Maria Brbic
@mariabrbic
Followers
1,386
Following
193
Media
13
Statuses
150
Assistant Professor of #computerscience at @EPFL @ICepfl @epflSV Previously @Stanford #machinelearning #compbio
Joined September 2020
Don't wanna be here?
Send us removal request.
Explore trending content on Musk Viewer
Luka
• 255960 Tweets
Caitlin
• 66054 Tweets
Dallas
• 61770 Tweets
Gobert
• 54161 Tweets
Mavs
• 51848 Tweets
#超超超超ゲーマーズ
• 33485 Tweets
Soto
• 26593 Tweets
Naz Reid
• 24408 Tweets
Anthony Edwards
• 22330 Tweets
Mavericks
• 21701 Tweets
DPOY
• 20399 Tweets
Sparks
• 17934 Tweets
Marlon
• 16750 Tweets
Draymond
• 15509 Tweets
Lively
• 12981 Tweets
マイフリ
• 12957 Tweets
サクナヒメ
• 11869 Tweets
グーフィー
• 10492 Tweets
クマフェス
• 10437 Tweets
Last Seen Profiles
Pinned Tweet
Our 🪐SATURN method is now out in
@naturemethods
!
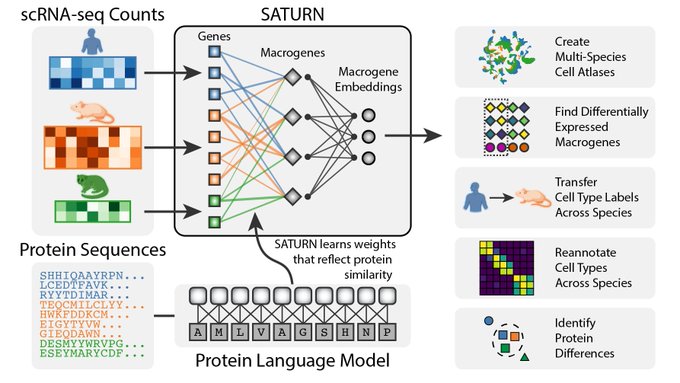
SATURN paves the way for universal cell embeddings, enabling integration of datasets across different species 🐒🐁🧍🐟🐸 Using protein language models, we encode biological meaning of genes in scRNA-seq datasets.
SATURN performs cross-species integration and analysis using both single-cell gene expression and protein representations generated by protein language models.
@jure
@YanayRosen
@mariabrbic
@yusufroohani
1
36
149
2
22
120
I'll be joining EPFL
@EPFL_en
as an assistant professor this fall! I'm immensely grateful to my mentors, collaborators, colleagues and everyone who supported me. Thank you all!
Excited to continue my research at the intersection of ML and biomedicine!
Thrilled to see brilliant minds joining
#EPFL
! Such an honour to welcome Maria Brbic (
@Stanford
), Daryl Yee (
@MIT
), Martin Hairer (
@imperialcollege
,
#FieldsMedal
2014) and Xue-Mei Li (
@imperialcollege
) to our faculty
via
@ETH_Rat
@Conseil_EPF
1/2
5
17
225
24
11
262
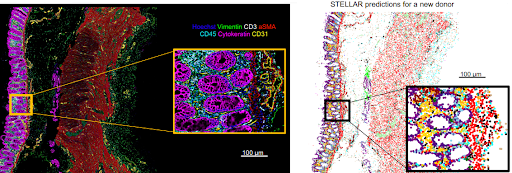
Proud to share that our STELLAR 💫 method for annotating spatially resolved single-cell data is now out in
@naturemethods
! Incredible collaboration and team
@kaidicao
@johnhickey22
@yuqitan3
@SnyderShot
@GarryPNolan
@jure
as part of the Human BioMolecular Atlas Program
@_hubmap
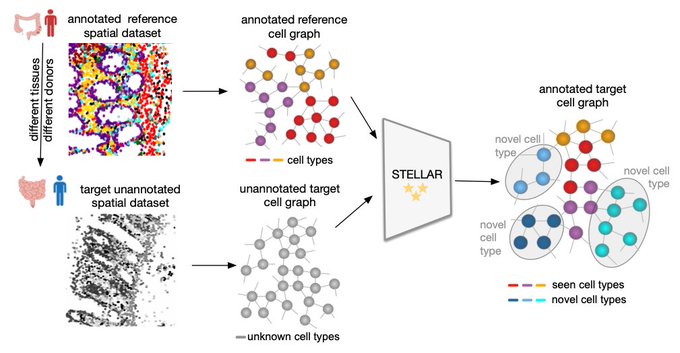
Introducing STELLAR: a geometric deep learning method for cell type discovery and identification in spatially resolved single-cell datasets.
@jure
@GarryPNolan
@SnyderShot
0
53
206
5
41
175
Excited to share our STELLAR 💫 method for annotating spatial single-cell data! STELLAR has been used to annotate CODEX
@_hubmap
data.
Wonderful collaboration with amazing team
@kaidicao
@johnhickey22
@yuqitan3
led by
@jure
@SnyderShot
@GarryPNolan
1
25
130
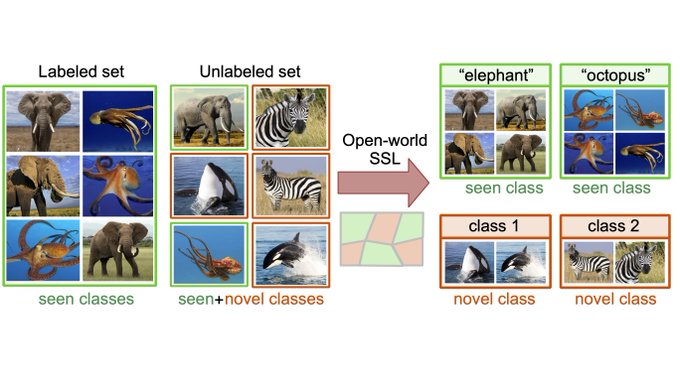
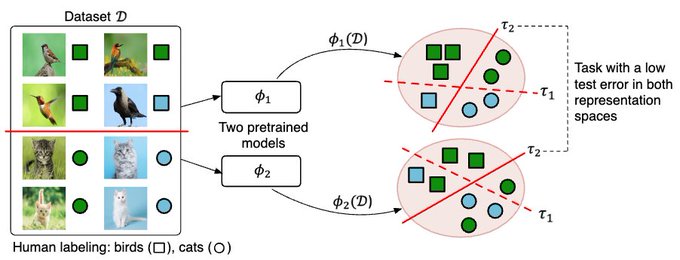
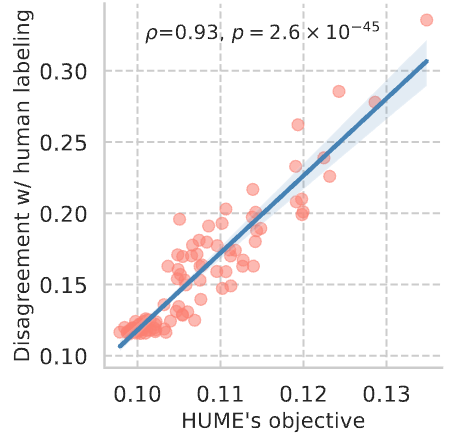
How to infer human labelling of a given dataset in a model-agnostic way?
Check our new method HUME accepted at
@NeurIPSConf
as
#spotlight
!🌟 HUME provides a new view to tackle unsupervised learning.
Kudos to my fantastic PhD student
@artygadetsky
!
Paper
1
22
112
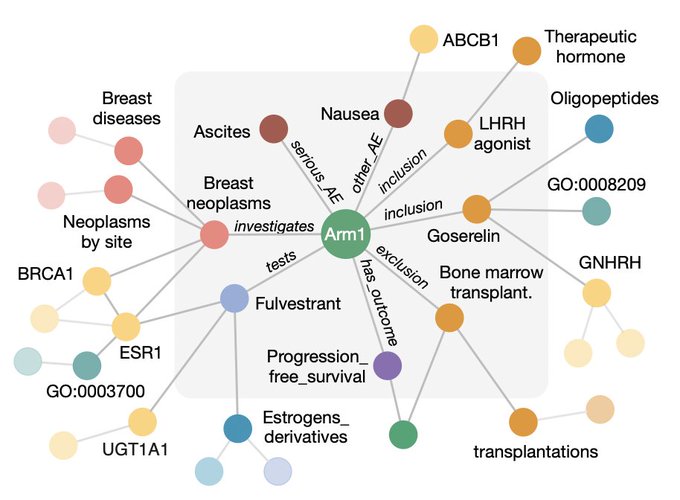
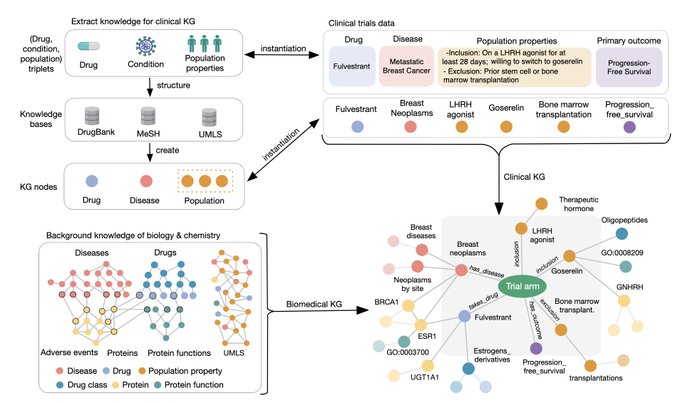
Excited to share our PlaNet 🌏method that reasons over population variability, disease biology, and drug chemistry using a massive clinical knowledge graph. We apply PlaNet to predict outcomes of clinical trials.
w/
@michiyasunaga
@agrwalprabhat
@jure
2
18
99
Absolutely thrilled that 3/3 submissions from our lab have been accepted at
#ICML2024
! A big shoutout to my fantastic students
@ShuoWen18
@artygadetsky
@GrcicMatej
@YulunJiang
for this achievement!
Stay tuned—we'll be posting the papers online soon!
3
7
97
Honored and humbled to receive the Early Career Award 🏆. Thank you
@ISBSIB
for this recognition!
Immensely grateful to my mentors and colleagues for the support through this journey🌟
For her research bridging machine learning and biomedicine and her strong involvement in promoting diversity, equity and inclusion in computer science, Maria Brbic (
@mariabrbic
) from
@EPFL_en
receives the 2023 Early Career Award! Congratulations!
@epflSV
#SIBawards
#bc2basel
1
5
10
19
8
97
Our tutorial on Meta-learning for Biomedicine is happening tomorrow at
#ISMB
! W/
@jure
@chelseabfinn
We made all slides available at
0
12
65
Interested in how to bridge labeled and unlabeled data in biology and biomedicine? We are organizing a tutorial on Meta-learning for biomedicine at
#ISMB
. Stay tuned for slides and updates!
w/
@chelseabfinn
and
@jure
More details at:
1
8
62
Super excited about our latest SATURN 🪐 method towards creating universal cell embeddings!
With amazing team
@YanayRosen
@yusufroohani
@KyleWSwanson
@jure
Can we create universal cell embeddings that capture the biological meaning of genes?
We present SATURN🪐 a method that integrates datasets across species by coupling RNA expression with protein embeddings.
w/
@mariabrbic
@yusufroohani
@jure
Preprint:
1
35
216
1
8
43
Thrilled to see our Aging Fly Cell Atlas now out in Science! Wonderful collaboration with
@HongjieLi5
@TzuChiaoLu
@StephenQuake
@HenriJasper
Our Aging Fly Cell Atlas is out. Do different cell types age at the same rate? No. Which cell types age faster? Fat, liver and muscle. Which cell types age slower? Neurons. Great collaboration with
@StephenQuake
@HenriJasper
@mariabrbic
@TzuChiaoLu
17
139
460
2
7
39
Video of our ISMB/ECCB tutorial on Meta-learning for Bridging Labeled and Unlabeled Data in Biomedicine is released by
@iscb
! With
@jure
and
@chelseabfinn
Link:
Tutorial website with slides and additional information:
1
11
32
We are accepting submissions for ICML Computational Biology workshop!
#ICML2023
#Hawaii
#ComputationalBiology
Your submissions are highly welcomed! Submission deadline is May 17th, 2023.
📢 Calling all
#CompBio
enthusiasts! Our ICML 2023 Computational Biology Workshop is now open for submissions! Don't miss this opportunity to showcase your cutting-edge research in this exciting field. check more details at
#ICML2023
#ComputationalBiology
0
34
104
0
5
34
Consider submitting your work to the Machine Learning for Genomics Explorations (MLGenX) workshop at
@iclr_conf
! Join us to unite the worlds of
#MachineLearning
&
#Genomics
🧬
⏰Submission deadline: February 4, 2024 (AOE).
📢🚨 Excited to announce the Machine Learning for Genomics Explorations (MLGenX) workshop at
@iclr_conf
2024.
⏰ Submission deadline: February 4, 2024 (AOE).
🔗 Call for papers:
🗓️ Date: May 11, 2024.
1
9
31
0
10
33
Looking forward to giving invited talk at
#KDD
Workshop on Knowledge Graphs today 5:30pm ET!
I will talk about our work on constructing knowledge graph for predicting outcomes of clinical trials.
1
6
27
First first whole-organism single-cell atlas of the fly is released! It was a wonderful experience to be part of the team. Special shout-out to
@HongjieLi5
for his contributions on this tremendous effort!
Data portal:
Paper:
0
3
20
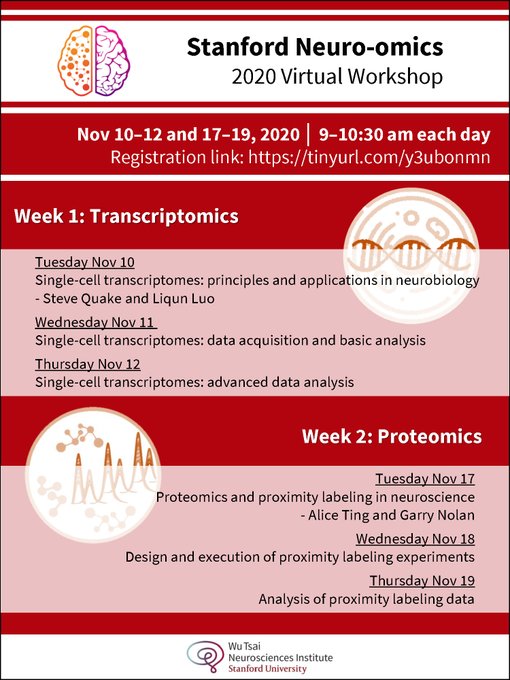
Stanford Neuro-omics initiative hosts a virtual workshop on transcriptomics and proteomics (Nov 10-12 and 17-19
@Stanford
). Liqun Luo, Steve Quake, Jure Leskovec
@jure
,
Alice Ting
@aliceyting
and Sergiu Pasca labs. Registration link:
0
3
17
Excited about our latest preprint! We present Aging Fly Cell Atlas 🪰
@HongjieLi5
@TzuChiaoLu
@drAOPisco
@jure
@lucksmith
@StephenQuake
Glad to share our Aging Fly Cell Atlas - our very first manuscript at Baylor
@bcmgenetics
@BCMFromtheLabs
.
@TzuChiaoLu
@mariabrbic
@drAOPisco
@jure
@lucksmith
@StephenQuake
7
27
155
0
0
10
Happy to give a talk
@MayoClinicCIM
on Meta-Learning in Single-Cell Genomics
Join us for the first of our
#AI
in Individualized Medicine Speaker Series on April 7! The webinar, featuring
@mariabrbic
& hosts
@JohnKalantari
& Kia Khezeli will focus on Meta-Learning for Novel Cell Discovery in Single-Cell Experiments. Register here:
1
4
0
0
3
9
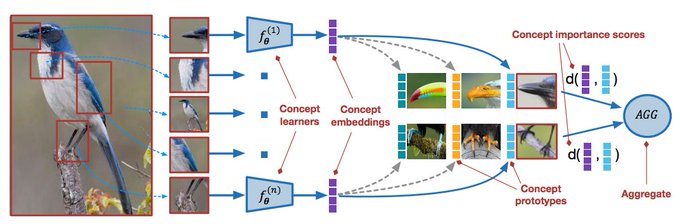
Excited that our COMET framework for concept-based meta-learning has been accepted to
#ICLR2021
! Joint work with
@caokd8888
@jure
Paper:
Website:
Code:
0
0
7
Congrats
@stephanedascoli
! 🎉We'll miss you at EPFL!
Thrilled to announce that I will be joining
@MetaAI
next month as a Research Scientist 😍
I will be working in the Brain & AI team on decoding language from neural activity, to hopefully help those which have difficulties to speak or type. Learn more here:
12
12
276
1
0
6
John is an amazing scientist and person! So happy for you
@johnhickey22
!! Can’t wait to see the work from your lab!
1
0
3
@RongFan8
@Nature
@YaleBme
@YaleSEAS
@YaleMed
@GoCasteloBranco
@karolinskainst
Wow, congrats
@RongFan8
!
0
0
2
@CirielloLab
@Nature
@elisa_oricchio
@RuxandraLambuta
@luca_nanni93
@dbc_unil
@unil
@EPFL_en
Congrats Giovanni!
0
0
2
@david_van_dijk
@EPFL_en
@epflSV
@ICepfl
@EPFLEngineering
It was a pleasure! Thank you for visiting us and for an inspiring talk!
0
0
1
@suinleelab
@twimlai
@bidumit
Having you as a speaker was an absolute delight! Thanks for joining us! 🙌
0
0
1
Congrats
@_camiloruiz
!!
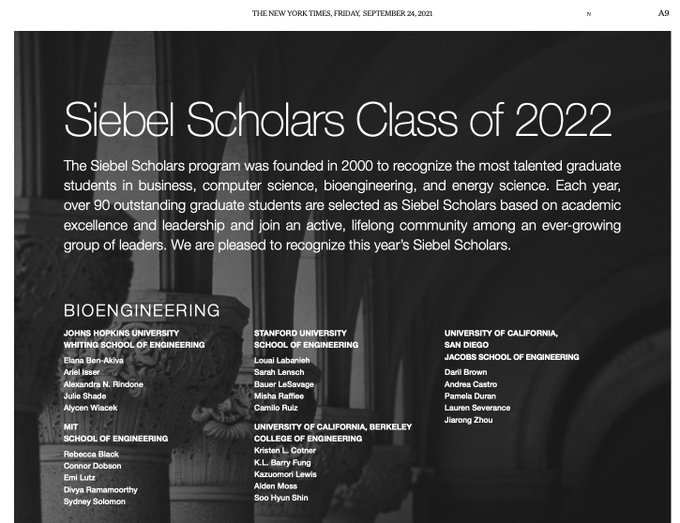
Honored to be named a 2022 Siebel Scholar for my PhD research with
@jure
! Picture from print edition of NYTimes.
5
5
70
0
0
1