Institute for Protein Design
@UWproteindesign
Followers
15,403
Following
664
Media
368
Statuses
2,963
We create proteins that solve challenges in medicine, technology & sustainability // Co-developers of @Foldit // @UWMedicine |
Seattle, WA
Joined September 2013
Don't wanna be here?
Send us removal request.
Explore trending content on Musk Viewer
Madonna
• 161964 Tweets
Amethi
• 94670 Tweets
Bucks
• 78713 Tweets
Knicks
• 68233 Tweets
#無責任でええじゃないかLOVE
• 61659 Tweets
Dame
• 59920 Tweets
Racing
• 57904 Tweets
Pacers
• 48751 Tweets
憲法記念日
• 45593 Tweets
Game 7
• 44485 Tweets
Sixers
• 43055 Tweets
Leafs
• 30624 Tweets
Brunson
• 27962 Tweets
dua lipa
• 25371 Tweets
Bruins
• 24380 Tweets
Rony
• 24126 Tweets
Giannis
• 22830 Tweets
Maxey
• 22797 Tweets
#PONDSXTZUYU
• 21843 Tweets
Estevão
• 19471 Tweets
Willy
• 17842 Tweets
Costas
• 15830 Tweets
76ers
• 15228 Tweets
Arias
• 12713 Tweets
設営完了
• 11757 Tweets
チャレンジクルー
• 11348 Tweets
Romulo
• 10798 Tweets
Buddy Hield
• 10681 Tweets
Last Seen Profiles
Pinned Tweet
Our latest AI tools for protein modeling & design were highlighted on the cover of
@ScienceMagazine
.
Read the open-access paper:
Use these tools for free:
RoseTTAFold All-Atom:
RFdiffusion All-Atom:
0
65
211
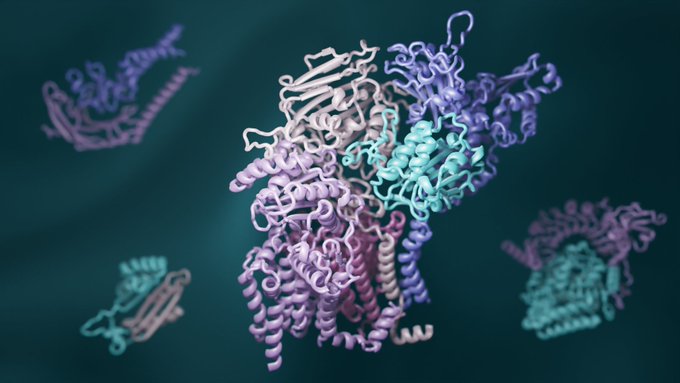
(1/8) Today we share the structures of protein complexes from nearly every core process in eukaryotic biology — from DNA repair to membrane trafficking.
This work builds on breakthroughs in coevolution & deep learning (
#Alphafold
,
#RoseTTAFold
).
2
277
761
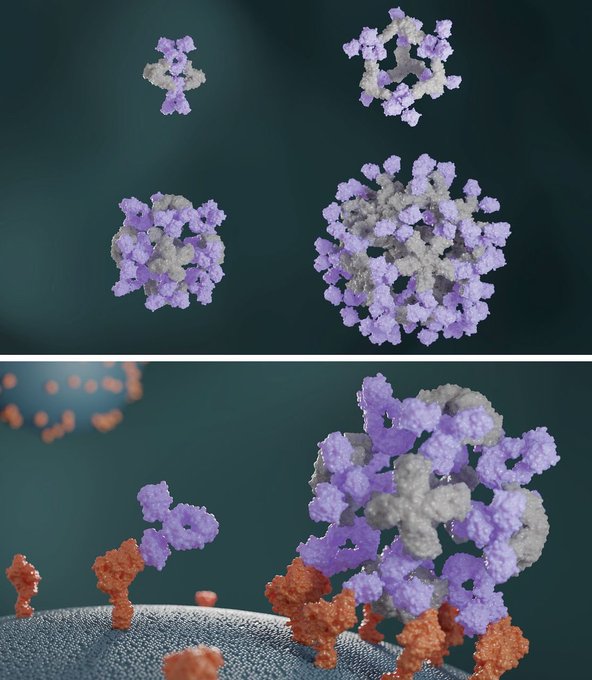
Generative AI for protein design is now one click away. We've launched a new platform that puts powerful tools behind a simple web interface, making
#proteindesign
accessible to anyone.
Try it here ➡️
Learn more on YouTube ➡️
7
200
677
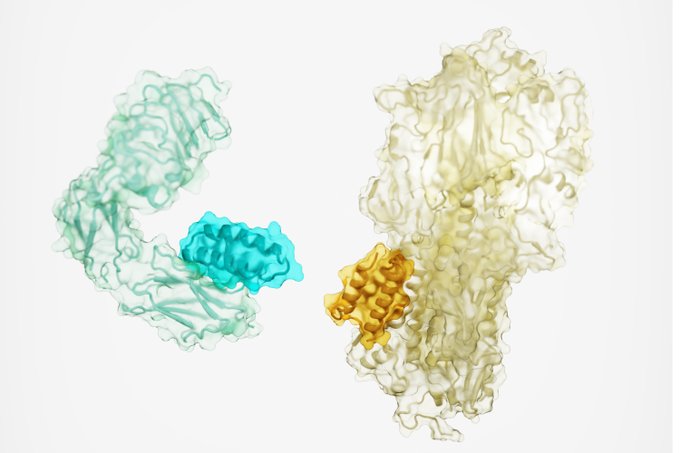
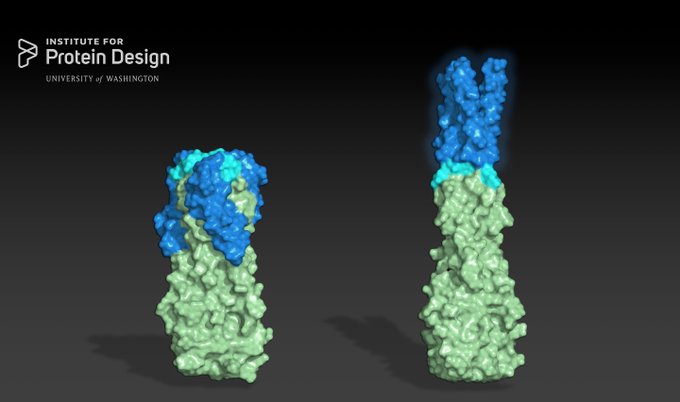
Design of protein binding proteins from target structure alone |
@Nature
Our approach now enables targeted design of binders to sites of interest on a wide variety of proteins for therapeutic and diagnostic applications.
#proteindesign
4
165
544
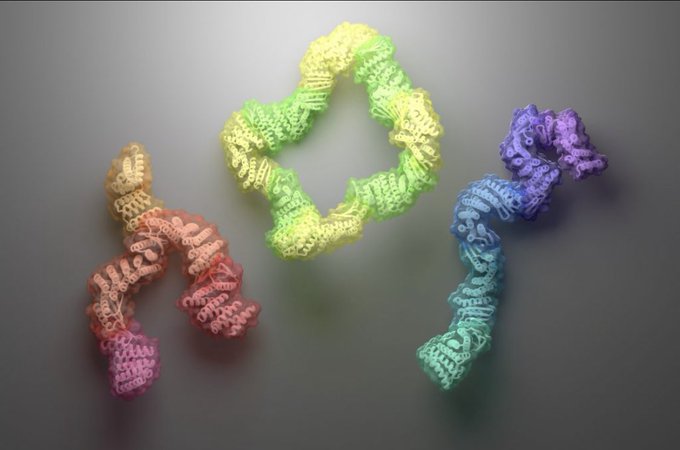
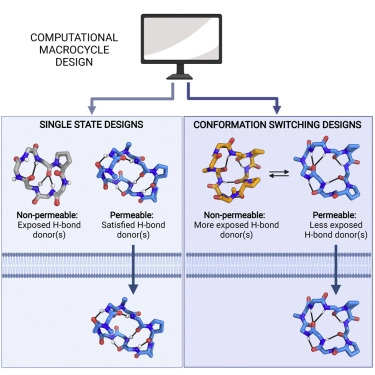
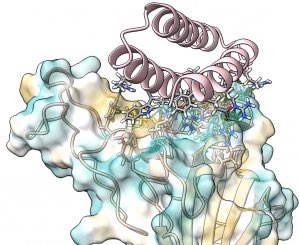
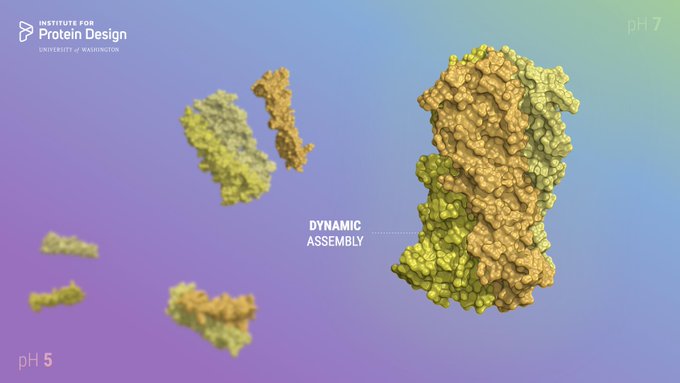
Scientists in the Baker Lab have designed new proteins that change shape upon binding a target molecule. This research opens the door to new applications in drug delivery, environmental sensing & more.
@FloPraetorius
,
@definitelyphil
4
101
407
“This really is a big deal” — David Baker on today's
#AlphaFold
news.
From our team to
@DeepMind
: Congratulations! You have electrified this important field. We look forward to what can be accomplished in the coming years.
0
74
332
Today we report the use of reinforcement learning, a strategy used to solve games like chess and Go, to a challenge in protein design. This milestone in using
#AI
for science may enable more effective cancer treatments & new biodegradable textiles.
5
87
293
Rosetta Commons is pleased to once again offer (paid) undergraduate internships and postbac fellowships in biomolecular structure prediction and design!
Learn about
#deeplearning
breakthroughs and how to apply them to create new therapies, vaccines & biotechnologies. 1/3
8
85
290
We are honored to see our work on protein structure prediction recognized as the 2021 Breakthrough of the Year.
We also applaud
@DeepMind
for invigorating the field and contributing so much to open science.
All areas of computational and molecular biology will be transformed.
AI algorithms can now churn out predictions for the 3D shapes of proteins with a precision matching that of painstaking laboratory techniques. The programs, and the blizzard of protein structures they have revealed, are Science’s 2021
#BOTY
.
45
451
1K
3
56
285
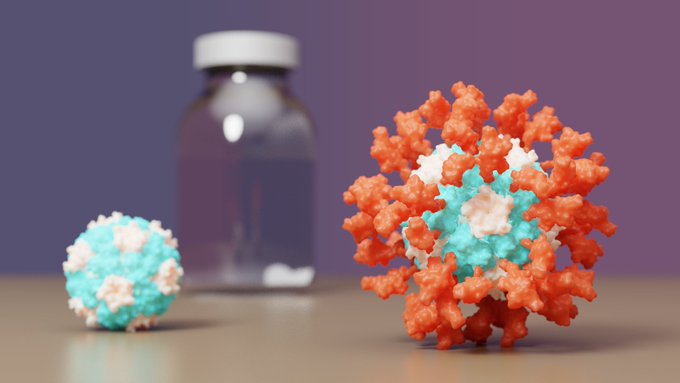
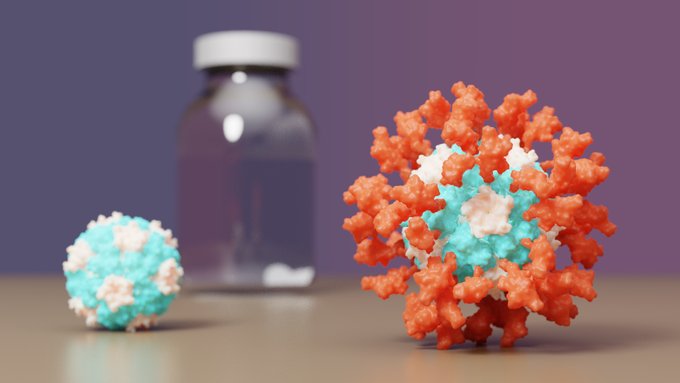
SKYCovione, a COVID-19 vaccine developed by
@KingLabIPD
&
@veeslerlab
, has just won full approval abroad!🇰🇷This protein-based vaccine outperforms Oxford/AstraZeneca's and does not require freezing. It is our first designed protein medicine.
Learn more:
2
92
280
Just as convincing images of cats can be generated using artificial intelligence, new proteins can now be made using similar tools. In a new report in
@Nature
, we describe a neural network approach for “hallucinating” proteins with new, stable structures.
2
75
281
David Baker will present the
@MRC_LMB
2021 Max Perutz Lecture, titled ‘The coming of age of de novo protein design’ at 8:00am (Pacific) on Tuesday, April 27. Attendance via Zoom is free & open to all.
3
73
268
This is a historic day for protein design: New computer-generated proteins fight
#cancer
with fewer side effects.
#immunotherapy
#proteindesign
#neoleukin
@nature
3
111
231
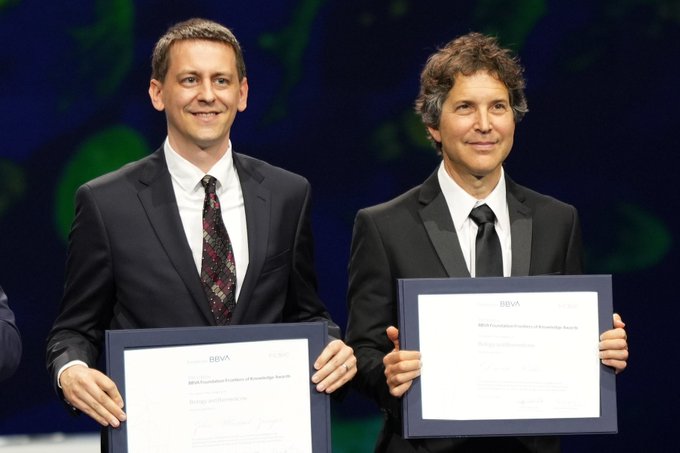
David Baker, together with Demis Hassabis & John Jumper of
@DeepMind
, has received the Frontiers of Knowledge Award from
@FundacionBBVA
for "contributions to the use of A.I. for the accurate prediction of the 3D structure of proteins."
2
37
237
#RoseTTAFold
: Accurate prediction of protein structures and interactions using a three-track neural network. By
@minkbaek
et al,
@ScienceMagazine
3
66
226
#ProteinMPNN
, which is now available free on the open-source software repository GitHub, will give researchers the tools to make unlimited new designs. “The challenge, of course, is what are you going to design?”
GitHub:
1
44
225
The Baker Lab has a new
#openaccess
paper in
@NatureComms
showing that dramatic improvements can be achieved when deep learning software is used to design protein-binding proteins.
Full paper ➡️
Short summary ⬇️
1
59
213
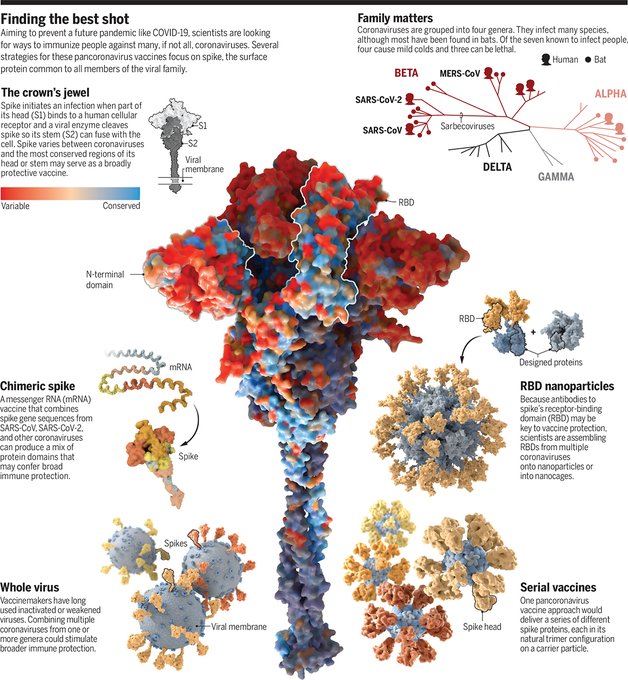
.
@ScienceMagazine
has published a beautiful profile of our efforts to design a vaccine that protects against many coronaviruses at once. A close collaboration between NIAID,
@KingLabIPD
,
@veeslerlab
,
@WardLab1
, & others.
The dream vaccine:
0
53
185
Today we report new
#deeplearning
approaches for functional
#proteindesign
. We use these to generate immunogens, receptor traps, metalloproteins, enzymes, and binder proteins. We validate the designs using in silico and lab tests.
2
58
175
Generative AI for protein design is now one click away. We've launched a new platform that puts powerful tools behind a simple web interface, making
#proteindesign
accessible to everyone.
Full video ➡️
Try it here ➡️
3
42
178
Have you had a chance to use RFdiffusion?
We would love to hear about it! Please share your story here or send us an email: contact
@ipd
.uw.edu
4
39
175
S. Korea has just approved a Phase 3 trial of the COVID nanoparticle vaccine we reported last year in Cell: All Phase 1/2 trial participants induced neutralizing antibodies.
@KingLabIPD
@veeslerlab
@UWMedicine
5
52
169
A
#deeplearning
approach to protein design: When combined with detailed physical models, AI can capture properties normally only accessible through large-scale simulations.
Protein sequence design by conformational landscape optimization |
@PNASNews
0
48
159
IPD director David Baker, together w John Jumper and
@DemisHassabis
of DeepMind, will receive the 2022 Wiley Prize in Biomedical Sciences for pioneering work on protein structure prediction. Their award lectures will be streamed April 1. Learn more at:
0
33
150
Dr. Anthony Fauci shared some news with Congress about our collaborative influenza vaccine research. Phase 1 trials are now underway!
Paper is by
@DanLikeProteins
,
@KingLabIPD
,
@kanekiyom
,
@BarneyGrahamMD
, & many more.
1
49
140
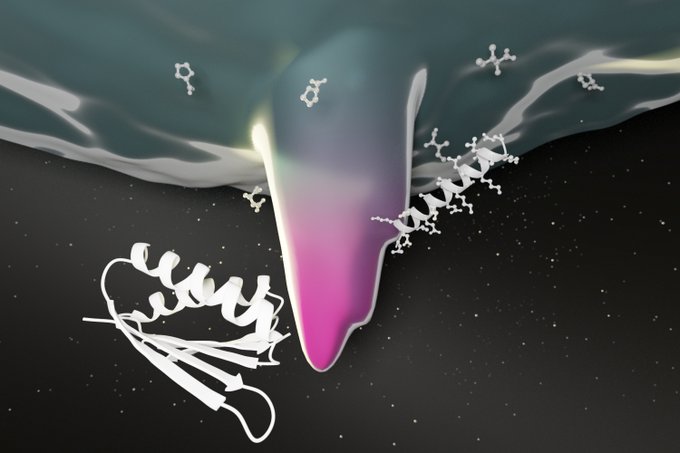
New protein-based biosensors have been designed from scratch to enable direct detection of
#SARSCoV2
particles. Additional sensors that detect COVID-19 antibodies are also described in our latest preprint: |
#ProteinDesign
1
48
133
In a new report, we describe the de novo design & characterization of selective ion channels, as well as larger pores that permit the passage of fluorescent molecules thru cell walls.
Computational design of transmembrane pores
| |
@nature
#ProteinDesign
2
38
115
The COVID19 vaccine developed by
@KingLabIPD
&
@VeeslerLab
has met its Phase 3 trial goals and is now under review for full authorization in Korea.
SK bioscience, the company leading clinical development abroad, found the vaccine outperformed Oxford/AstraZeneca’s in the clinic.
1
37
108
New in
@NatureComms
: "Improved protein structure refinement guided by
#deeplearning
based accuracy estimation"
0
37
107
Our trRosetta
#deeplearning
method has been applied to all protein families of unknown structure, yielded 6,370 new protein models. All now available in
@PfamDB
, with 3D structure views and contact maps in
@InterProDB
. Happy exploring! 🔬
1
27
108
Peptide-binding specificity prediction using fine-tuned protein structure prediction networks | PNAS
@AMotmaen
,
@UWBiochemistry
0
23
106
Deep learning has revolutionized protein structure modeling. Here, David Baker and
@minkbaek
relate
#AlphaFold
and
#RoseTTAFold
to classical approaches to structure prediction and discuss the areas of structural biology that are likely to be transformed.
1
25
104
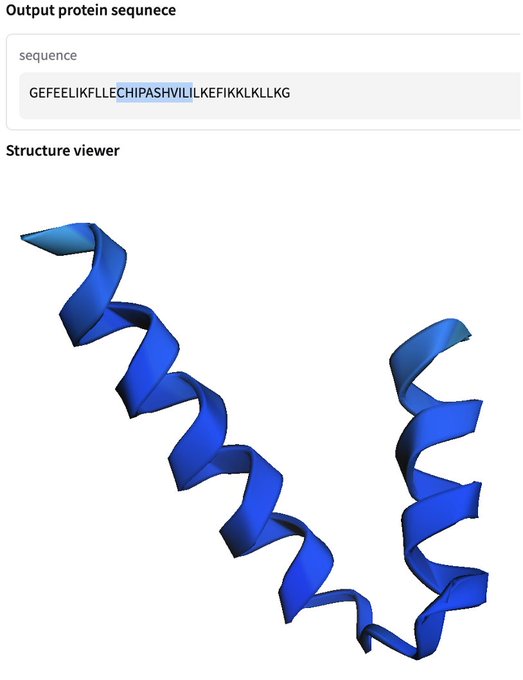
Tired: Writing your name on a grain of rice ✍️
Wired: Writing your name into an AI-generated protein 🧬
Try it for free in your browser ➡️
0
16
101
RF Diffusion was developed by a team of computational biologists from
@UWProteinDesign
,
@Columbia
&
@AIHealthMIT
.
Our thanks to
@sokrypton
for implementing it onto the ColabFold platform.
0
25
99
From modeling viral proteins to creating candidate vaccines, our scientists are taking on COVID-19.
• 11,000+ de novo antiviral proteins are in testing
• Nanoparticle vaccines already in animals 🐁 (
@KingLabIPD
,
@veeslerlab
,
@NIH
)
Learn more:
1
34
99
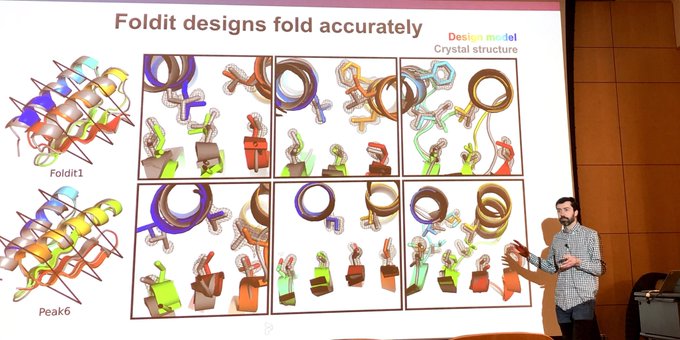
Today's an exciting day!
After many years of work, we're thrilled to report our latest advance in citizen science:
@Foldit
@UWBiochemistry
@uwcse
@RutgersU
@dartmouth
@Northeastern
@HHMINEWS
#citizenscience
#proteindesign
@nature
2
47
99
Honored to see our work on
#deeplearning
for protein structure prediction featured on the cover of
@ScienceMagazine
. Research was led by the tremendously talented
@minkbaek
.
Article:
Commentary:
0
20
93
A COVID19 nanoparticle vaccine designed
@UWMedicine
(
@KingLabIPD
,
@veeslerlab
) has achieved its Phase 1/2 clinical goals! In a trial led by SK Bioscience, the vaccine was found safe and generated high levels of neutralizing antibodies. Ph3 is underway.
4
36
92
Deep networks trained to predict native protein structures from their sequences can be inverted to design new proteins |
@biorxivpreprint
1
30
94
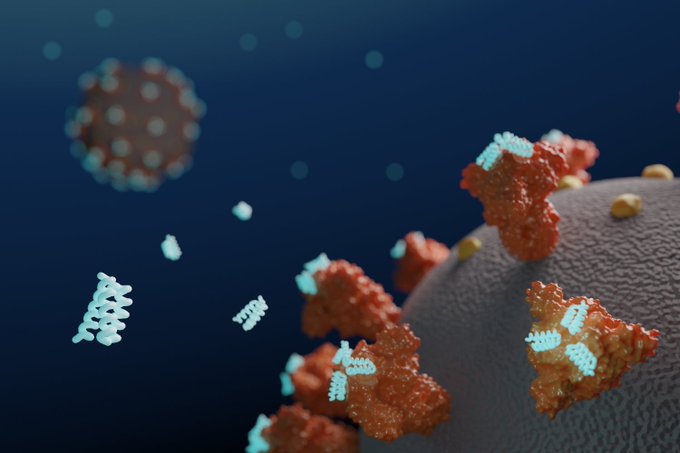
New: Synthetic proteins designed to bind Spike on SARS-CoV-2 provide starting points for COVID-19 therapeutics.
@UWMedicine
@veeslerlab
@WUSTLmed
De novo design of picomolar SARS-CoV-2 miniprotein inhibitors |
1
50
92
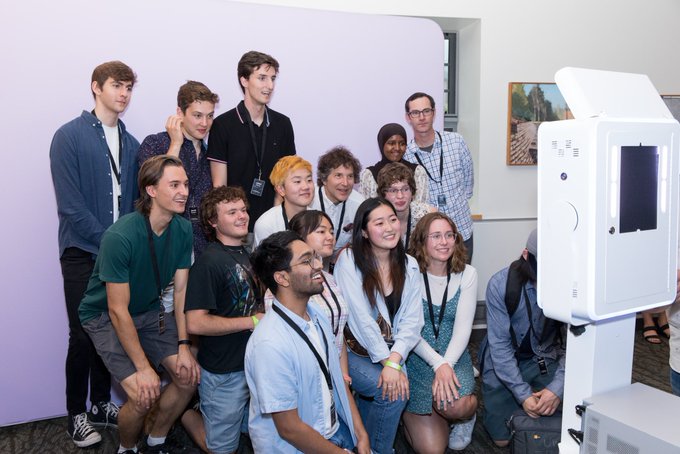
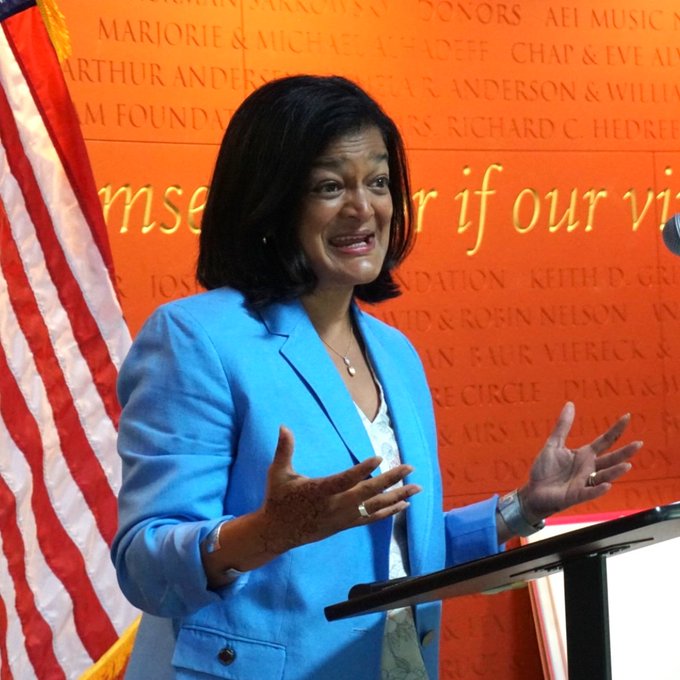
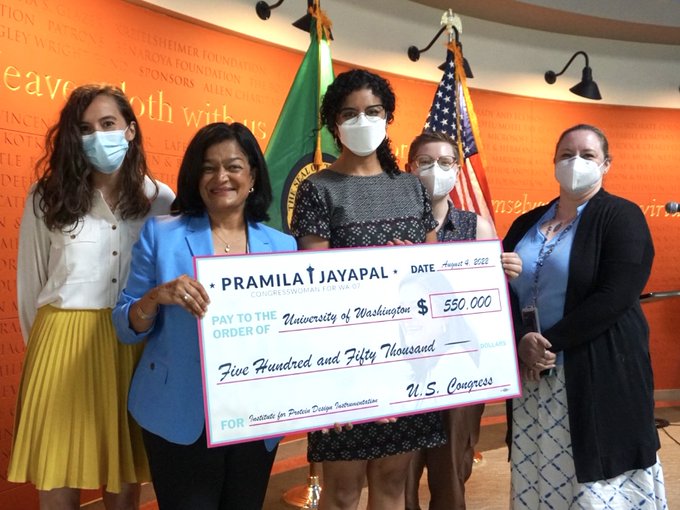
Today our admin team met with
@RepJayapal
to celebrate new Federal investments in 10 key projects across the Seattle area. Our request will enable the purchase of advanced laboratory equipment that will help us discover new cures for current and future pandemics.
2
13
81
Today we report the design of protein sequences that adopt more than one well-folded structure, reminiscent of viral fusion proteins. This research moves us closer to creating artificial protein systems with reliable moving parts.
#ProteinDesign
1
23
81
Protein designers can't make molecules that move, right?
Out today: "De novo design of tunable, pH-driven conformational changes"
@sciencemagazine
#proteindesign
1
26
77
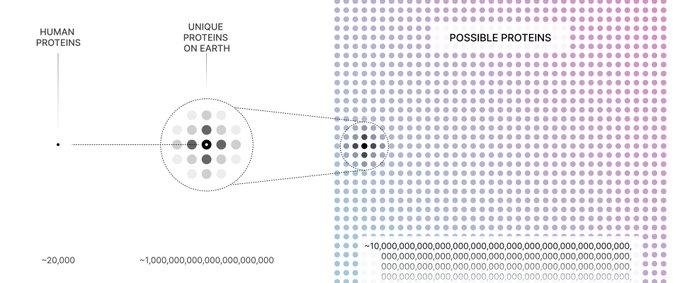
@_biojmg
We think it's the third option: three billion years of natural selection is not nearly enough time to sample even one percent of the vast protein universe. There have got to be remarkable —and useful — molecules still out there.
1
11
78
There’s a new profile of our research out today in the
@NewYorker
. Features perspectives from
@francesarnold
& other luminaries in the field
#ProteinDesign
#MachineLearning
#SARSCoV2
@Foldit
@RosettaAtHome
|
@SilverJacket
1
35
74
New this week:
@definitelyphil
and
@FloPraetorius
have created “hinge” proteins that adopt two shapes: one with ligand, one without.
This control over molecular motion sets the stage for new applications in smart medicines and beyond.
0
17
74
Preclinical data out today: Our nanoparticle vaccine candidate for COVID-19 elicits ten times more neutralizing antibodies than vaccination with Spike alone.
Led by
@KingLabIPD
,
@veeslerlab
|
#ProteinDesign
1
26
72
Congrats to Brian Koepnick, Baker lab, for successfully defending his PhD today!
Brian has helped a community of over 700k citizen scientists gain access to advanced tools for protein design.
@Foldit
#citizenscience
2
13
71
Reminder: Researchers on our COVID-19 vaccine design team will soon be answering questions live for a Reddit AMA!
They'll be joined by
@Foldit
scientists working on antivirals.
Monday, Mar. 30
11:00 AM PT
2:00 PM ET
6:00 PM GMT
Look for us:
1
19
69
De novo design of modular and tunable protein biosensors |
@Nature
New sensors detect coronavirus proteins and antibodies |
@UWMedicine
0
26
71
Dive inside our efforts to design second-generation COVID-19 vaccines & treatments with this new feature in Scientific American.
@coronalexington
,
@veeslerlab
,
@KingLabIPD
,
@UWMedicine
,
@BarneyGrahamMD
|
@sciam
,
@rowanjacobsen
0
24
71
Regulators in the UK have granted full approval to SKYCovione, the computationally designed COVID-19 vaccine pioneered by
@KingLabIPD
and
@veeslerlab
|
@UWMedicine
0
18
66
Congratulations to IPD founder David Baker for being named a 2018 Highly Cited Researcher. Dr Baker's publications are among the world's most influential, having achieved the highest percentile of peer citations in Biology & Biochemistry
#HighlyCited2018
@webofscience
0
13
67
Today in
@Nature
: Programmable design of orthogonal protein heterodimers
cc:
@UWBiochemistry
0
41
65
In this new video, the team at
@SciShow
explains how A.I. is transforming protein science.
"The future of biology is starting to take shape." —
@hankgreen
WATCH:
#RoseTTAFold
#AlphaFold
#AlphaFold2
1
22
65
New reporting on our de novo designed COVID-19 antivirals: "When spritzed up the noses of mice & hamsters, it also appears to protect animals from becoming seriously sick."
#ProteinDesign
0
22
60
Our friends at CellSpace are bringing beauty to this dark moment with a galley of art focused on the molecular biology of SARS-CoV-2. Included in the collection is this piece by
@dr_opatel
depicting our antiviral miniproteins (CC-BY-4.0)
Full gallery:
1
14
63
New from our spinout
@neoleukin
: "De novo design of potent and resilient hACE2 decoys to neutralize SARS-CoV-2" |
@ScienceMagazine
#ProteinDesign
0
13
62
Meet LOCKR, a new synthetic protein switch. Its design and initial applications are detailed in dual reports out today in
@Nature
.
@UWBiochemistry
@UCSF
@HanaScientist
@langanbiotech
@andrewng_synbio
1
25
59
New: Designed biosensors that can detect neutralizing antibodies against SARS-CoV-2 in blood.
These quantitative devices may aid in monitoring vaccine durability.
#proteindesign
with
@msdiamondlab
@veeslerlab
1
24
60
The University of Washington’s Institute for Protein Design has become a hub to galvanize de novo protein engineering, citizen science and much more since its founding in 2012 |
@NatureBiotech
0
19
59
Our director David Baker recently shared his thoughts on the impact of AI on life sciences and healthcare startups. He spoke with Jenny Cronin at a
@UWCoMotion
event.
0
10
58
Ultrapotent miniproteins targeting the SARS-CoV-2 receptor-binding domain protect against infection and disease.
Collaborative work w/ Diamond Lab
@WUSTL
,
@WUSTLmed
|
#proteindesign
,
@TheAudaciousPrj
1
18
58
Are you (or someone you know) interested in postdoctoral research in protein design? We are always accepting new
#postdocs
!
Join us for an informational open house — Jan 13, noon PT
Sign up here for Zoom link: |
#ProteinDesign
#postdocjobs
3
24
57
Deep
#neuralnetwork
that outputs relative distances & orientations of all residue pairs in a
#protein
aids in predicting protein structures — especially for proteins that have been designed (rather than evolved).
#proteindesign
#machinelearning
@PNASNews
0
28
57
Congratulations to Frances Arnold on her
#NobelPrize
in
#chemistry
! She is true pioneer in showing the world the awesome power of proteins.👩🔬 📷:
@Caltech
0
16
53