Ryan Flynn
@raflynn5
Followers
5,012
Following
620
Media
336
Statuses
6,862
RNA, Glycans, Space, Renewable Energy, and all technologies making lives more interesting. Now @BCHStemCell + @HSCRB . he/him
Boston, MA
Joined May 2009
Don't wanna be here?
Send us removal request.
Explore trending content on Musk Viewer
Pedro Sánchez
• 378290 Tweets
Wizkid
• 276358 Tweets
woozi
• 204742 Tweets
#CDTVライブライブ
• 169392 Tweets
Scotland
• 105502 Tweets
The Gift
• 72699 Tweets
Thiago Silva
• 68061 Tweets
#キュウゴー
• 58275 Tweets
#TimnasDay
• 52868 Tweets
アンセム
• 51570 Tweets
Mufasa
• 45893 Tweets
Uzbek
• 34856 Tweets
Matisse
• 32074 Tweets
#アンメット
• 20787 Tweets
Blue Ivy
• 20257 Tweets
ドラゴンボール
• 17461 Tweets
Medina
• 17186 Tweets
UNIT SONGS
• 11917 Tweets
Rock in Rio
• 11868 Tweets
Wasit
• 11838 Tweets
Qちゃん
• 11377 Tweets
鈴木千裕
• 10934 Tweets
Last Seen Profiles
Pinned Tweet
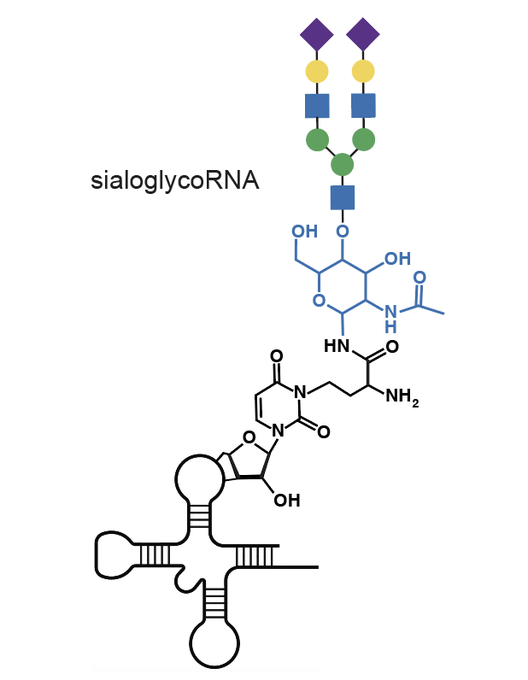
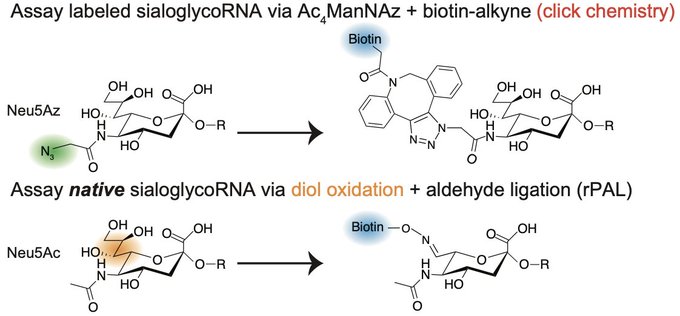
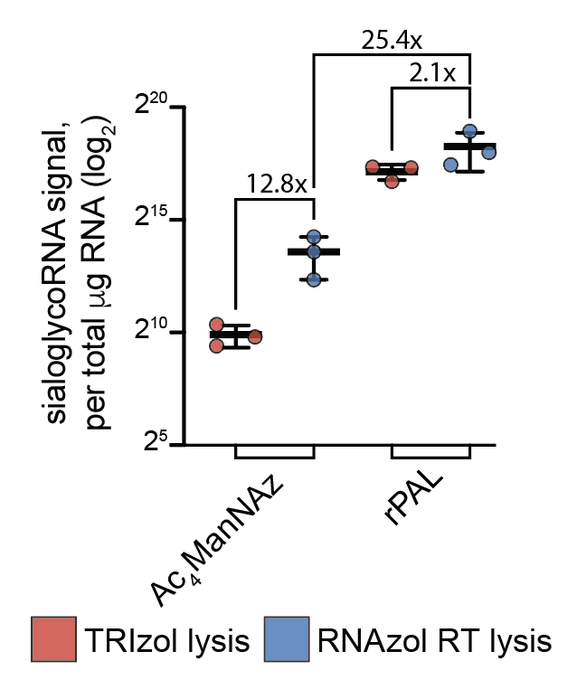
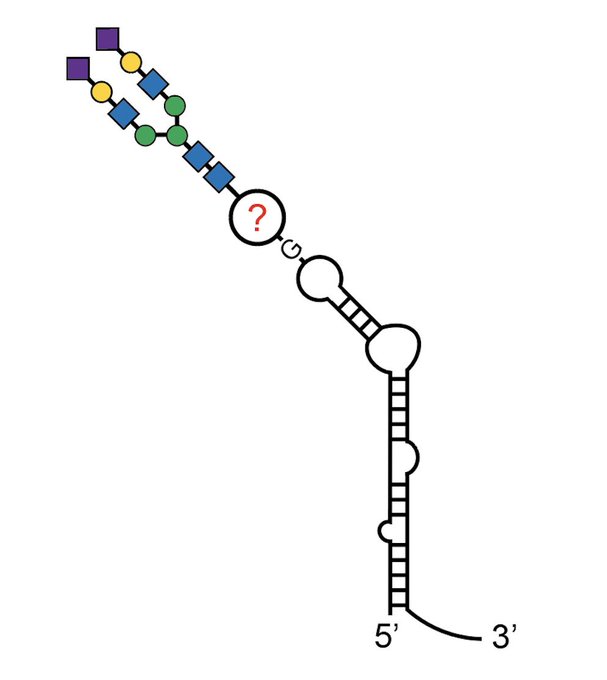
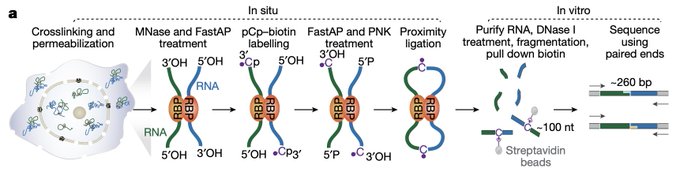
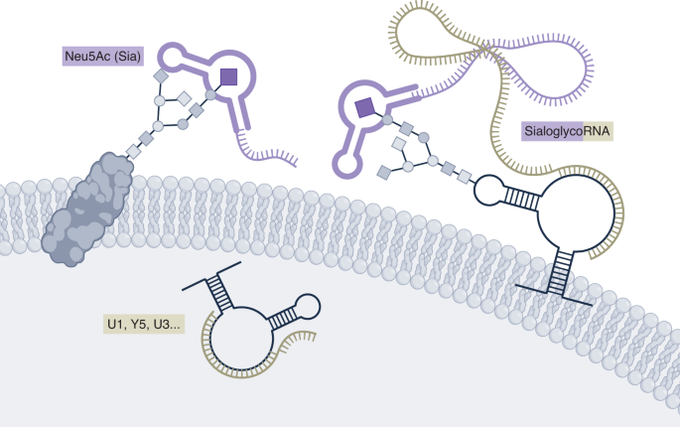
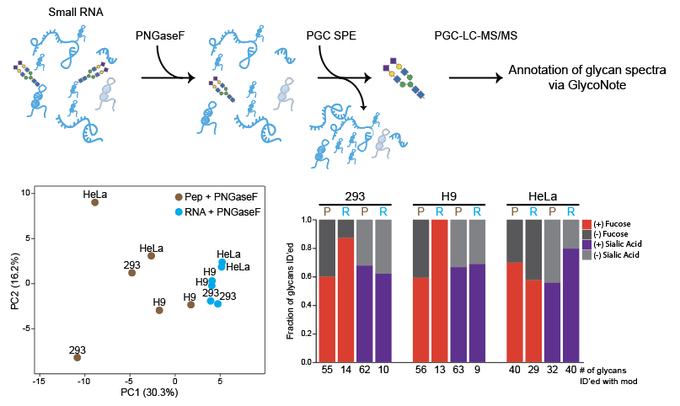
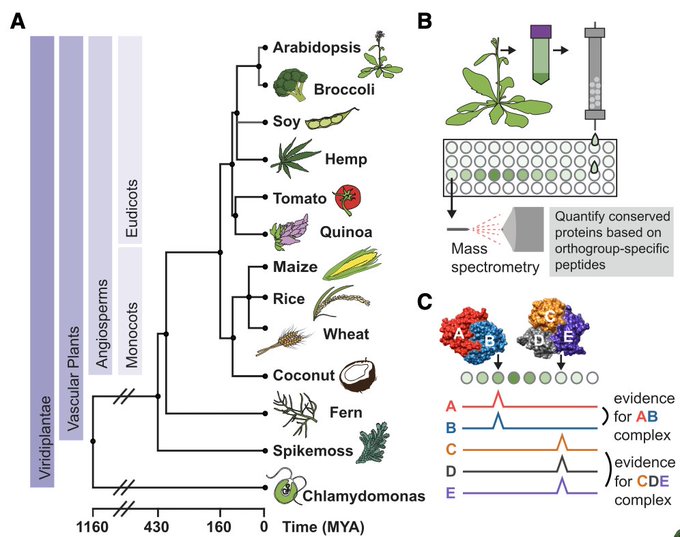
Excited to share our results that the RNA base acp3U is an attachment site for N-glycans in glycoRNA by coupling native glycoRNA labeling (rPAL) w/SWATH-MS
Led:
@axe_xie
@HelenaHemberger
Critical collab
@GarciaLabMS
@CarolynBertozzi
,
@BCHStemCell
@HSCRB
24
119
517
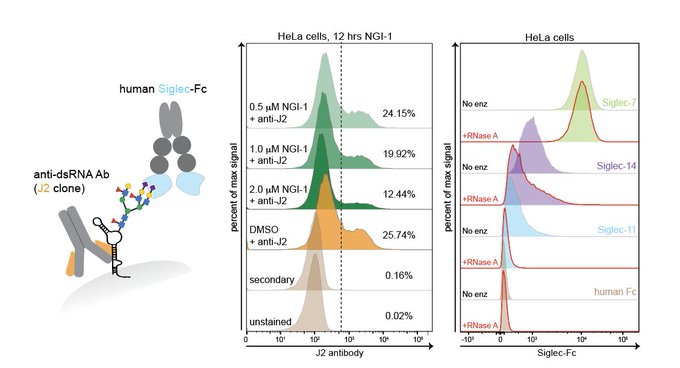
Excited to share my work w/
@CarolynBertozzi
describing the discovery of glycoRNA today
@CellCellPress
. Since our
@biorxivpreprint
we uncovered an unexpected twist: glycoRNAs are on the surface of cells+some Siglec receptors have RNA-dependent cell binding
87
715
3K
Singularly the best news - the Chemistry
@NobelPrize
to
@CarolynBertozzi
!!!! with Sharpless and Meldal
Deepest congrats Carolyn, so so well deserved and excited for you.
8
57
553
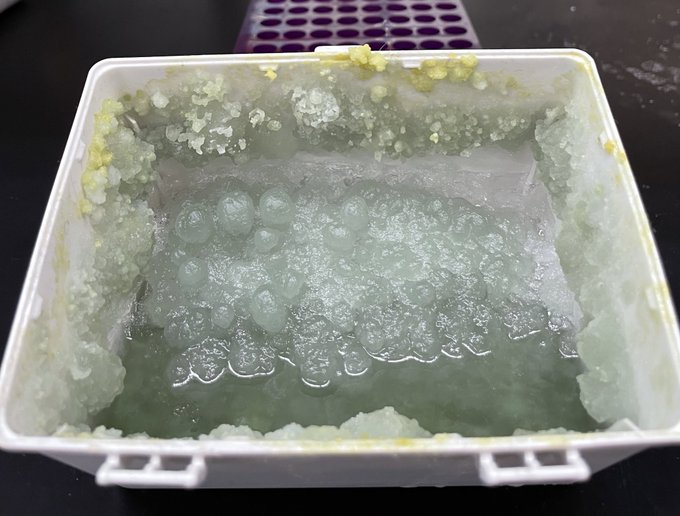
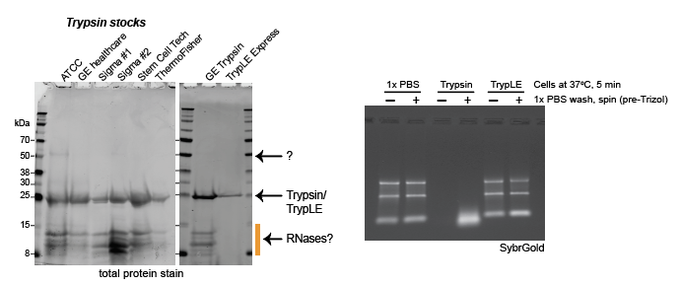
Short "know your reagents" PSA:
Cell culture trypsin has A LOT of RNase activity+is active on ice.
So if you have interest in cell surface
#glycoRNAs
D O N O T use trypsin
[promoting Thermo isn't great but] TrypLE is very clean and has no RNAse activity that I could detect
14
69
479
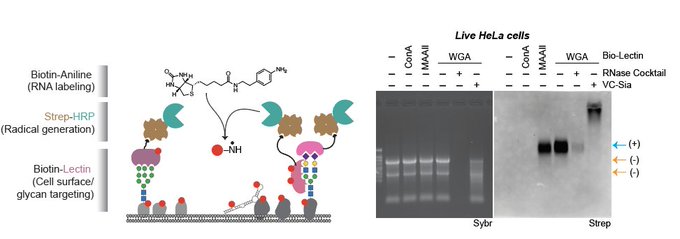
Sharing the first research effort from the Flynn Lab “Rapid and sensitive detection of native glycoRNAs”
#glycoRNA
#glycotime
#preprint
@biorxivpreprint
@BCHStemCell
@HSCRB
14
83
463
“The Ryan Flynn Lab for kids who can’t read good and want to learn to do cool science stuff good too”
@BCHStemCell
11
10
425
Coming from the RNA world, I learned about the world glycans from
@CarolynBertozzi
. We wondered, why are these two great biopolymers not talking to one another? Turns out, they are, and we’re excited to share our work on glycoRNA
16
139
402
After a decade in the bay, I’m excited to be heading back this winter to Boston to start my lab
@BostonChildrens
+
@HSCRB
! Thinking about why and how molecules like RNA and glycans work (together) to regulate human biology. []
76
19
352
Holy. Guacamole. Congrats to Aviv Regev and Genetech/Roche! Big loss for
@broadinstitute
/Boston
"will join Genentech as the new Head of gRED and will become a member of the enlarged Corporate Executive Committee as of 1 August 2020."
3
86
344
Very happy for the Flynn Lab to have support from
@NIGMS
through the R35/MIRA ESI grant! This will accelerate our basic understanding of
#glycoRNAs
, towards precisely defining their biogenesis and structure.
@BCHStemCell
@HSCRB
34
8
310
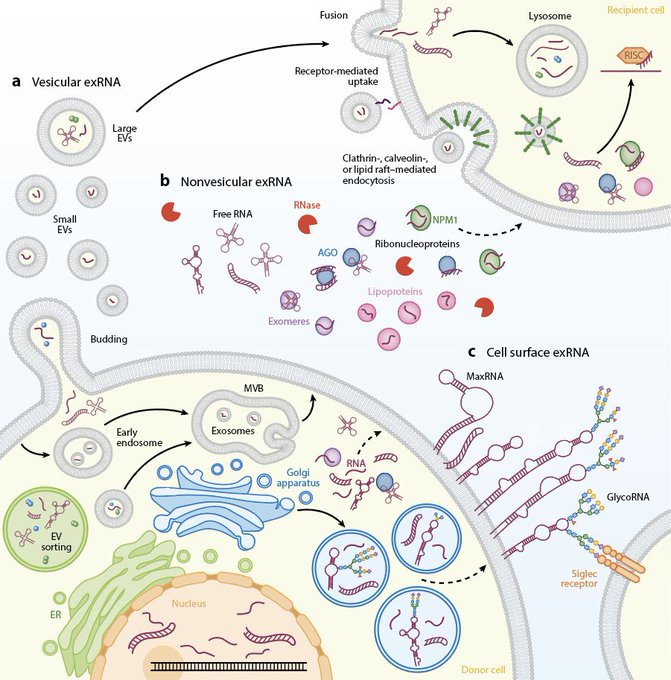
Our review is out!
“RNA Crossing Membranes: Systems and Mechanisms Contextualizing Extracellular RNA and Cell Surface GlycoRNAs”
written with Peiyuan
@PeyoChai
and Char
@catsofmisa
in the lab
@AnnualReviews
@BCHStemCell
@HSCRB
3
63
285
Excited to share work lead
@jonbperr
- we show that RBPs are common members of cell surface proteomes, form nano-scale domains with csRBP+glycoRNAs, and loss of csRNA/RBP clusters impacts entry of the TAT CPP
@BCHStemCell
@HSCRB
@biorxivpreprint
6
72
276
Very excited to have our cover art selected for this issue of
@CellCellPress
@CarolynBertozzi
Thinking about new ways to look at the cell surface.
Thanks also to
@Emily_M_Eng
for the illustration!
The new issue is out! Featuring the deep population history of northern East Asia, cardioids that recapitulate chamber-like structures, and RNAs modified with N-glycans!
#NewIssue
3
51
294
7
16
231
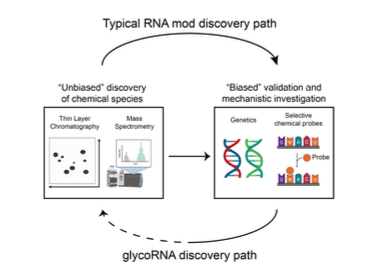
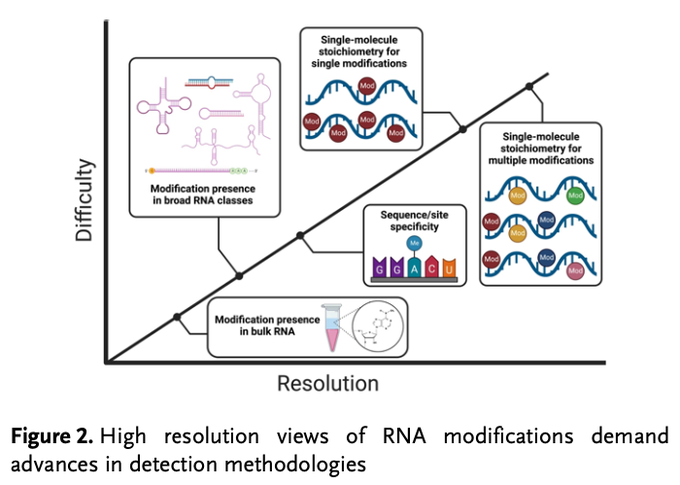
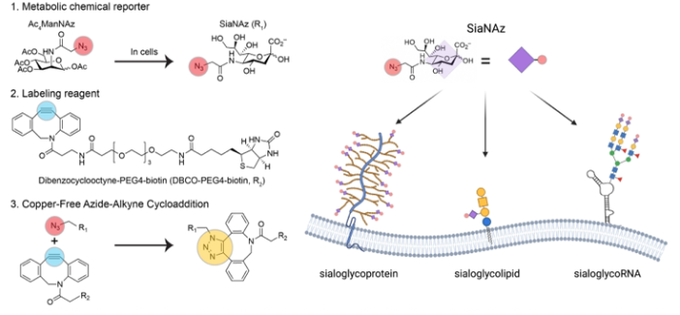
First Review Article from the Flynn Lab (!) w/
@reesecald
written in honor of
@CarolynBertozzi
winning the 2022 Wolf Prize in Chemistry! (relevant for other Chemistry prizes) We discuss RNA PTMs and how we used bioorthogonal concepts to discover
#glycoRNAs
6
42
204
We’re super excited to get this support from
@DamonRunyon
which will help us push our
#glycoRNA
work closer to clinical translation!
@raflynn5
’s research aims to define RNA as a new cell surface molecule that could have unique structures on acute myeloid leukemia (AML) cells. With this knowledge he will develop antibodies to selectively detect cancer cells and enable tumor killing.
1
0
12
21
7
191
Excited and honored to be a part of the 2022 class of Rita Allen Foundation Scholars. Their support will help us uncover more
#glycoRNA
biology!
24
15
182
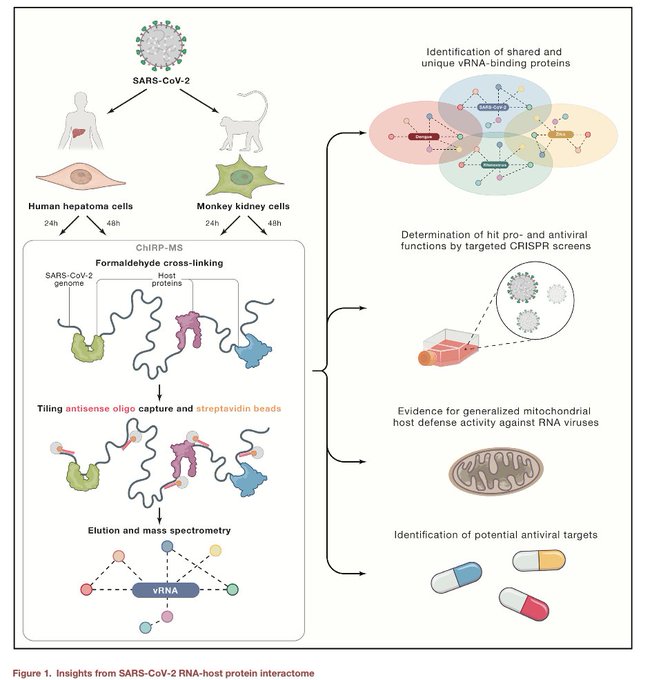
We can finally hear the ChIRPing of the SARS-RNA-interactome! Very collaborative work w/
@juliabelk
@Satpathology
@CarolynBertozzi
@WilenLab
JanCarette
@HowardYChang
others! |
@yaleibio
@Stanford_ChEMH
@StanfordMed
@BWFUND
12
57
178
"New Vistas for Cell-Surface GlycoRNAs"
Great piece from Siggy Nachtergaele and
@KrishnanYamuna
about
#glycoRNA
@NEJM
4
37
157
The Flynn Lab is looking for curious people interested in figuring out
#glycoRNA
stuff.
Skills moving solids, liquids, ions, or electrons around are all welcome.
Boston is sweet, and also part of the larger
@BCHStemCell
and
@HSCRB
communities
4
58
152
Flynn Lab is looking for postdocs!
Topics: A = RNA, B = Glycans → A, B, A+B, A*B, etc
Please email/DM with interest
#glycoRNA
#glycotime
@BCHStemCell
@HSCRB
4
46
146
Had a great time visiting
@MIT
/
@kochinstitute
to discuss our lab’s recent work on
#glycoRNA
- thanks to Phil Sharp for hosting
0
4
133
ChIRP-MS of
#SARSCoV2
+ RBP KOs +
#Mitochondria
KO screen in 7 viruses
Many RBPs and Mito factors work as host-protective during RNA virus infections!
@CellCellPress
w/
@juliabelk
@Satpathology
@CarolynBertozzi
@WilenLab
Carette
@HowardYChang
Horvath
10
29
123
Very excited to have been selected as a 2020 CAMS awardee! Thankful to
@BWFUND
forward getting excited about
#glycoRNA
#glycotime
and what is to come. Shout out to
@CarolynBertozzi
for supporting and cultivating this so far!
28
2
121
Excited to share my first paper with
@CarolynBertozzi
. A great collaboration with Jan Carette,
@KarimMajzoub
, and
@yaw_shin
using RNA-centric tools to learn about new ways flaviviruses hijack ER-resident RBPs
8
30
119
Excited for HiChIRP to be out! It's the HiChIP, for RNA: and congrats to
@MaxMumbach
@JeffreyGranja
@HowardYChang
@WJGreenleaf
1
51
118
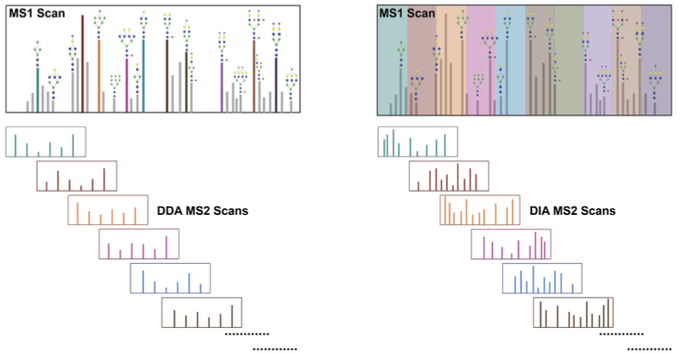
GlycanDIA is out!
Congrats to
@axe_xie
@GarciaLabMS
@CarlitoLebrilla
GlycanDIA can ID+Quant glycans with high sensitivity/accuracy and distinguish composition + isomers of glycans.
This could enable new access to glycomics for low abundance samples
4
20
116
Happy to welcome
@lauren_kageler
as a new PhD student in the Flynn Lab! Lauren is in the Chemical Biology Program
@chemical_phd
@harvardmed
and will be thinking with us all about
#glycoRNA
biology.
2
3
108
Excited for Jon Perr’s (
@jonbperr
) first day as the *first* grad student in the Flynn Lab.
#glycoRNA
MCO/
@MCB_Harvard
@HSCRB
@BCHStemCell
1
2
103
We’re excited to welcome
@ChehadaDina
to the Flynn Lab as our newest graduate student! She’s in the
@chemical_phd
and will be developing new tools to understand
#glycoRNAs
in mammalian systems
@BCHStemCell
@HSCRB
3
4
100
Very excited to officially welcome Dr. Benson George
@BensonMGeorge
to the Flynn Lab. Having completed the beginning of his Heme/Onc Fellowship
@DanaFarber
he will be helping us better connect glycoRNAs to outstanding clinical problems in blood cancer.
@BCHStemCell
@HSCRB
5
4
99
Fun to see our News & Views out written with
@pphristov
about the ARPLA method developed by
@yuan_ma_
@weijieguoguo
and
@LuLabUT
to visualize
#glycoRNAs
on single cells
1
16
98
Welcoming the newest post-doc to the Flynn Lab, Peiyuan Chai
@PeyoChai
-- lots of
#glycoRNA
and
#glycotime
work coming!
1
4
97
Very grateful to Rick and
@sontagfdn
for this award and enabling the Flynn Lab to explore
#glycoRNA
mechanisms in the context of brain tumors.
Exciting things to come and open positions in the lab
@HSCRB
@BCHStemCell
12
11
97
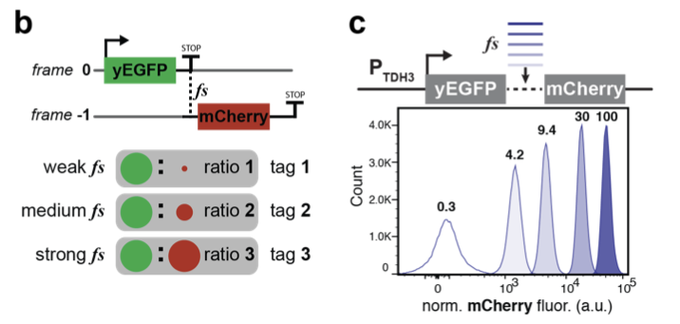
“genetically encoded fluorescent barcodes *FRAME-tags* that break a scalability barrier by encoding barcode identity as unique FP expression ratios” super cool idea and very awesome looking flow plots from Anzalone, Jimenez, and V. Cornish
@ChemColumbia
0
11
93
Very excited that our very own Reese Caldwell
@reesecald
was selected for the
@NSF
GRFP! Reese is wrapping up an incredible undergrad thesis effort
@HSCRB
. It’s a well deserved honor and we’re excited to see what he does next! (picture of him smiling, likely thinking about RNA)
1
0
91
We’re excited to welcome Jennifer Porat to the Flynn Lab as our newest postdoc! Coming from the Bayfield lab where she carefully uncovered new functional roles of small RNAs, here she will help us better understand mechanistic aspects of
#glycoRNAs
@BCHStemCell
@HSCRB
2
0
87
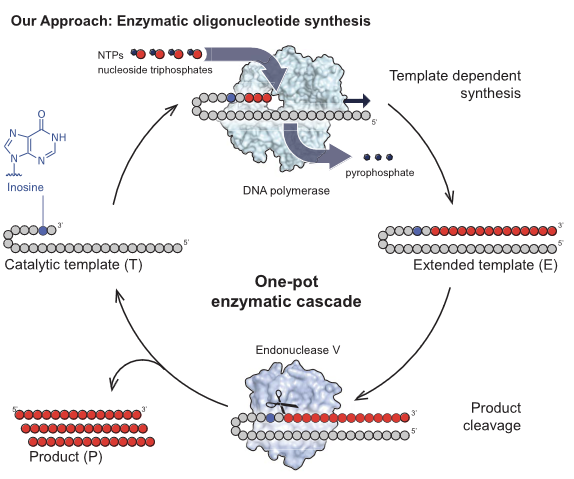
Really cool work from
@LovelockLab
"An enzyme cascade enables production oftherapeutic oligonucleotides in a single operation" demonstrating useful scale and purity
@ScienceMagazine
2
6
81
DNA-PK functionally assembles on nucleolar RNA! Happy to share our very nice collaboration between
@CaloLabMIT
and Zha lab
@ColumbiaMed
-- "DNA-PKcs has KU-dependent function in rRNA processing and haematopoiesis"
3
12
78
Excited to have Petar
@pphristov
join the Flynn Lab (
@BCHStemCell
@HSCRB
) as a new PhD student! Petar is in the Department of Chemistry and Chemical Biology
@HarvardCCB
and will be working on methods to better understand
#glycoRNA
biology
1
2
70
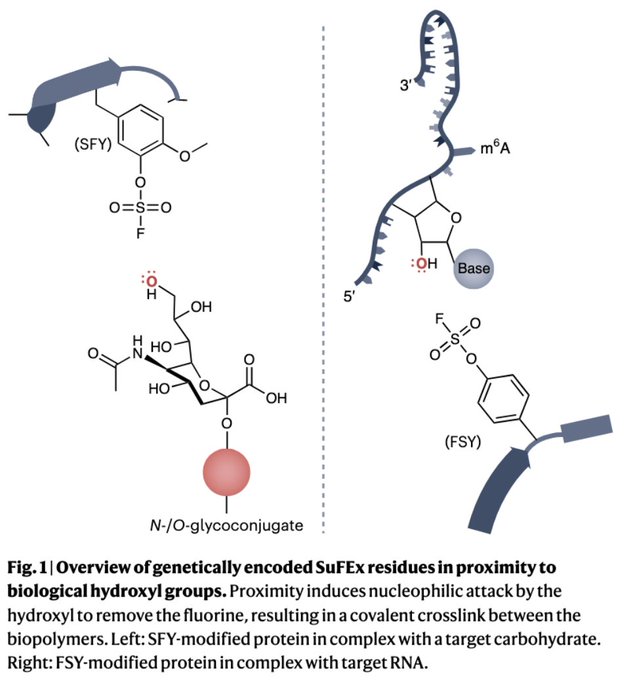
Happy to see our News & Views
@NatureChemistry
come out, written with postdoc Chris Watkins
@BCHStemCell
@HSCRB
about 2 new papers from Lei Wang’s
@UCSF
on SuFEx-based chemical Xlinking methods – useful for RNA-protein and glycan-protein interactions!
1
10
69
Have you ever wanted to decide the “group” vs “lab” naming? -- come join me in 2021
@BCHStemCell
and
@HSCRB
and help me decide! On the side we can think about glycoRNAs and the broader interface of RNA biology and glycobiology too…
#glycotime
#glycoRNA
1
19
65
Quick thanks / shout out to
@disney_lab
for writing a great Preview on
#glycoRNA
@CellCellPress
@CarolynBertozzi
1
7
63
Had a really great time visiting
@OdedRechavi
@TelAvivUni
. Fun to share new things in
#glycoRNA
methods and to talk about the cool science going on here!
2
1
60
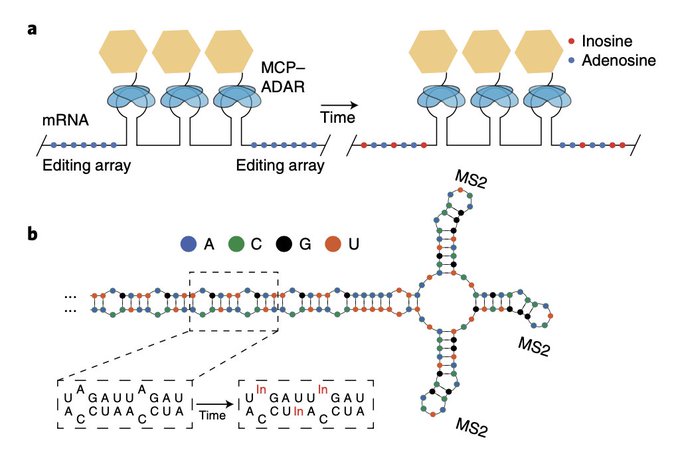
Using ADAR to repeatedly edit RNAs, allowing for the tracking of their "age" from Fei Chen
@eboyden3
@HSCRB
@broadinstitute
0
8
56
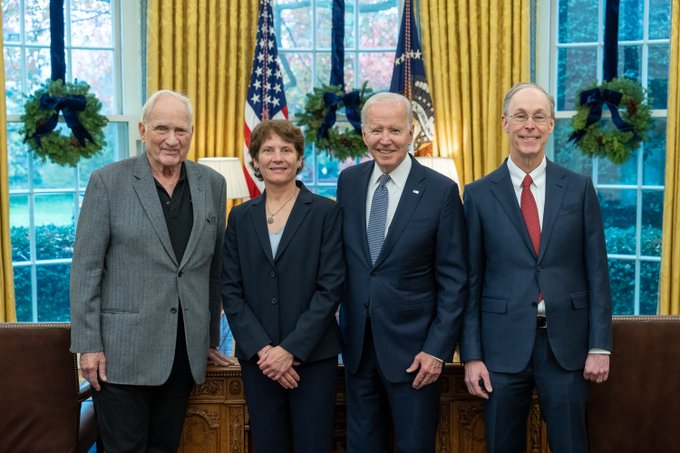
Thanks Carolyn and also for all the help with this 5+ year effort!!
During his postdoc
@raflynn5
discovered the existence of
#glycoRNA
on cell surfaces. Now
@BostonChildrens
@Harvard
he has determined the structure of one of the attachment sites: the modified base acp3U, in collab with
@GarciaLabMS
!
1
37
230
0
0
53
Check out Andrew’s work on the HUSH complex now out in
@MolecularCell
“Co-transcriptional genome surveillance by HUSH is coupled to termination machinery” Awesome thesis work in Joanna Wysocka’s lab
@StanfordMed
0
9
52
@eyal_maori
@qichen_lab
@CarolynBertozzi
@CellCellPress
@biorxivpreprint
We're thinking a lot about this now but you could imagine scenarios where glycoRNAs may play roles in parallel to other cell surface biomolecules like glycoproteins/lipids but with different dynamics, stabilities, or configurations
5
6
51
Thanks to Pamela Stanley for the invitation to share some of our new
#glycoRNA
work to the Cell Biology department. It was also great to hear some of the other exciting work going on
@EinsteinMed
1
1
49
Great to have Char
@catsofmisa
on the Flynn Lab team, working with us to better understand why cells make
#glycoRNA
!
1
0
43
"SARS-CoV-2 RNA: Exclusive friends and common foes"
@CellCellPress
Thank you to
@emmavleblanc
and
@colpittsc
for the very nice preview that includes discussion of our (
@Satpathology
@juliabelk
@WilenLab
) SARS-CoV-2 ChIRP-MS work!
1
13
43
@davidrliu
@QIAGEN
@QIAGENscience
usually...
RNA lysis buffer: 6M guanidine thiocyanate, 4% sarcosyl, and 4% Titron X-100 +/- 100mM β-mercaptoethanol
RNA wash buffer 1: 4M guanidine hydrochloride in 80% EtOH
RNA wash buffer 2: 80% EtOH
h/t to
@HentzeTeam
Asencio et al 2018
2
14
42
This is sick
0
2
41
Excited to have Chris Watkins
@CwatkinsChem
join us as a new postdoc! He’s coming from Tao Pan’s group where he developed molecular tools to better understand tRNA biology.
@BCHStemCell
@HSCRB
1
1
41
And now, we’re back to work [and lots to do]
@BCHStemCell
and
@HSCRB
exploring new mechanisms, tools, and functions of glycoRNAs
4
2
39
Check out this very cool new method to directly image *specific* native glycoRNA transcripts!
Excited about GlycoRNA which was discovered by
@CarolynBertozzi
and
@raflynn5
? Come and check our most recent paper on the development of glycoRNA in situ imaging tool, ARPLA, leverages the power of aptamer and ISH.
@LuLabUT
@NatureBiotech
4
12
52
1
4
40
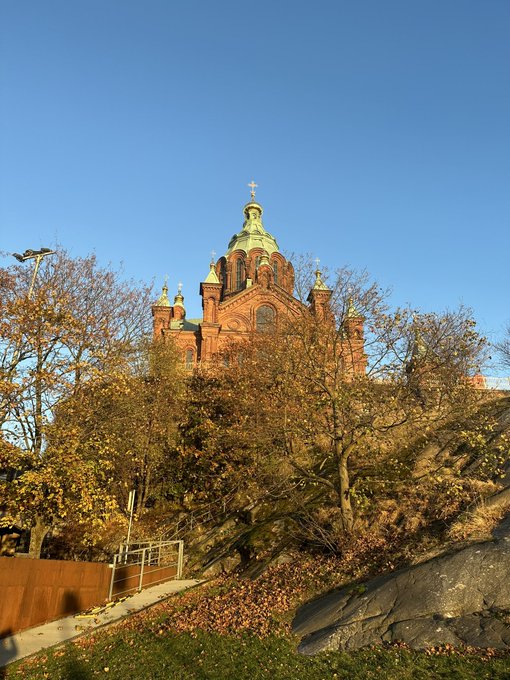
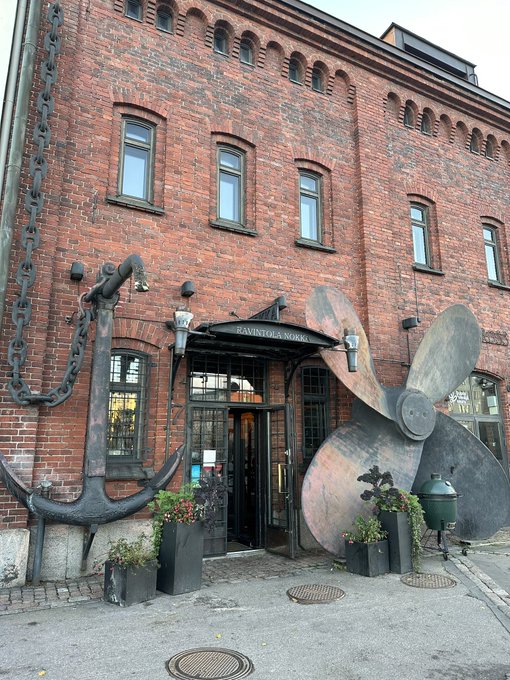
Thanks to
@v_paavilainen
for a great visit to
@LifeSciHelsinki
@HelsinkiUniMed
@HiLIFE_helsinki
nice to meet many of the labs here and see a bit of the city!
3
0
37
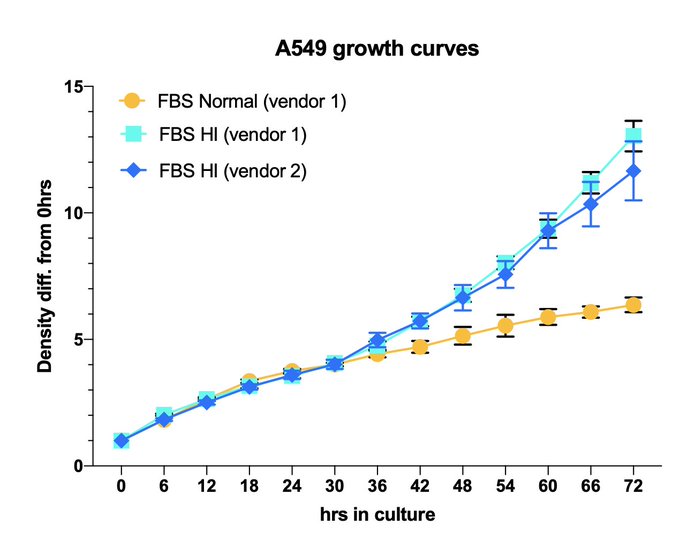
Ended up testing cell growth w/DMEM+reg. vs HI FBS. Was not expecting such an obvious difference… Can’t really say that 1 is “better”. Just highlights the need to move towards expanding the HPLM idea from
@JasonRCantor
and
@DMSabatini
but for many more cell types..
3
2
37
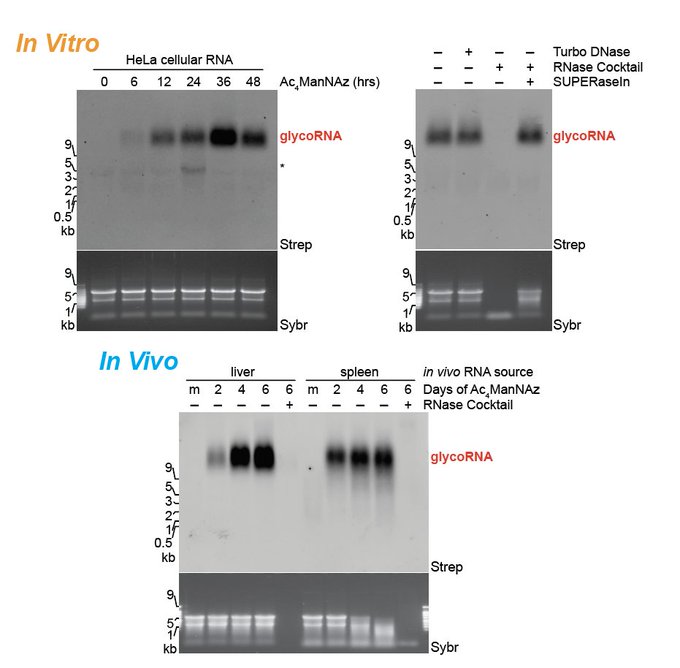
Enabling this was
#bioorthogonal
chem from
@CarolynBertozzi
: ManNAz could be incorporated into RNA from mammalian cells in vitro and in vivo. The RNAs are a collection of sno, sn, t, and Y RNAs that are similarly selected across two cell types
3
4
34
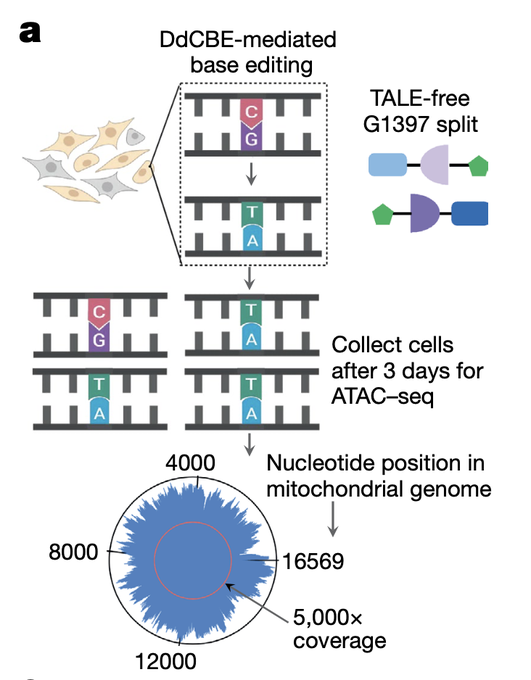
As many have seen, really cool work from the
@davidrliu
and Mougous' labs on mtDNA editing.
I particularly like how they casually use the background of ATACseq for essentially targeted seq of the mtDNA in Fig. 5a
@nature
1
5
34
Celebrating Helena’s
@HelenaHemberger
very important contributions to the Flynn lab before she heads off to BU for grad school!
1
2
34
First edition of the
#SmartinelliChallange
with Jon
@jonbperr
officially becoming a PhD candidate!
The challenge begins with a glass of Smartinelli’s and ends the same way, but one is a domain expert (and + a PhD) at the end of the challenge
@BCHStemCell
@HSCRB
@MCB_Harvard
0
2
32
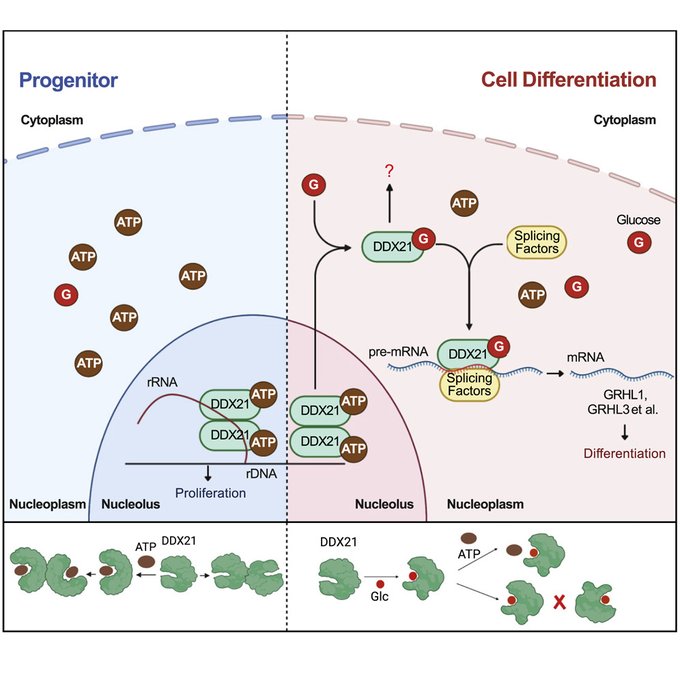
Nucleolar DDX21 sensitive to [glucose], where high [Glc] results in splitting of DDX21 dimers, nucleolar exit, leading to modulation of splicing
By Weili + Douglas in the Khavari lab
@StanfordMed
happy to have made a small contribution
@CellCellPress
0
4
31
Wish we could do hematopoietic stem cells transplants without ANY chemo/radiation? Now we have a path towards it courtesy of Benson George and Irv Weissman
Enabling MHC-mismatched hematopoietic stem cell transplants and organ graft tolerance without chemotherapy or radiation
#bioRxiv
0
10
17
1
11
31
Cool
@CellCellPress
Resource paper from
@edward_marcotte
New meaning to "deep proteomics profiling" -- 14,520,970 interpretable peptide mass spectra from 2,111 individual fractions
1
9
30
Congrats to my PhD mentor
@HowardYChang
and the other 18 excellent PIs for becoming HHMI Investigators
0
2
29
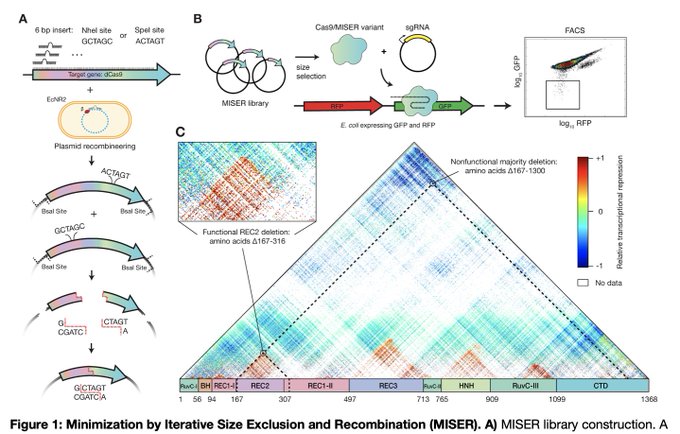
Neat idea to systematically explore the ability to lose all parts of a protein, but retain activity.
@realarikshams
+ Sean Higgins in
@SavageCatsOnly
and
@doudna_lab
1
8
28