Ellen Zhong
@ZhongingAlong
Followers
7,243
Following
908
Media
75
Statuses
703
Assistant Professor @PrincetonCS . #proteins247 #ml #cryoem #cryodrgn ❄️🐉 #compbio #ai4science Prev: @MIT @DeepMind @DEShawResearch
Joined May 2011
Don't wanna be here?
Send us removal request.
Explore trending content on Musk Viewer
No Riley
• 199816 Tweets
Ayuso
• 193950 Tweets
Hawk Tuah
• 99003 Tweets
Netherlands
• 95833 Tweets
Sean
• 94345 Tweets
#NEDFRA
• 90631 Tweets
Austria
• 82541 Tweets
#LondonTSTheErastour
• 74636 Tweets
Francia
• 68228 Tweets
Olise
• 61099 Tweets
Kante
• 54382 Tweets
Bayern
• 52683 Tweets
Griezmann
• 42139 Tweets
Rabiot
• 39610 Tweets
فرنسا
• 33132 Tweets
Thuram
• 32408 Tweets
オランダ
• 30944 Tweets
Doña Segunda
• 30343 Tweets
Holanda
• 28376 Tweets
THE BLACK DOG
• 27763 Tweets
Dembele
• 26332 Tweets
Deschamps
• 25722 Tweets
HITS DIFFERENT
• 21244 Tweets
Maroon
• 20987 Tweets
كانتي
• 19166 Tweets
DIAN
• 16996 Tweets
Barcola
• 16873 Tweets
COME BACK BE HERE
• 14483 Tweets
Depay
• 14279 Tweets
Xavi Simons
• 11751 Tweets
Dumfries
• 11034 Tweets
Maignan
• 10471 Tweets
Last Seen Profiles
Pinned Tweet
We present a new method, DRGN-AI 🐉🤖, for fast ab initio reconstruction of modern (i.e. messy and large) cryo-EM datasets! The culmination of an amazing collaboration with superstar
@axlevy0
.
We operate on unfiltered versions of EMPIAR datasets and discover new states. See 🧵!
We present DRGN-AI for fast, ab initio cryo-EM reconstruction!
* learns a neural field from unposed images,
* designed for single-shot reconstruction of unfiltered datasets,
* finds new states missed by prior approaches!
Teamwork led by
@ZhongingAlong
1/
2
48
212
2
22
168
I am thrilled to share that I will be starting as an Asst Professor of Computer Science at
@Princeton
this summer! Really excited for the future of machine learning in structural biology and honored to have the opportunity to lead a group in this exciting area!
#proteins247
1/
130
68
2K
I am recruiting PhD students
@PrincetonCS
interested in ML applications in bio!
Our group aims to define new problems for AI in molecular & structural bio. We work at the intersection of many areas incl 3D vision & generative modeling for proteins. Plus we make useful tools!❄️🐉
16
228
1K
What do dynamic protein complexes look like *inside the cell*? Excited to share our work, led by superstar
@ramyarangan
, on cryoDRGN-ET, a new approach in the ❄️🐉 software for reconstructing and visualizing structures from in situ cryo-ET data. 1/
6
163
677
We present a new algorithm, cryoFIRE ❄️🔥, for fast ab initio heterogeneous reconstruction, appearing at
@NeurIPSConf
this Dec!
Really proud to share our latest work, led by the amazing
@axlevy0
, and my first paper
@PrincetonCS
#EZlaboratory
Paper: 1/
8
124
588
CryoDRGN2 ❄️🐉 is now out at
@ICCV_2021
! We present an algorithm for *ab initio* reconstruction of heterogeneous protein structures. We focus on techniques for fast + accurate pose inference to achieve state-of-the-art reconstructions on all of our fav, real cryo-EM datasets 1/
4
105
474
I just officially started my appointment
@PrincetonCS
and couldn't be more excited! Current status: wandering around Princeton figuring out how to buy GPUs
#newPI
16
2
416
Our cryoDRGN ❄️🐉 paper is out! A deep generative model for reconstruction and analysis of protein structures from cryo-EM images, including discovery of new conformations and visualizing continuous dynamics. 1/
CryoDRGN: a deep neural network based approach for heterogeneous single particle cryo-EM reconstruction
@jhdavislab
@ZhongingAlong
2
40
153
6
84
368
The Machine Learning in Structural Biology workshop
@workshopmlsb
is on again
@NeurIPSConf
, in-person/hybrid in New Orleans this Dec! Stay tuned for our call for papers 📜
More info here:
Sign up for our mailing list:
2
69
283
A monumental work by John Ingraham and team
@generate_biomed
on Chroma✨ for protein design. Congrats!!! 🔥🎉🔥🎉
So excited to see it out, fully open source and with experimentally validated xtal structures! Perhaps cryo-EM of the giant complexes next?
2
29
259
Amplifying this GitHub thread on cryoDRGN ❄️🐉integration with
@UCSFChimeraX
! Please comment if you have any feedback/desiderata or just a 👍/🎉 would be appreciated
4
40
252
Above and beyond seminar trip to the Institute of Protein Design
@UWproteindesign
and seeing the Baker lab in action. Thanks for having me!
3
8
233
Pleased to announce the speaker lineup for our
@NeurIPSConf
workshop, ML in Structural Biology, featuring keynotes from John Jumper
@DeepMind
and Jane Richardson and invited talks by
@mmbronstein
,
@BarzilayRegina
,
@LucyColwell37
@CecClementi
,
@keating_lab
,
@Dereklowe
3
33
207
In a fantastic collaboration with
@karsten_kreis
,
@timudk
, and Zihao Li, we extend cryoDRGN ❄️🐉 for generative sampling of cryo-EM structures via latent diffusion models. We'll be presenting this work
@workshopmlsb
@NeurIPSConf
Sat, 9am!
#EZlab
Paper: 1/
6
41
191
The Machine Learning in Structural Biology workshop
@workshopmlsb
at
@NeurIPSConf
is happening again! Our website is still out of date () but stay tuned for our speaker lineup and call for papers for 2021!
Preregister here:
1
41
165
Release the cryoDRGN ❄️🐉 (on biorxiv)! Instead of static protein structures, we develop a neural network architecture to reconstruct diverse conformations and molecular motions from real cryo-EM image data. Joint work with Tristan Bepler, Bonnie Berger, and
@jhdavislab
.
3
40
143
This is incredible work on protein generative modeling by John Ingraham and team
@generate_biomed
. Innovative ML (novel diffusion, sub-quadratic scaling arch) and insanely impressive results of diverse, designable proteins (incl. 96,000 res symmetric complexes)!
#MLSB
is on 🔥
Today we introduced Chroma, a generative model that creates new proteins & protein complexes given geometric & functional constraints. It learns to transform unstructured, random 3D shapes into
#protein
molecules, which can have tens of thousands of atoms.
13
230
1K
2
18
126
CryoDRGN ❄️🐉 in the wild! A trajectory of 120 reconstructed structures sampled from the latent space of the generative model helps to understand the conformational dynamics of this megaenzyme ✨
0
22
115
I am so excited to see this out!! It was an amazing experience working with the team on new directions in the midst of the
#AF2
release, and very thankful to be able to contribute. Fun fact, I called my model denoisaur 🦖🦕😂
PS Part of my decision to join
@PrincetonCS
was the
Announcing AlphaFold 3: our state-of-the-art AI model for predicting the structure and interactions of all life’s molecules. 🧬
Here’s how we built it with
@IsomorphicLabs
and what it means for biology. 🧵
215
2K
7K
3
10
114
I'm prepping a new talk on machine learning in biology for a rising stars symposium at
@IMPvienna
! Excited for fresh slides and story on how ML e.g.
#cryoDRGN
,
#alphafold
,
#LLMs
can impact biological discovery.
7
6
115
Happy Monday! AlphaFold for protein complexes to be released soon! 🥳
1
21
113
Insane progress these days in ML for structural biology 🔥🔥🔥 The team at Meta AI just released an atlas of 617M+ protein structure predictions that include metagenomic proteins with no/shallow MSAs
0
28
109
Excited to give two same-same-but-different talks this week: a structural biology-oriented ❄️🐉 talk today at 11am ET hosted by
@structbiolbxl
and an ML-oriented talk tmrw (Fri) at 10am ET at the
@iclr_conf
workshop on Deep Generative Models for Highly Structured Data
2
8
106
A profile of our ❄️🐉 collaboration between
@MITBiology
@MIT_CSAIL
@MITMath
Solve is a little strong 😅 but proud to contribute to a truly interdisciplinary problem with
@tbepler1
@jhdavislab
and
@lab_berger
Grad student
@ZhongingAlong
reached across departmental lines to solve a long-standing problem in electron microscopy. Co-advised by
@jhdavislab
and
@lab_berger
, she helped devise software to reconstruct molecules in motion. Article by
@saimamaysidik
:
0
4
34
2
12
96
A nice paper from
@chloehsu0
,
@adamlerer
,
@alexrives
, and others from
@MetaAI
on learning an inverse folding model (predicting sequence from structure), supervised on millions of
#AlphaFold
predictions. It's an exciting time for protein design!
0
17
94
Very excited to co-organize the 4th Machine Learning for Structural Biology workshop
@workshopmlsb
@NeurIPSConf
again this year, happening in New Orleans this Dec!
Stay tuned for our call for papers 📜
Website:
The Machine Learning in Structural Biology workshop will be back at
#NeurIPS
once more! MLSB will be an in-person workshop held in New Orleans in December.
Website & Speaker Lineup:
Mailing List:
4
21
104
1
11
90
400k+ more
#Alphafold
predictions released... including T-Rex and woolly mammoth haha
🎉New
#AlphaFold
data! With
@DeepMind
, we’ve more than doubled the size of the database & added predictions for most of the manually-curated
@uniprot
entries in UniProtKB/SwissProt.
That's >400,000 new protein structure predictions for you to explore!
7
188
585
4
5
87
Alphafold will be open sourced 🔥💯❤️
Structure prediction ✅
#proteins247
Brief update on some exciting progress on
#AlphaFold
! We’ve been heads down working flat out on our full methods paper (currently under review) with accompanying open source code and on providing broad free access to AlphaFold for the scientific community. More very soon!
70
700
4K
2
7
84
Very inspiring to be in Stockholm and honored to open
#CryoNet
2023 with our latest work on cryoDRGN(-ET) ❄️🐉
0
6
82
I'm in such awe of the
#AlphaFold
team and feeling so privileged to be able to work with them this summer at
@DeepMind
. Very optimistic about the future scientific discoveries from this transformative technology 🤩
1
3
79
Had a fun convo on AI and drug discovery with
@kchonyc
@kpowgerade
and
@RichBonneauNYC
at the Atlantic Festival today!
Sponsor Content:
@zhongingalong
at
@PrincetonCS
shares what good data entails to build foundational models. This session is produced by
@genentech
.
#TAF23
0
1
8
1
7
77
Just read this incredible work on DNA LLMs by
@exnx
,
@pdhsu
,
@BrianHie
et al!
So many fun things like fine-tuning on CRISPR seqs to generate protein-RNA complexes, zero-shot gene essentiality prediction, genome-scale DNA generation given bacterial species prompts, oh my! 🔥🚀👏
2
6
77
It's the dawn of a new era of biology 🥹
From ~100k unique protein structures in the PDB about a year ago to 200M+
#AlphaFold
structures today
Today in partnership with
@emblebi
, we’re releasing predicted structures for nearly all catalogued proteins known to science, which will expand the
#AlphaFold
database by over 200x - from nearly 1 million to 200+ million structures: 1/
107
2K
6K
1
8
75
Our data from the cryoDRGN-ET paper is now publicly released on EMPIAR!
Pioneering efforts of
#teamtomo
’s
@AbhayKot
from
@thermofisher
led to this spectacular lamella from S. cerevisiae, the entry includes all metadata and annotation of ribosomes! Published
@biorxivpreprint
().
#empiar
Data:
0
10
41
0
10
74
MLSB is tmrw! Excited to hear from John Jumper
@DeepMind
- his
@workshopmlsb
talk goes more in depth into the ML behind
#AlphaFold
than previous lectures; and absolutely honored to hear from Jane Richardson, pioneer of ribbon drawings (see her bio at )
Featuring keynotes:
John Jumper: “Highly accurate protein structure prediction with AlphaFold”
Jane Richardson: “The early days of structural biology before ML”
And invited talks from
@BarzilayRegina
@mmbronstein
@LucyColwell37
@CecClementi
@keating_lab
@Dereklowe
w live Q&A! 2/
1
2
12
1
7
73
Diffusion generative models now applied to 3D protein structure in the IPA module of Alphafold2 by
@namrata_anand2
:) 🔥 Interesting downstream applications and I'm really optimistic on these types of models to extend to real world tasks in structural bio
0
9
70
Congratulations
@BodrugTanya
! Check out our work on reconstructing time-resolved conformational landscapes of the APC/C complex with cryoDRGN ❄️🐉. Joint with Brown lab
@UNC_PHCO
and
@HaselbachLab
, now published in
@NatureSMB
Exited to share my thesis project! Huge thanks to co-mentors Nick Brown at
@UNC_PHCO
and
@HaselbachLab
. Also couldn’t have made it happen without major input and contributions from
@ZhongingAlong
(cryodrgn❄️🐉)
@Hahn_Lab
(microscopy)
@WeiZhangTO
(ubiquitin variants) and others 🙏
5
9
40
0
8
69
Not static bricks indeed! 😍 A VAE for direct cryo-EM reconstruction of atomic protein conformation *and image pose* evaluated on MD simulation data.
Proteins are not static bricks! Feasibility study to infer a continuous distribution of all states using an end-to-end model from Cryo-EM images to atom coordinates: .
@danrsm
,
@GarneloMarta
,
@MichaelZielins
,
@JonasAAdler
,
@arkitus
,
@CarlDoersch
,
@pushmeet
8
168
646
2
9
66
"cryodrgn gave me by far the best classification results so far (compared to a lot of RelionClass3D and CS 3DVA or the new 3Dclassification)"
Nice to wake up to emails like this 😊
(Obligatory caveat emptor - your mileage may vary)
#deeplearning
1
1
65
That's a wrap for my first
@CVPR
! Had an awesome time at the neural fields tutorial, poster sessions, speed mentoring, and even a swamp tour! Lots of exciting parallels between cryo-EM and the vision community to explore
1
2
65
Just landed in New Orleans for
@NeurIPSConf
! Please reach out if you want to meet and chat about anything ML in structural biology, grad school/ joining my new group or more! Find me at our poster on ❄️🐉🔥 Tues 4-6p and at
@workshopmlsb
this Sat
We present a new algorithm, cryoFIRE ❄️🔥, for fast ab initio heterogeneous reconstruction, appearing at
@NeurIPSConf
this Dec!
Really proud to share our latest work, led by the amazing
@axlevy0
, and my first paper
@PrincetonCS
#EZlaboratory
Paper: 1/
8
124
588
1
2
61
Nice commentary on the synergies between
#AlphaFold
/
#rosettafold
structure prediction and cryoEM/ET structure determination
@cryoem_UBC
.
My additional 2 cents is that there are exciting possibilities specifically for new ML methods at this intersection. V exciting times!
Sriram Subramaniam and Gerard Kleywegt discuss how structure prediction and cryo-EM/cryo-ET will synergize to tackle challenging biological questions regarding conformational dynamics and in situ structure analysis.
@cryoem_UBC
@UBC
@emblebi
@PDBeudrope
0
23
76
1
6
57
I had a great conversation with
@mcianfro
,
@kellogg_liz
&
@mimi1inh
on ML for structural biology and cryo-EM, and honored to be the first guest on The Plunge!
(P.S. I stuck around the filming location that day, and can confirm the rest of the episodes will be amazing!)
We teased this project during
#MMPortland2022
and are thrilled to finally share it with the world!
Introducing
#ThePlungePodcast
w/ co-hosts
@mcianfro
,
@kellogg_liz
&
@mimi1inh
!
Tune into E1 ft.
@ZhongingAlong
re:
#MachineLearning
,
#CryoEM
&
#newPI
life:
0
1
7
0
5
55
Interdisciplinary research = PhD BOGO 💸💸💸
Which is challenging but crucial for impactful ML applications in computational (and structural) biology
0
7
56
Had a great time in Heidelberg this past week at the EMBL Symposium on AI and Biology!
Pictured:
@alexrives
thinking deeply about working in large groups of people, and the party at the conference venue...
#EESAIBio
'AI and biology' day 4️⃣🙌🏻
Discussing 'Industry vs Academia' with:
🔹Oliver Stegle
🔹Wolfgang Huber
🔹Alex Rives
🔹Jennifer Listgarten
🔹Ellen Zhong
🔹Joshua Pan
@alexrives
: "There are some things that can only be accomplished by working in a large group of people"
#EESAIBio
0
0
30
2
3
55
I am incredibly excited to host John Jumper, lead of the AlphaFold2 team, at
@PrincetonCS
next week, Monday 4:30p ET. (And excellent timing with the new variant prediction model released today 🎉
#AlphaFold
#AlphaMissense
)
Talk details:
Please join us in person or virtually this Monday, Sept 25 at 4:30pmET. John Jumper from
@GoogleDeepMind
will present his talk: Highly accurate protein structure prediction with
#deeplearning
. Hosted by
@ZhongingAlong
, details here➡️
0
2
5
2
5
52
Check out
@axlevy0
's
@SLAClab
public lecture on our work for an accessible introduction to cryo-EM, AI, and structural biology! 👏👏👏
"Capturing molecular motion using artificial intelligence"
@fredericpoitev1
@GordonWetzstein
0
9
50
One of the highlights of 2021 has been organizing MLSB with my colleagues! And wow so many cool stories, science, and people across ML and structural biology yesterday! Talks are recorded but the gather town will live on in memory ✨
That's a wrap for MLSB 2021! We hope you enjoyed the talks, panel, and poster sessions! On behalf of the organizers, a huge thank you to our speakers, authors, reviewers, attendees, sponsors, and the
@NeurIPSConf
organizers for making this day happen!
1
3
35
0
7
48
This is so cool. Proteins can make balloons 🫧 Had a fascinating conversation at a poster session once about the biomechanics of these gas vesicles (though I think with someone from a different lab ?). Congrats to the authors!
1
8
49
Someone in the audience shared this ❄️🐉 pic with me from a talk at BPS. I wasn't in the session, but so cool!! A great first Biophysical Society meeting
#BPS2023
, plus motorbike tour of
@salkinstitute
/ La Jolla with
@PallavKosuri
and his lab
1
0
44
A huge congrats to my former mentor John Jumper and Demis Hassabis for the 2023 Breakthrough Prize in Life Sciences for AlphaFold!!
And incredible timing as we are talking about the
#AlphaFold2
papers in my class today with guest talk by
@mfigurnov
from the AlphaFold team! 🤩
“Change the world” is a phrase we hear a lot. But Demis Hassabis and John Jumper
@DeepMind
did. AlphaFold has solved the protein structure prediction problem, one of the central challenges in biology and medicine. They win the 2023 Breakthrough Prize in Life Sciences.
19
327
2K
0
3
42
I suppose cryoDRGN2 ❄️🐉 at
#iccv2021
counts as a NeRF paper! We revisit the problem of ab initio reconstruction (camera pose inference w conditional latent code in CV terminology). Separate thread and code coming soon!
1
0
42
First application paper of cryoDRGN ❄️🐉 to analyze dynamic ciliary motor complexes. Learned so much interesting biology (and chimerax tips) from
@alanbrownhms
,
@miao_gui
, et al. Thank you for being early adopters (v0.0.19 😯) of the method!
This work was led by
@miao_gui
in a wonderful collaboration with
@RuiZhangWUSTL
and
@DutcherLab
. Thanks also to
@ZhongingAlong
for getting us started with cryoDRGN - such a powerful tool for analyzing dynamics.
2
1
22
0
3
42
Great to see all the interest in using cryoDRGN ❄️🐉! This protocols paper describes the steps for analyzing the assembling ribosome dataset from the
@naturemethods
paper. Lots of hard work by
@LaurelKinman
and
@barrettmpowell
to create this comprehensive resource.
Want to try
#cryoDRGN
, but unsure how to get started?
@LaurelKinman
,
@barrettmpowell
,
@ZhongingAlong
, and
@lab_berger
provide step-by-step guidance (and new features) in a new protocol pre-print
1
73
293
1
5
41
Amazing! 🔥🔥🔥 cryoSPARC's new 3D classification can sort 100+ structures, now w/ fixed consensus poses. Curious how dependent the ensemble is on the initial model(s)
❄️ Just in time for the holidays:
#cryoSPARC
v3.3 is now available! ⚡
New 3D classification (BETA), auto-tuning blob picker, Ewald sphere & anisotropic magnification correction, performance improvements, bug fixes, and more for
#cryoEM
processing!
1
54
189
0
4
39
Recently did an interview about
#cryoEM
and
#cryoDRGN
❄️🐉 for a Dutch TV documentary -- just released (if you speak Dutch 🤗)! Thanks for the opportunity and fun chat about proteins and their impact on health and medicine
Cool to see a 3D printed model of a protein structure I solved (MAG, PDB 5LF5) on Dutch national television in a program about "tomorrow's medicine": (program mostly in Dutch). Also featuring
#cryoEM
,
@ZhongingAlong
with ❄️🐉 and ofcourse
#alphafold
!
1
0
6
4
4
40
Our review of "Deep Generative Modeling for Volume Reconstruction in Cryo-Electron Microscopy" is now published in JSB!
Paper:
Glad to have been able to contribute to this review!
✨New paper published in the Journal of Structural Biology!
We provide a methodological survey of deep learning methods for molecular volume reconstructions in cryoEM.
By
@ClaireDonnat
@axlevy0
@fredericpoitev1
@ZhongingAlong
@ninamiolane
0
3
27
1
3
38
Hugely excited about the potential of ML to continue driving transformations in structural biology. Check out the recording of the talks+panel from our
@NeurIPSConf
workshop!
And please leave us feedback!
0
5
38
Never had such a high production value science chat before (thanks
@thermosciEMSpec
), but a super fun convo nevertheless! Thankful to have amazing colleagues in cryo-EM who made this happen
@mimi1inh
@kellogg_liz
@mcianfro
Behind the scenes at
#MM2022Portland
with some incredible people in
#cryoEM
! We can’t wait to share what we have in the works with
@mcianfro
,
@mimi1inh
, and
@kellogg_liz
who kickstarted our special project today with
@zhongingalong
. More to come!
0
6
62
0
2
38
Register at to attend our ML for structural biology workshop
@workshopmlsb
on Mon Dec 13.
I'm ready for 🤯. Fantastic lineup of contributed talks and posters. Schedule coming soon:
Thrilled our work “Structure-aware generation of drug-like molecules” was selected for an oral presentation at
@workshopmlsb
:)
We generate molecular graphs *and* conformations conditioned on protein binding pockets
2
21
128
0
9
37
A belated post from some of our group (and friends
@sainingxie
) attending NYC vision day earlier this month!
2
2
37
So exciting!
#AlphaFold
your proteins!
Or just look it up if you work on homo sapiens/ other model organisms
Today with
@emblebi
, we're launching the
#AlphaFold
Protein Structure Database, which offers the most complete and accurate picture of the human proteome, doubling humanity’s accumulated knowledge of high-accuracy human protein structures - for free: 1/
100
3K
7K
0
3
36
A hypothesis generating machine for regulatory genomics 👩💻🧬👩🔬
Model and variant predictions openly available:
Published today in
@naturemethods
together with colleagues from
@calico
: Enformer - a transformer model that has led to greatly increased accuracy in predicting gene expression from DNA sequence.
Blog:
Paper: 1/
5
236
791
1
1
36
Congratulations Zeming!
#ESMfold
I defended my thesis! Big thanks to my committee
@ylecun
@rob_fergus
@kchonyc
@ZhongingAlong
@RichBonneauNYC
! Thanks to
@alexrives
and the entire Meta AI protein team as well,
@proteinrosh
@BrianHie
@TomSercu
and too many others to name.
Now on to more fun science!
23
11
194
0
0
36
Variant predictions from our covid spike language model from where we frame viral escape prediction as an NLP task. Note:
grammaticality == pseudo-log likelihood
semantic change == latent embedding delta
In line w what we currently know about Omicron
1
2
34
Gave my first in person talk earlier this week since NeurIPS 2019 and it was AMAZING! 😭 So excited to attend the upcoming in-person 💉💉 3DEM GRS/GRC and make up for all those virtual meetings. Looking forward to discussions on
#cryoEM
, new ❄️🐉features,
#AlphaFold
and more!
0
0
34
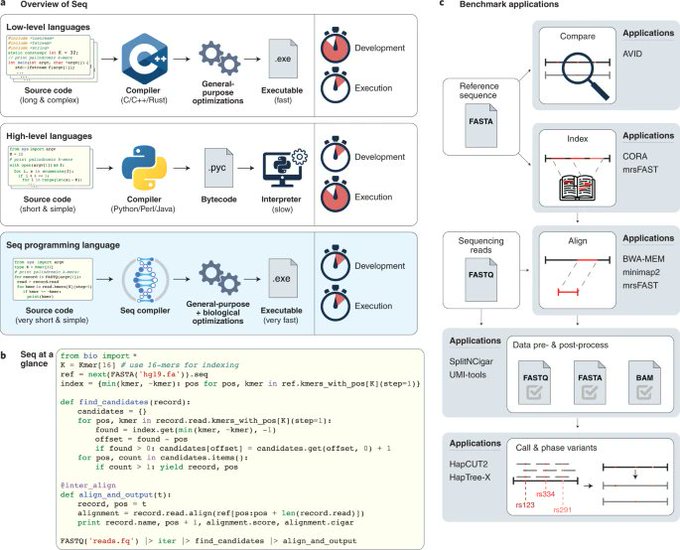
One of the coolest projects in
@lab_berger
! A compiled programming language specialized for genomics, but a drop-in replacement for Python. Congrats Ariya et al!
0
7
34
Talk to
@adamlerer
about anything from multi agent RL for games,
@pytorch
, molecular dynamics hardware, our recent cryoem reconstruction papers ❄️🐉2, and now large scale graph embedding tools for single cell genomic analysis! Congrats
@hd7chen
@lucapinello
et al!
Today we're releasing SIMBA, a tool for single-cell omics built on top of PyTorch-BigGraph. It’s been fun collaborating with
@hd7chen
,
@lucapinello
et al at the Broad to apply PyTorch-BigGraph, originally designed for large-scale web interaction data, to biology!
2
19
76
0
4
33
This was a fantastic collaboration with
@axlevy0
and
@fredericpoitev1
, who I just had the pleasure of hosting at Princeton last week,
@GordonWetzstein
, and
@jnpmartel
. 6/6
1
1
31
With the advent of
#AlphaFold
methods for structure prediction and rise of
#cryoEM
/
#cryoET
imaging, it is an incredible moment right now in structural biology. My lab will be focused on next-gen machine learning methods at this intersection. 3/
1
0
31
Yes! CryoDRGN in the ☁️☁️☁️
🚨New tool alert🚨
#cryoDRGN
is now on our gateway, allowing you to run
#cryoDRGN
on your
#cryoEM
particle stacks! A great tool provided to the community by
@ZhongingAlong
@jhdavislab
as well as great interactive visualization tools 👌
1
11
45
0
0
32
Please reach out if you're interested in joining the group, collaborating, or talking AI, proteins, and science! A huge thanks to everyone at
@Princeton
for the warm welcome! 5/
2
1
31
Witnessed an amazing seminar last week by
@vincesitzmann
@PrincetonCS
on progress and predictions for 3D vision foundation models. Thanks so much for visiting!
1
1
31
I just became affiliated with the QCB PhD program at Princeton. Apply to work with me! Catch the virtual open house today Fri 11/11 at 2pm ET for more info
Our interdisciplinary QCB PhD program
@Princeton
is amazing!
Great training and research opportunities.
Come join us to work with me and/or my colleagues!
Attend the open house on Fri Nov 11 to learn more:
1
3
16
0
5
30
Something I've touched on in every recent seminar -- we're in such an exciting era where new technologies around cryo-EM/ET structure determination can complement
#AlphaFold
/ML for structure prediction. So many new possibilities and 🆒 directions to watch! 👀
Exciting and interesting outlook on new tech for 2022
@Nature
- eg. on structure (and cryo-ET), long-read sequencing but also spatial multi-omics.
0
57
206
0
0
30
😍 cryoDRGN in the wild again! Beautiful structures of nucleosomes bound to the ALC1 (Amplified in Liver Cancer 1) protein ᗤ
Congrats on the preprint and a huge thank you for all your contributions and help triaging issues in the ❄️🐉github
@Guillawme
Would not have been nearly as interesting without the wonderful cryoDRGN ❄️🐉 from
@ZhongingAlong
!
0
1
4
2
1
30
Announcing our
@NeurIPSConf
workshop
@workshopmlsb
specifically focused on ML for proteins and macromolecular structure! Targeting methods for protein design, folding, ensembles, structure determination, and more!
Check it out:
#MLSB2020
#NeurIPS2020
Machine Learning in Structural Biology (
@workshopmlsb
) is accepted at
#NeurIPS2020
! Come check out the exciting line-up of speakers and dates for the call for papers at (more details coming soon). Register interest at
0
35
106
0
12
29
Good morning Berkeley! Excited to attend the AI=Science workshop
@SimonsInstitute
this week. Thank you
@jlistgarten
@ask1729
for organizing!
(Also we're slightly behind on tweeting about some of our recent papers... Stay tuned!)
1
0
30
A didactic review of the neural volume architecture fundamental to cryoDRGN ❄️🐉 placed in the broader context of other computer vision tasks etc
(I still tend to call it "coordinate MLP" but could be convinced on "neural fields")
Our review article on neural fields is out!
W/
@yiheng
@yongyuanxi
@psyth91
@orlitany
Shiqin Yan () Numair Khan ()
@fedassa
Sunny Li ()
@jtompkin
@drsrinathsridha
(equal advising). (1/n)
2
26
129
0
4
29
A flow generator for modelling protein motion from single particle cryo-EM.
A cool new approach for heterogeneous reconstruction from
@a_punjani
@fleet_dj
@structurabio
. Great to see more DL in the space! 🔥👏
Continuous flexibility is the next frontier for
#cryoEM
.
We are thrilled to introduce 3D Flexible Refinement, a new deep generative model that solves both detailed non-rigid motion, and high-res structure of flexible proteins!!
1/
12
184
751
0
7
29
I'll be going to New Orleans next week for
#CVPR22
🎷🎶🦐! Excited to give a talk about proteins, cryo-EM, and neural 3D reconstruction, Mon @ 4pm. Reach out if you'd like to chat and thank you
@vincesitzmann
and the other organizers for the invite!
Excited to be co-organizing a tutorial on Neural Fields in Computer Vision
@CVPR
. The agenda: try to provide a structured introduction/refresher of the latest advances in neural fields. Monday, June 20, Great Hall D. Please come! (1/3)🧵
7
93
451
0
2
28
We have been missing our weekly spliceosome reconstruction sightings (
@alkin_kaz
) :) Very cool work, congrats! 🔥
3
6
28
Amazing talks, posters, and energy at
@workshopmlsb
so far and excited to moderate a panel with our invited speakers at 2:50p CST! Please submit your questions here:
We'll also have an in-person panel discussion with our invited speakers
@fleet_dj
,
@alexrives
,
@wellingmax
, and
@tfgg2
from 2:50p-3:50p CST, moderated by
@ZhongingAlong
. Please submit any questions you would like to ask here: 4/
1
0
5
0
2
27
❄️🐉 in the wild! Very cool to learn about their structural adventure into the epitranscriptome ✨
0
2
27
🚨❄️🔬 cryo-EM twitter: Come join us at
@Princeton
! I've been blown away by the fantastic environment and support across all levels/departments/etc so far
The Department of Molecular Biology at Princeton University is seeking a cryoEM and/or cryoET specialist to serve as Manager of Biomolecular Electron Microscopy.
#cryoEM
#cryoET
#teamtomo
#structuralbiology
0
5
11
0
2
27
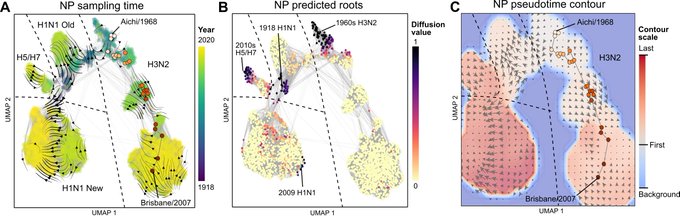
Very cool research on understanding the evolutionary roots of protein families from mining the openly available ESM-1b protein language model from
@facebookai
In some fun recent work with
@KevinKaichuang
and Peter Kim, we show that by using masked language models to predict local mutational effects, we can construct an evolutionary "vector field" -- kind of like RNA velocity, but for protein evolution!
9
68
304
1
4
26
Nice work by
@ebetica
and team
@MetaAI
introducing ESMFold, a single seq to structure model, which swaps the input MSA to
#Alphafold
with a large language model. Less performant than the full AF2 system, but faster + more convenient inference, with potential extensions to design.
We have trained ESMFold to predict full atomic protein structure directly from language model representations of a single sequence. Accuracy is competitive with AlphaFold on most proteins with order of magnitude faster inference. By
@MetaAI
Protein Team.
20
317
1K
0
5
26
You can find me at
#NeurIPS23
🎷🎶 this week! I will be
@workshopmlsb
💖 on Fri and the
@genbio_workshop
and the Deep Learning & Inverse Problems workshop on Sat. Please reach out if you want to chat!
0
3
26