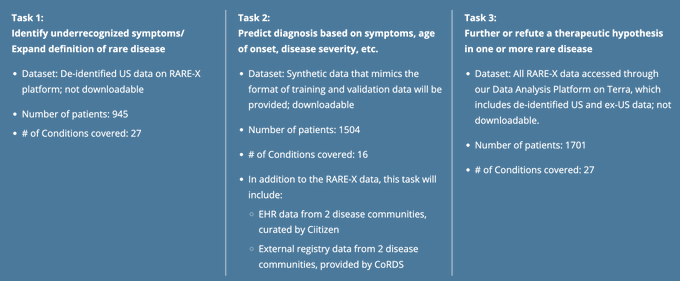
Only 5% of 10,000+
#RareDiseases
have FDA-approved treatments
Excited about
#XcelerateRARE
data challenge to utilize patient-provided data and help solve these critical unknowns
@GlobalGenes
@RARE_X_
0
4
36
New preprint: Machine Learning for Integrating
#Data
in
#Biology
and
#Medicine
: Principles, Practice, and Opportunities - w/ Francis Nguyen, Bo Wang,
@jure
, Anna Goldenberg,
@michaelhoffman
0
25
33
Looking forward to an exciting program at
@NSF
workshop on future directions in
#network
#biology
#NetBio
@ProfMilenkovic
@anthonygitter
@haiyuanyu
@ProfMonaSingh
@sroyyors
@compbiologist
@benjraphael
@xanderpico
@GysiDeisy
@bozdags
@pedjagogue
++
2
10
35
Join us
@NeurIPSConf
2021 "AI for Science" Workshop — Monday, Dec 13, 8a-6p ET
@AI_for_Science
Great day to celebrate AI achievements in scientific discovery and highlight open challenges that need to be addressed to move the field forward
#AI4Science
0
7
35
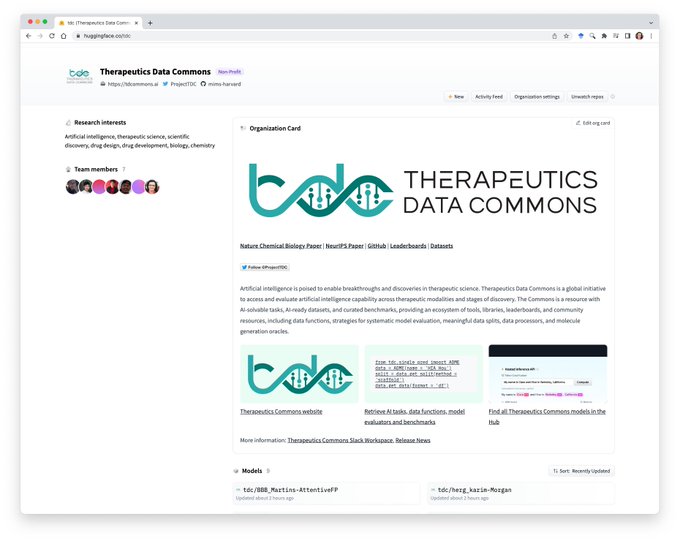
Therapeutics Commons
@ProjectTDC
now has its own
@huggingface
🤗 model hub
📢📢1/ Thrilled to share
@ProjectTDC
Therapeutics Commons 0.4.0
We have a new interface allowing users to easily access and leverage large pre-trained models for direct prediction or fine-tuning on downstream tasks
Check out our
@huggingface
Hub
2
18
45
0
3
35
Excited to share how combining biological LLMs with knowledge graph AI enables zero-shot prediction in drug discovery 🎯💊
Thanks
@HarvardCMSA
for organizing this conference
1
5
35
We are very excited to be co-organizing a workshop at
#AAAI2021
on Trustworthy
#AI
for
#Healthcare
! We have a stellar lineup of speakers. Details to follow soon!
@RealAAAI
w/
@cmuptx
, Byron Wallace, Jennifer G. Dy, and Eric P Xing.
1
4
34
Had a great time today giving a seminar
@cziscience
@czbiohub
about Therapeutics Data Commons
@ProjectTDC
,
#AI
-driven
#therapeutic
action, drug
#repurposing
, and much more that is in the works with students & collabs
0
2
34
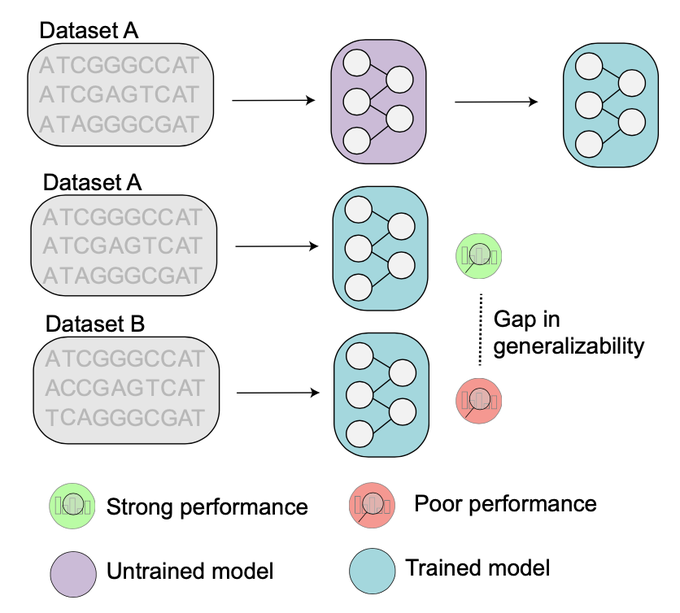
How well do your AI models perform on new molecular sequences?
@YEktefaie
🧵 👇
Understanding generalizability - how well an AI model works on new data - is crucial in biology. This challenge grows with foundation models, large pre-trained models that promise to better predict…
0
6
33
One big question in
#AI4Science
@axios
: Can we build more than a scoop machine?
Instead of having AI predict discoveries and who will make them, the researchers tuned the model to "avoid the crowd," and it found scientific blindspots — surprising combinations and predictions…
0
6
33
Congrats to graduate student Michelle Li
@_michellemli
@HarvardDBMI
on being awarded
@NSFGRFP
Graduate Fellowship! Well deserved! Thrilled to see what awesome things you do next.
1
1
33
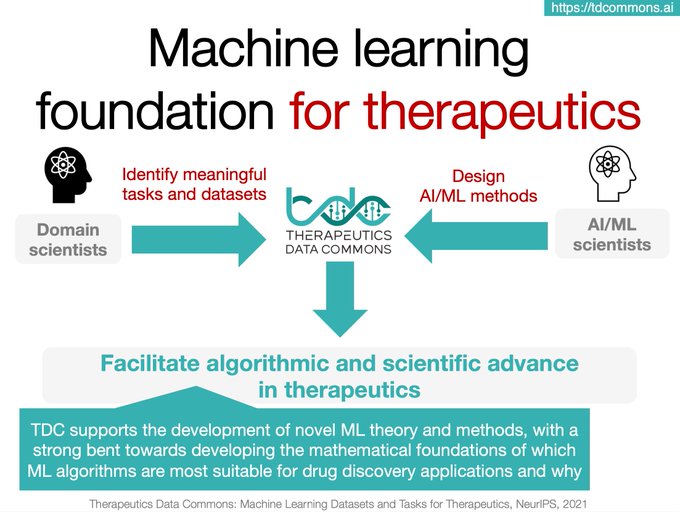
In our Nature Chemical Biology
@nchembio
paper, we describe Therapeutics Data Commons as providing
#AI
foundation for
#therapeutic
science
#drugdiscovery
@ProjectTDC
@HarvardDBMI
Collab
@cwcoley
@MIT
@jure
@stanford
@jimeng
@UofIllinois
C Xiao & students
Excited to share our new paper in Nature Chemical Biology
@nchembio
AI is poised to transform
#therapeutic
#science
The Commons is an initiative to access and evaluate
#AI
capability across therapeutic modalities and stages of discovery 1/4
1
19
72
2
6
32
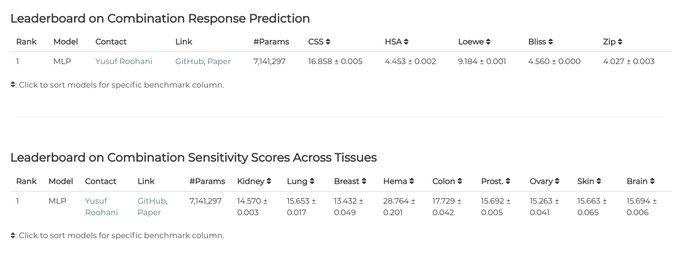
We are glad to announce Therapeutics Data Commons
@ProjectTDC
now has the leaderboard for cancer drug combinations!
Kudos to
@yusufroohani
@KexinHuang5
@WenhaoGao1
@yzhao062
Tianfan Fu
0
7
33
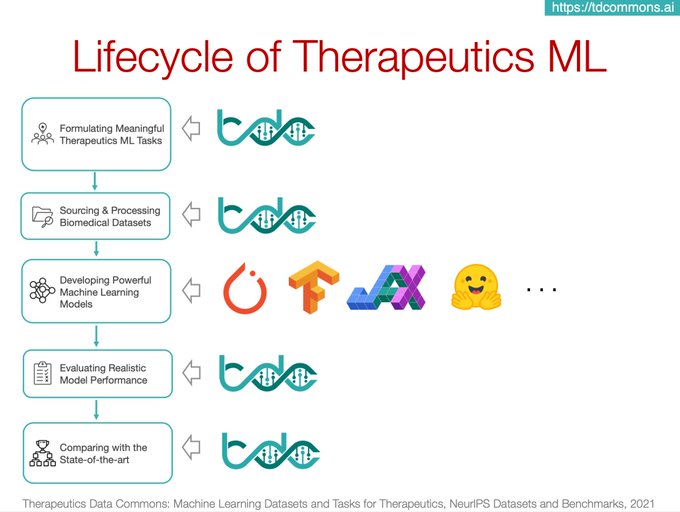
Therapeutics Data Commons is a collection of
#ML
tasks spread across areas of
#therapeutics
22 tasks
67 datasets
Therapeutics ML offers incredible opportunities for innovation and impact
TDC informs ML model dev, valid and transition into production & clinical implem. Join us!
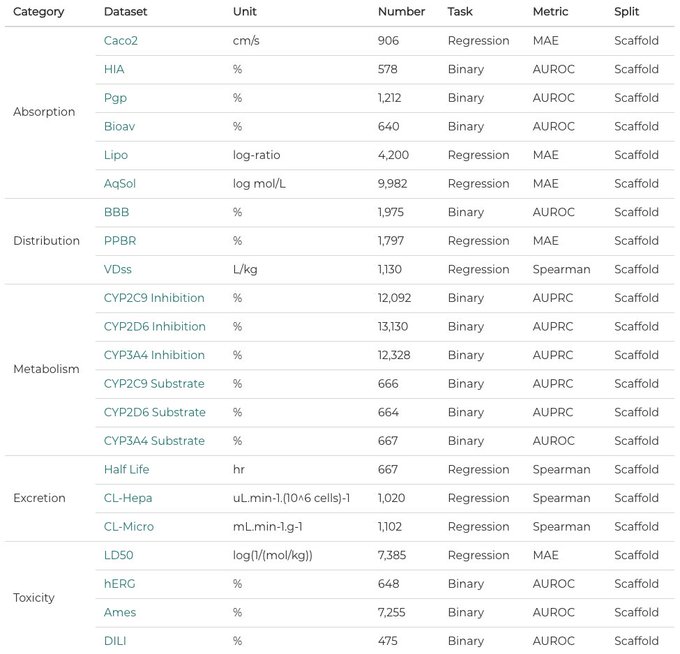
Happy to share the first leaderboard of TDC: 22 new & important ADMET property prediction datasets for molecular ML model! Submit your model to TDC!
More info:
GitHub:
@marinkazitnik
@TianfanFu
@WenhaoGao1
@yzhao062
@yusufroohani
0
9
28
0
7
31
@xiangzhang1015
@MITLL
@Harvard
@HarvardDBMI
Results suggest models can be trained on a source TS dataset and deployed on a range of target TS datasets without retraining, resulting in performance that is better than that of state-of-the-art & state-of-practice models 2/3
1
1
29
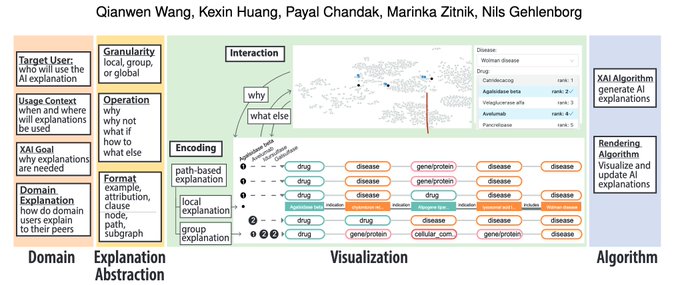
Congrats
@WangQianwenToo
for winning Best Paper Honorable Mention award
@ieeevis
🏆
Fusing
#visualization
with
#ML
is increasingly important to ensure
#AI
outputs are actionable, support transition into biomedical implementation, and more
Thankful for collab w
@ngehlenborg
Explaining "why and what else" in hypotheses generated in AI-driven (Graph Neural Network) drug repurposing is a UI challenge taken on by
@marinkazitnik
@ngehlenborg
@IEEEXplore
#machineLearning
#pathways
0
9
16
0
3
29
@xiangzhang1015
@MITLL
@Harvard
@HarvardDBMI
Paper:
Data:
Code:
Enjoyed working on this project, which is led by postdoc
@xiangzhang1015
@HarvardDBMI
with invaluable help from
@Harvard
undergraduates and
@MITLL
collaborators 3/3
2
4
28
Looking forward to it, thanks
@trustworthy_ml
!
📢 1/ We are delighted to have Marinka Zitnik
@marinkazitnik
from
@Harvard
with us for the next TrustML seminar on Thursday, July 22nd, 12pm ET! 🎉🎉
Register here: .
Check this thread for the speaker and talk details! 👇
1
4
22
0
2
27