Howard Chang
@HowardYChang
Followers
6,110
Following
73
Media
2
Statuses
170
Explore trending content on Musk Viewer
#บางกอกคณิกาep4
• 329240 Tweets
WE MUV ML
• 104117 Tweets
Colin
• 65649 Tweets
Apolinho
• 59180 Tweets
RPWP CONCEPT PHOTO 3
• 52503 Tweets
#세븐틴만_아는_SPELL로_주문거네
• 50741 Tweets
Silvio Luiz
• 40903 Tweets
KING KONG CONCEPT SPOILER
• 37476 Tweets
Ultimate Team
• 33138 Tweets
JUNHUI
• 28088 Tweets
Boebert
• 27889 Tweets
Blake
• 24223 Tweets
D-3 TO CHEESE
• 18029 Tweets
#岩本照誕生祭2024
• 17134 Tweets
2NE1
• 13590 Tweets
#SelahattinDemirtaş
• 11149 Tweets
Last Seen Profiles
Delighted to share our new paper
@MolecularCell
, discovering circular RNA translation start elements. Congrats to first author Chun-Kan Chen, with
@WeingartenShira
,
@segal_eran
,
@SnyderShot
,
@JswLab
,
@PeterKJackson
, and more.
4
68
255
X chromosome silencing by viral mimicry! Xist lncRNA pretends to be a virus to get the female cell to shut down the second X chromosome. Great work by Ava Carter, Jin Xu
@stanfordmed
and with Wuttke lab
@CUBIOCHEM1
.
The latest research from HHMI Investigator
@HowardYChang
and colleagues is an important step towards understanding how female cells carry out dosage compensation in mammals.
0
15
30
0
70
212
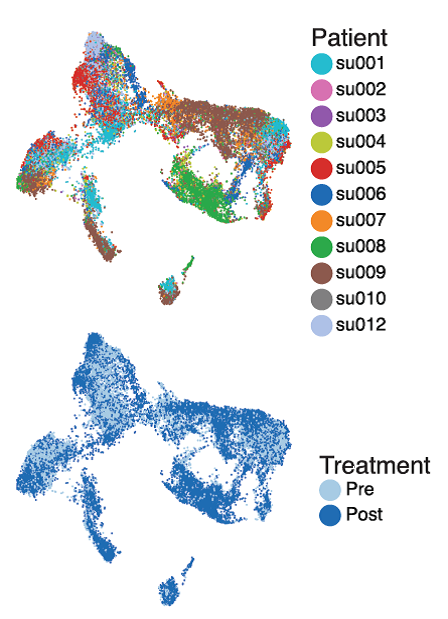
Delighted to share this work with
@WJGreenleaf
and
@10xGenomics
Massively parallel single-cell chromatin landscapes of human immune cell development and intratumoral T cell exhaustion
0
61
148
Clonal replacement of cancer fighting T cells by checkpoint blockade. Delighted to share our latest paper. Great team work with Katie Yost,
@Satpathology
, Dr. Anne Chang, and
@parkerici
!
After anti-PD-1 therapy in basal or squamous cell carcinoma patients, the CD8+ T cell response largely consists of an expanded and exhausted repertoire of new
#Tcell
clones |
#ImmunoTherapy
@HowardYChang
2
79
146
0
32
126
Excited to share our preprint on BABEL, a computational method to translate and synthesize multiomic profiles in single cells. . Wonderful work by grad student Kevin Wu and with
@james_y_zou
.
0
38
121
Celebrating this special honor with all of my mentors, lab members, and supporters who made it possible.
@HowardYChang
of
@StanfordMed
earns 2018
@theNASciences
Award in Molecular Biology for work on long noncoding RNAs!
#MolecularBiology
#epigenetics
18
12
109
Cancer genes beyond chromosomes--delighted to contribute to this new study. Great teamwork with Paul Mischel, Roel Verhaak, Vineet Bafna, and Bing Ren.
Along with 23 pairs of chromosomes, some of our cells have mysterious circles of DNA. Here’s my
@nytimes
story on their emergence—and how they may accelerate cancer.
14
212
467
1
11
72
Congratulations to Ava Carter, super star genome scientist and stem cell biologist!
Ava Carter, 27 on the
#ForbesUnder30
2019 Science list via
@forbes
0
9
38
Joan Steitz
@SteitzLab
pioneer of noncoding RNAs and a personal hero.
#shedidthat
#11February
@MolecularCell
1
9
37
@the_asci
I am grateful and humbled by this honor, especially for mentorship. Thank you to my own mentors, collaborators, and all the trainees who lifted me up.
4
0
35
@Noncodarnia
I am grateful to all my lab members and mentors. Delighted to represent and share the honor with all of you!! 🙏
1
0
25
@JasonSynaptic
@CedricFeschotte
@KeystoneSymp
Thanks! Our provocative idea: X inactivation by viral mimicry. Delighted to preprint: Spen links RNA-mediated endogenous retrovirus silencing and X chromosome inactivation
3
15
26
Beautiful new paper by
@raflynn5
on RNA virus biology, combining ChIRP-MS and CRISPR screens. Congratulations!
Excited to share my first paper with
@CarolynBertozzi
. A great collaboration with Jan Carette,
@KarimMajzoub
, and
@yaw_shin
using RNA-centric tools to learn about new ways flaviviruses hijack ER-resident RBPs
8
30
119
1
5
23
Delighted to announce
#opencrispr
challenge at
#eternagame
. Online game to improve gene editing.
0
17
22
Very cool story!
Histone H4 lysine 16 acetylation continues to surprise us. Do you want to know how mothers instruct their kids' gene expression? Check out our latest work. Great collaboration with
@Iovino_Lab
and
@leonidmirny
5
101
325
0
2
19
This work expands the world of circRNA encoded proteins. Many thanks to
@HHMINEWS
,
@StanfordMed
1
1
17
Well done Rob and team!
NEW PREPRINT:
@ENCODE_NIH
supported "Transcriptome-Wide Combinatorial RNA Structure Probing in Living Cells"
Great collaboration with
@UCIrvine
colleagues
@calizavi
and
@YongshengShi
. Help from the lovely and talented
@raflynn5
too!
12
45
117
1
3
9
Congratulations Furqan!!
Thank you, it is a great honor from the
@RNASociety
! Especially grateful for the support from my postdoc (
@HowardYChang
) and graduate (Steven Block) mentors.
6
3
41
0
0
9
Great collaborative opportunities for Chang lab, represented by
@KatieEYost
!
Saul Priceman
@cityofhope
Cristina Puig-Saus
@UCLAJCCC
@andrewjrech
@pennmednews
@MelodySmithMD
@sloan_kettering
Stephanie Dougan
@DanaFarber
Jeffrey Ward
@SitemanCenter
@KatieEYost
@stanfordmed
3/3
1
3
9
0
0
9
Congratulations Cigall!!
Extremely honored and humbled today to be named a Blavatnik National Awards finalist among these talented peers. My laboratory members and I look forward to many productive and exciting years to come!
@BlavatnikAwards
@DanaFarberNews
@broadinstitute
@harvardmed
20
12
177
0
0
8
Amazing. Congrats to
@raflynn5
and
@CarolynBertozzi
.
Postdoc
@raflynn5
and I are keen for feedback on preprint posted
@biorxivpreprint
. He made the shocking discovery that Y RNAs + other small RNAs are modified in the secretory pathway with N-glycans. Thus begins the new world of “glycoRNA”.
21
149
381
1
0
7
Congrats Xuebing!
Incredibly honored & humbled to be named a 2020 Pew-Stewart Scholar for Cancer Research! Now hiring postdocs to work on the folding and misfolding of the cancer transcriptome thanks to
@pewtrusts
& stewart trust &
@columbiacancer
29
7
153
0
0
5
@VashKurdistani
@ScienceMagazine
@NarsisAttar
@OA_Campos
Amazing discovery. Congratulations
@VashKurdistani
!
0
0
4
@CharlesSwanton
@NatureMedicine
@TakKarasaki
@davidallanmoore
@NickyMcGranahan
@MariamJHanjani
@SwantonLab
Congrats to the TRACERx team for this remarkable set of papers!
0
0
3
Well done and well deserved!!
0
0
3
@HarmitMalik
Looking forward to seeing you at NAS! I am coming with my daughter. This list is great for me--thanks for public service:)
0
0
1