Nicky Whiffin
@nickywhiffin
Followers
3,628
Following
761
Media
234
Statuses
1,931
Associate Prof & Henry Dale Fellow @bdi_oxford @HumanGeneticsOx | Computational Rare Disease Genomics | Non-coding | @StAnnesCollege | @nickywhiffin .bsky.social
London, UK
Joined November 2016
Don't wanna be here?
Send us removal request.
Explore trending content on Musk Viewer
Ireland
• 365634 Tweets
#WeAreSeriesEP8
• 352151 Tweets
Norway
• 315117 Tweets
Spain
• 272714 Tweets
#MissUniversePhilippines2024
• 155834 Tweets
Apple Music
• 134961 Tweets
PondPhuwin WeAre EP8
• 130430 Tweets
Blonde
• 89099 Tweets
LOST TEASER
• 88714 Tweets
KATY PERRY
• 67086 Tweets
Irlanda
• 54961 Tweets
Noruega
• 49693 Tweets
PREPARE FOR MLMU
• 47265 Tweets
#RCBvsRR
• 45866 Tweets
Rishi Sunak
• 39710 Tweets
KING KONG MV TEASER
• 38163 Tweets
Thriller
• 37402 Tweets
Atalanta
• 36760 Tweets
Lauryn Hill
• 32936 Tweets
مورينهو
• 31154 Tweets
Maxwell
• 24297 Tweets
Frank Ocean
• 24034 Tweets
TPAB
• 15569 Tweets
ガッキー
• 14937 Tweets
$BULL
• 14525 Tweets
Sokak Köpekleri Toplatılsın
• 12115 Tweets
#GeneralElection
• 12051 Tweets
GKMC
• 11748 Tweets
Ashwin
• 10345 Tweets
Last Seen Profiles
Pinned Tweet
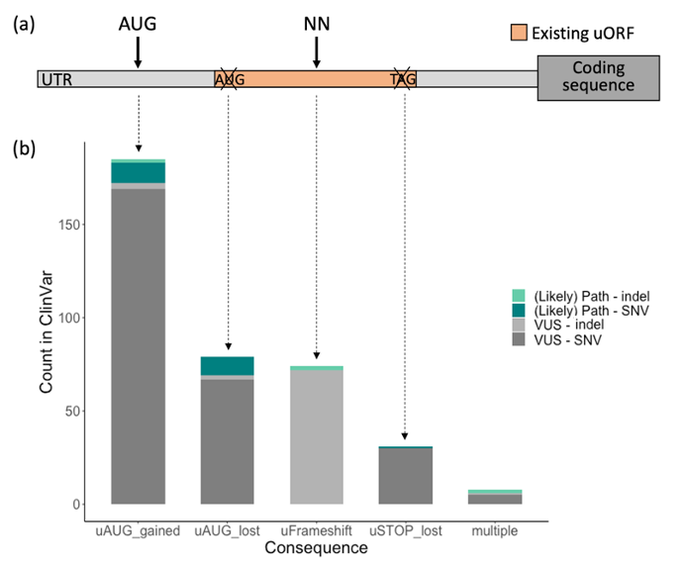
Would you believe me if I told you that a single variant in a non-coding RNA explains ~0.5% of all undiagnosed individuals with neurodevelopmental disorders (NDD) in
@GenomicsEngland
???
I didn’t initially either, but here is the story of RNU4-2 🧵1/9
15
213
643
Hugely excited to announce that I have been awarded a
@wellcometrust
Sir Henry Dale Fellowship and will move to
@HumanGeneticsOx
@UniofOxford
in September (hopefully physically, if not then virtually) to set up my lab!!! 1/3
59
15
428
🎉Out today in
@GenomeMedicine
: our recommendations for clinical interpretation of variants in non-coding region of the genome
A huge team effort, co-led by the super awesome
@j_ellingford
!
🧵
8
117
344
Public release of first ~150k
@uk_biobank
whole genomes will be in early November! 😀
1 variant every 4-5 bps, 44% singletons
8.1% coding variants not in WES (85.3% UTR variants)
STRs and SVs also called
Improved regulatory variant annotation hugely needed for analysis!
#ASHG21
4
79
314
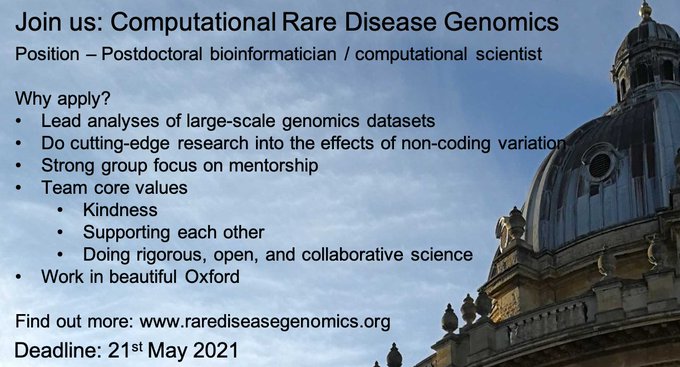
**Postdoc & PhD opportunities**
Interested in using large genomic datasets to discover and analyse non-coding variants that cause rare diseases?
Come work
@HumanGeneticsOx
@UniofOxford
Please share and RT!
But why should you join my team? A thread 1/4
3
173
216
Today the Computational Rare Disease Genomics group, or Whiffin lab
@HumanGeneticsOx
@NDMOxford
becomes a reality 🎉
This isn't at all how I imagined this day would go (without even leaving my house!), but I am excited to meet lots of new colleagues, even if virtually for now 👋
10
12
187
16 months since the first was on
@biorxivpreprint
, and just over a year since the first was accepted, the 7 gnomAD papers are finally out in
@nature
,
@NatureMedicine
and
@NatureComms
:
Incredibly excited to have been part of this effort!
#teamScience
(1/5)
6
41
141
🚨 New preprint 🚨
"Non-coding variants are a rare cause of recessive developmental disorders in trans with coding variants"
Fantastic team effort co-led with incredible trio
@DrJennyLord
,
@hilsomartin
and
@ProfDBaralle
🧵1/6
9
50
140
Our paper on MEF2C non-coding region variants in developmental disorders is out today in AJHG
@AJHGNews
🎉
Blog post on our key findings and their impact:
1/2
4
32
137
Loving the celebration of awesome women in science at
#ESHG2020
- Leena Peltonen Prize =
@GosiaTrynka
- Mendel Lecture = Aviv Regev
- ESHG Award Lecture =
@wendy_bickmore
#womenInSTEM
😍
0
16
123
Finally, there is an awful lot of data across these 7
#gnomAD
papers - I have written a blog post that drills into some important lessons they teach us, specifically about clinical variant interpretation:
I hope it is useful! (5/5)
1
49
118
Me, actually at my new desk in
@HumanGeneticsOx
😄 Only took me 8 months to get here! Seriously big shoes to fill as the last occupant was
@annagloyn
🤩 (
@ProfJohnATodd
and I have questions about the whiteboard unicorn...).
4
2
117
Delighted that my team will join
@bdi_oxford
in November. I will co-lead the BDI genomics theme with the amazing
@astheeggeggs
. We will remain associated with
@HumanGeneticsOx
- excited to build stronger links between the two institutes and genomics more broadly across Oxford🧬💫
4
4
114
Attending
#ESHG2021
this weekend?
Check out the on-demand session 'Variant Interpretation in the Clinic' with me and the incredible
@HeidiRehm
I will talk about our work on 5'UTR variants and give a preview of how we are developing guidelines for non-coding region variants.
3
21
109
An incredible trio of kick-ass women in science gracing the
#ASHG22
awards stage this evening. Congratulations
@HeidiRehm
(Curt Stern),
@EimearEKenny
(Early career 2022) and
@cristenw
(Early career 2021). You are all awesome and hugely deserving 🌟🍾
#roleModels
3
11
108
I absolutely love my work, but I also know that my weekends are hugely important to rest and reset so I can give my all to both my science and my team the next week. Please let's stop normalising this unhealthy culture of overwork!
#AcademicChatter
#MentalHealthAwareness
2
13
108
It has been somewhat of an overwhelming afternoon. I would like to take a moment to thank all of those who support and uplift me, my mentors, and my absolutely wonderful team - I wouldn't be here without every one of you. Thank you ❤️
Now time to open some 🍾
proud of my faculty
@bdi_oxford
who are smashing it in the recognition of distinction activities - following hot on the heels of last years success with two more associate professors
@nickywhiffin
&
@jeromekelleher
you are both fabulous. I am so proud to have you on the team
2
2
31
26
3
105
Today it is a year since I moved to
@HumanGeneticsOx
and started the Computational Rare Disease Genomics Group 🥳🍾
To mark this, I have written a blog post summing up our first year and reflecting on setting up a group in the midst of a pandemic:
4
10
98
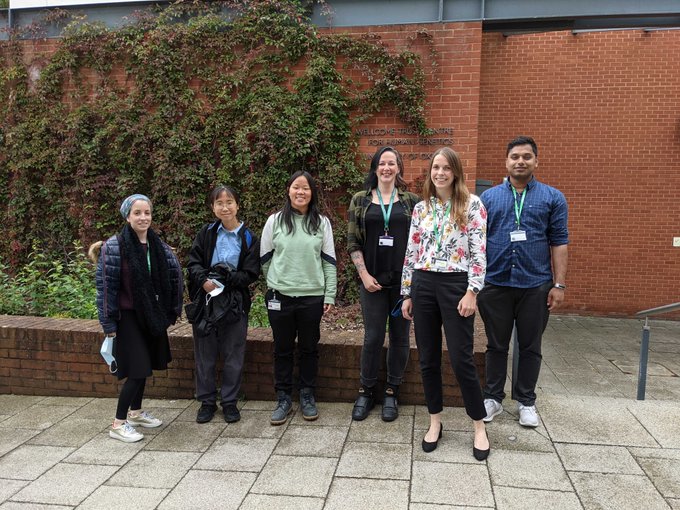
New team photo in our new home
@bdi_oxford
Love this group of amazing, talented, and inspiring individuals ❤️
2
6
93
Introducing the next gnomAD pre-print - in collaboration with
@23andMeResearch
we use highly curated genomic and phenotypic data from >4 million individuals to characterise the effects of life-long LRRK2 reduction in 1,358 loss-of-function carriers: 1/5
1
41
90
We are getting amazing emails from people who have identified individuals with RNU4-2 variants ❤️
If this is you, and your families are interested in meeting others or being part of a community, then please direct them to
@Unique_charity
(
@swynn_unique
).
1
26
90
Update on
@uk_biobank
whole-genome sequencing - ~150k samples sequenced already (will be made available to all in 9 months!). Working on a cloud-based environment to host the huge amounts of data created. Initiatives to broaden access to data (incl. free access models)
#UKBCONF20
0
27
85
First team photo 🥰
@Alextremophile
@NechamaWieder
Today we finally learnt how tall eachother are!
Thank you to fab photographer
@Jane_Pernes
😊
6
2
86
Friday thought:
Even a single person can influence team culture - small messages to say "you got this!" before a talk/meeting, or to tell people how great their presentation was have a knock-on effect - people begin to pass them on to others
#supportEachother
#PositiveVibes
1
6
81
📢📢 Join our team and study for a PhD in an amazing research environment in beautiful Oxford
@HumanGeneticsOx
@UniofOxford
Key focus on mentorship, teamwork, kindness, collaboration, and exciting science.
Two exciting opportunities - see 🧵👇
Please share and retweet! 1/3
4
39
78
Four years ago
@GeneticsSociety
established an 'Early-Career Award'. All four of the awardees to date have been male.
Please join me in nominating some kick-ass female geneticists for this year’s award
1/3
5
14
79
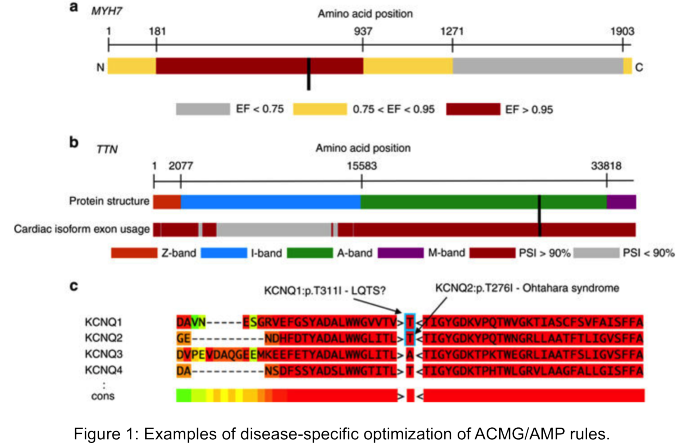
Our paper describing is now online
We show the importance of refining the ACMG/AMP guidelines for individual genes and diseases and demonstrate the power of automation for fast and accurate variant interpretation.
@GIMJournal
0
32
76
Just wanted to say a huge thank you to my Twitter network for all your job advert retweets - I am super excited to have an awesome DPhil student and fantastic postdoc both starting next month! 🎉
@HumanGeneticsOx
I will let them reveal themselves so as not to spoil all the fun.
3
1
75
Sad to be leaving such an awesome bunch of people
@CVGenomics
@ImperialNHLI
this week 😭
Thank you for my AMAZING start codon necklace (as close to uORFs as they could get). I love it! 😍🤓
Will miss you all terribly!
Thanks for making me the person and scientist I am today!
2
1
70
Excited to join
@StAnnesCollege
from today as a Junior Research Fellow working with the Centre for Personalised Medicine
@CPMOxford
🧬🩺💊
8
1
70
We are out of seats in the variant classification session at
#ESHG2023
where
@HeidiRehm
and
@stevieharri
are showing a preview of the next iteration of the ACMG/AMP (/CAP/ClinGen) guidelines. Moving to a more granular points based system.
4
10
70
**Job opportunities**
Interested in driving analyses of large genomic datasets for insights into causes of rare disease?
Openings for a postdoc and a research assistant
@HumanGeneticsOx
@UniofOxford
Please share and RT!
But why should you apply? 1/3
1
60
69
On
#rareDiseaseDay21
I remember my cousin Jonathan, who died from DMD aged 18. His diagnosis started my interest in genetics (why him?) and still fuels my drive today.
My awesome Aunt set up the J's hospice in his memory, which is still going strong today:
0
5
69
📢 Calling all early-stage PIs across
@UniofOxford
📢
Are you interested in networking and collaborating with others across the university? Would you value a peer support network?
I am on a mission to set up a network to join us up. DM/email me to sign-up.
Please share and RT!
3
36
68
For anyone still around at
#ASHG23
#ASHG2023
(and alive after last night's gala) I am speaking in the next session "Genetic diagnosis of neurodevelopmental disorders beyond the exome"🧬 about collaborative work with
@DrJennyLord
@ProfDBaralle
and
@hilsomartin
4
4
66
The future of rare disease diagnosis is multi-omic. Available in 2024: 7,500
@GenomicsEngland
individuals with long-read sequencing (
@oxfordnanopore
), metabolomics (
@metabolons
), transcriptomics, and proteomics, along with of course genome sequencing
#GERS2023
2
8
65
We are looking for a postdoc to join our awesome team
@HumanGeneticsOx
Apply here:
Please share and RT!
#AcademicTwitter
#ASHGtrainee
#ScienceJobs
0
57
64
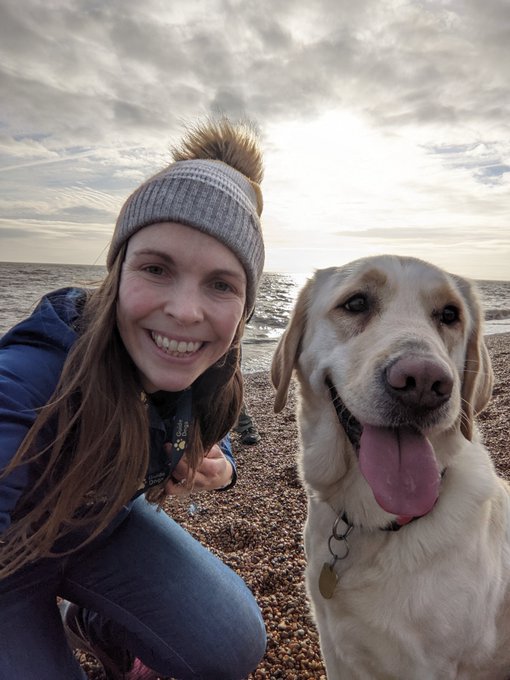
The hardest start to a week. After 6 months we just said goodbye to beautiful Kelsey. Although being a Guide Dog wasn't for her, she is heading to be a buddy dog for a child with sight-loss. She will be amazing, but I will miss her giant, fluffy cuddles! 😢🐾🐕🦺
#lifeChanger
2
1
64
The email auto-reply is on and it is time for 2 weeks of holiday ☺️ (just as the nice weather has disappeared...) When I come back I will be a full blown PI 😱
#grownUpScientist
#shitJustGotReal
1
1
61
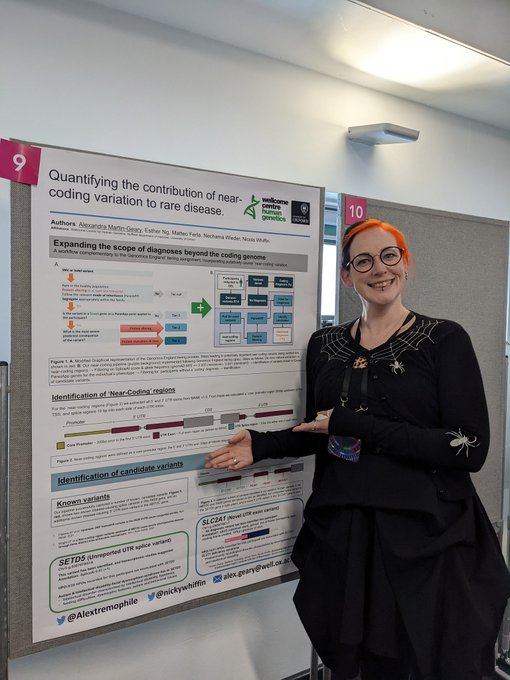
At the
@GenomicsEngland
research summit? Check out super awesome postdoc
@Alextremophile
's work at poster number 9. She has been systematically looking for 'near-coding' diagnoses in rare disease patients 🤩🌟
#GERS2022
2
6
61
Our UTRannotator tool paper is now online at Bioinformatics:
It annotates ~1/3rd of ClinVar (Likely) Path 5'UTR variants as potentially uORF-perturbing.
Thread from
@xiaolei_gene
on preprint👇(we have since added annotations for uSTOP-creating variants).
Want to identify regulatory variants disrupting protein translation with a large effect? With Matt Wakeling
@drjamesware
@nickywhiffin
, delighted to present UTRannotator - a VEP
@ensembl
plugin to identify high-impact 5'UTR variant perturbing upstream ORFs
2
36
97
0
15
60
I want to take a moment this morning to highlight four incredible women -
@ceclindgren
@hilsomartin
@GosiaTrynka
@Alexis_Barr
- we have never co-authored papers, and yet you have had a profound impact on me, and I wouldn't be where I am without your advice and support! 🥰
4
1
60
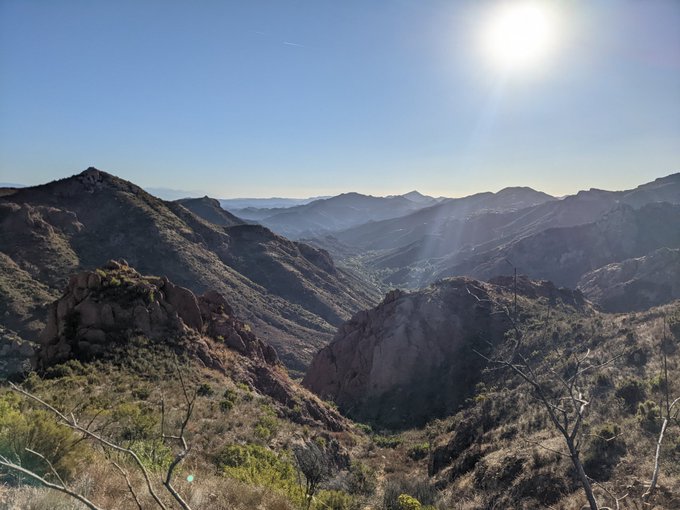
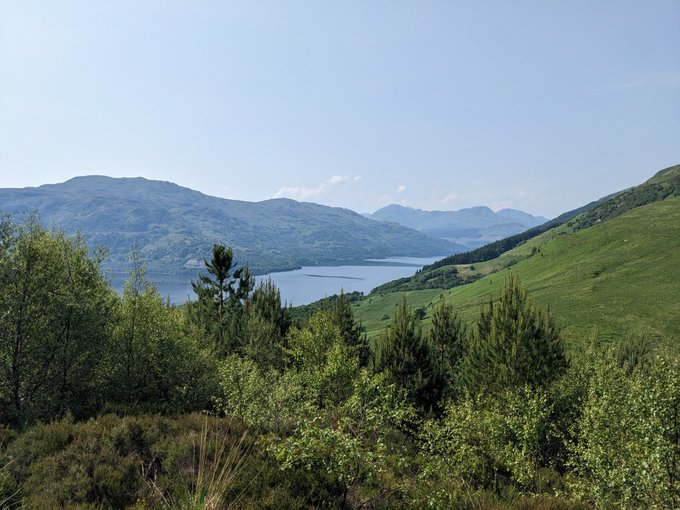
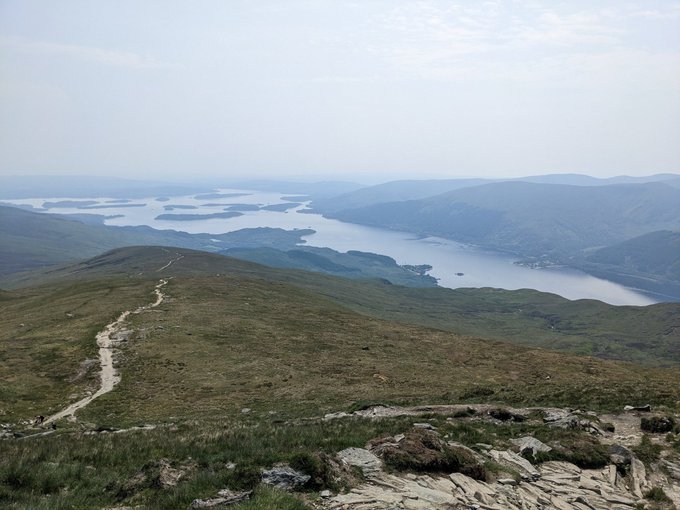
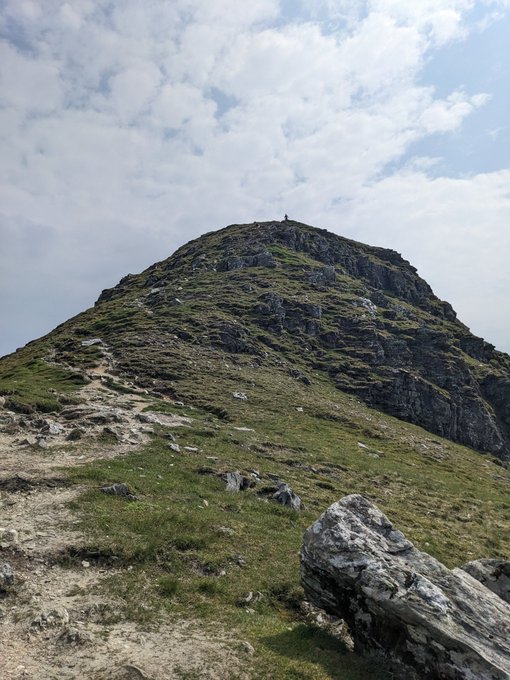
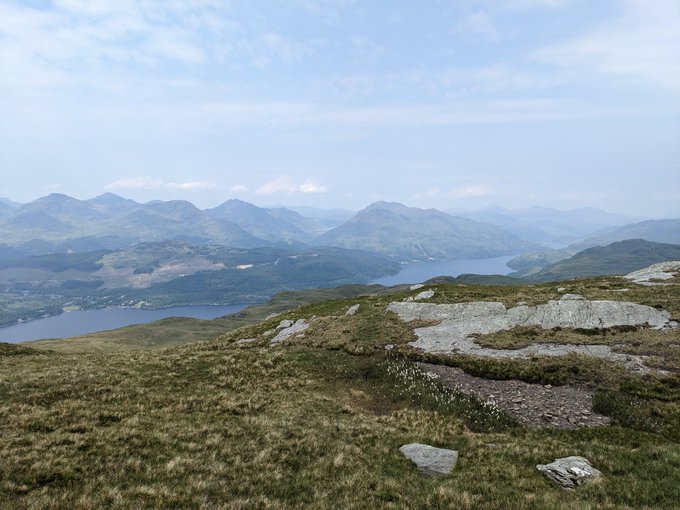
Fantastic time at
#ESHG2023
in Glasgow. Awesome science and great catch-ups, but there is never quite enough time (sorry if I missed you). I couldn't come all this way, however, without getting out into the surroundings and climbing a mountain. First Munro bagged: Ben Lomond ☀️⛰️
1
0
56
Love hearing about impact of
@GenomicsEngland
100k project!
Damian Smedley - 2,183 pilot probands - 26% diagnostic rate
Median length of diagnostic odyssey pre diagnosis = ~6yrs, 68 appointments!
24% diagnoses had immediate clinical utility
4% non-coding SNVs/indels
#ESHG2020
1
10
53
After multiple previous traumatic experiences (including completely losing my vision) I am pretty terrified of injections...
Today was a big day!
For anyone who may still be feeling hesitant - if I can do it, so can you!
#vaccinated
#doingMyBit
#feelingGrateful
0
1
52
📢 Looking for postdocs to lead an exciting new industry collaboration with
@joannahowson
@novonordisk
and an awesome team including
@ceclindgren
and
@carolinefwright
🌟
Early-career postdoc:
Senior role:
Please share and RT!
1/4
1
37
48
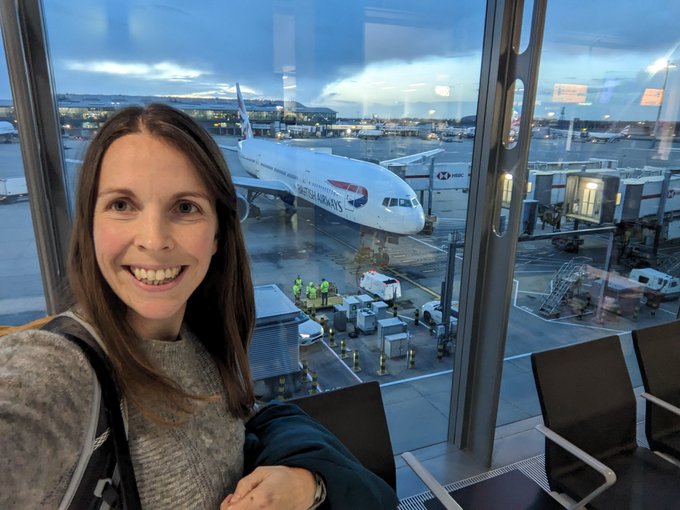
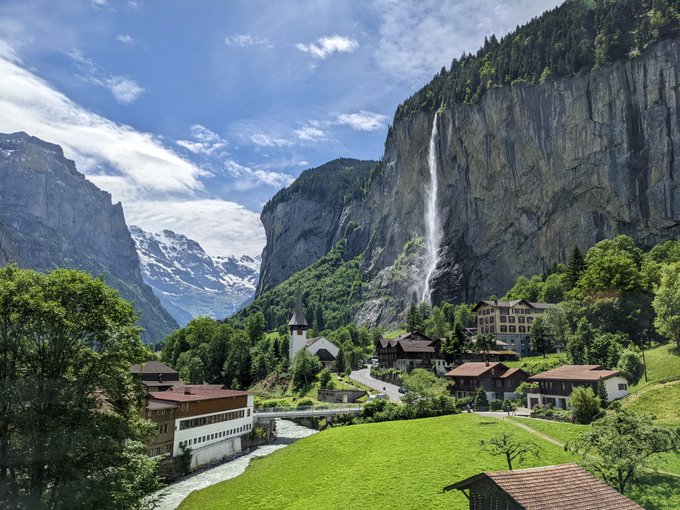
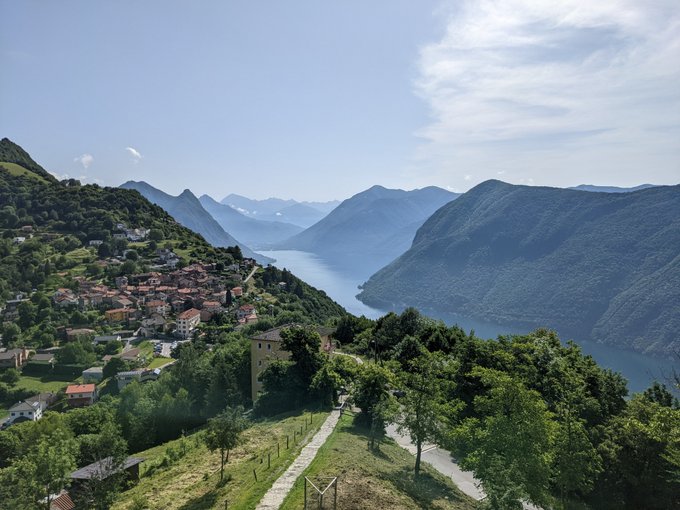
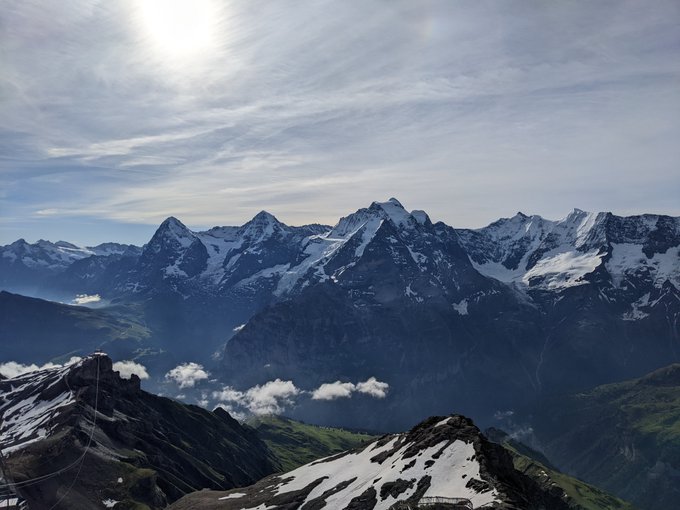
A trip first planned for May 2020, I finally got to experience the stunning views of Switzerland🇨🇭
Some stats:
17 days 📆
10 hotels 🏨
1 yurt 🛖
5 countries 🇨🇭🇫🇷🇮🇹🇦🇹🇱🇮
30 trains 🚆
7 cable cars 🚡
319,039 steps🚶♀️
Highest point: 2,970 m (Schilthorn) 🏔️
#mountains
#relaxation
5
0
49
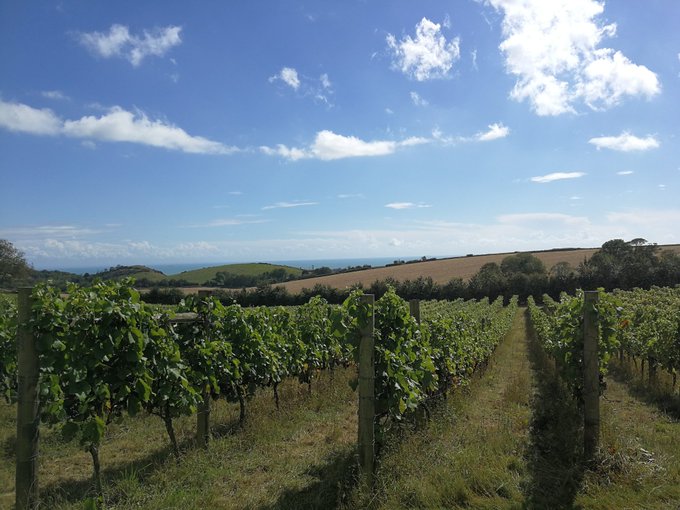
Second totally valid holiday interruption - to find out a paper has been accepted (our uORFs VEP plugin) - good job we picked up a lovely bottle of English sparkling wine whilst at
@Terlinghamwines
earlier today! Yes that is the sea, and if you look careful maybe a bit of France.
5
1
49
The number of people I have failed to recognise at
#ASHG23
is already embarrassingly high 🤦♀️ I am sorry everyone! A quick reminder that face-blindness (prosopagnosia) makes big events like this a struggle for many (~3-5% of people!). Please don't be offended ❤️
2
1
48
UTRannotator, our VEP plugin to identify and annotate uORF-perturbing variants is now 'finished'.
Find it here:
Paper here:
Please let us know any thoughts/suggestions
Full credit to superstar PhD student
@xiaolei_gene
🌟
Want to identify regulatory variants disrupting protein translation with a large effect? With Matt Wakeling
@drjamesware
@nickywhiffin
, delighted to present UTRannotator - a VEP
@ensembl
plugin to identify high-impact 5'UTR variant perturbing upstream ORFs
2
36
97
0
17
46
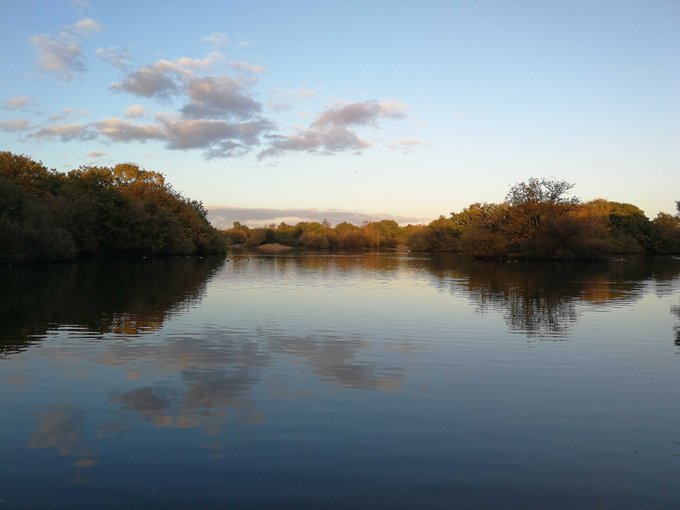
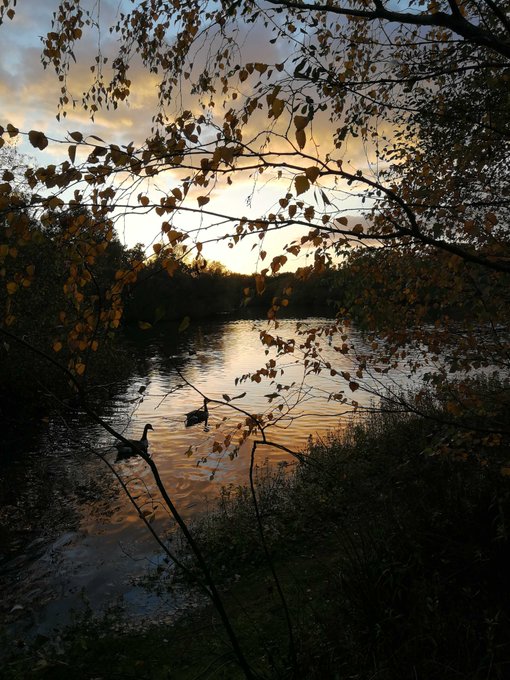
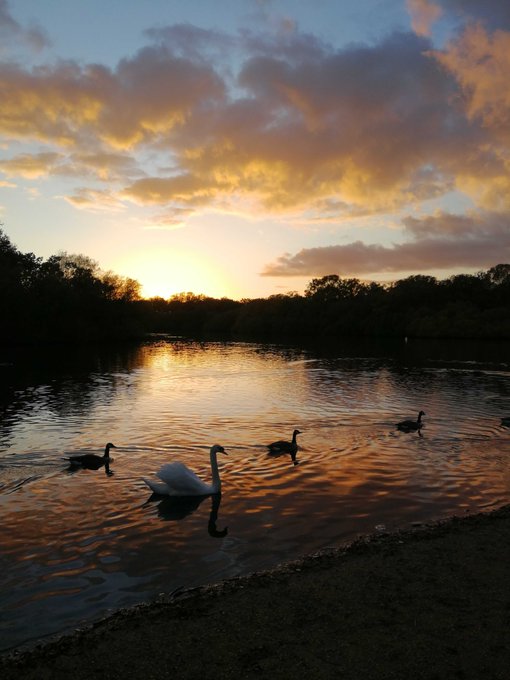
Interrupting your twitter feed with some calming pictures from my afternoon walk, just in case they are helpful
#autumncolours
#sunset
#reflections
#selfcare
1
2
47
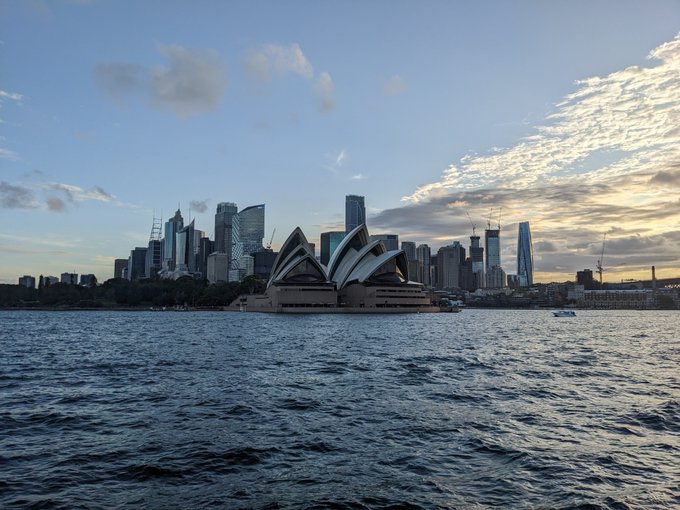
They _almost_ convinced me to stay... sad to leave beautiful Sydney behind (thank you for having me
@jodieingles27
and
@dgmacarthur
!) Now arrived in Hobart ready for
@GeneMappers
tomorrow. It is wet and windy, so here is sunny Sydney instead 👇
4
1
47
🚨 Crowd-sourcing input 🚨
The wonderfully talented
@elston_neil
has created a web-app to visualise 5'UTRs, uORFs, and variants within them. So far, our uber catchy name is "5'UTR Visualisation Application".
Please save us from ourselves and send any better suggestions!
13
14
46
Huge huge congratulations to the absolutely incredible
@GosiaTrynka
- winner of the Leena Peltonen Prize
#ESHG2020
for an outstanding young researcher in the field of human genetics 👏🌟
2
2
45
Last night we got to hang out with the dinosaurs whilst amazing DPhil student
@NechamaWieder
was awarded the
@NDMOxford
prize for "outstanding work outside of degree"! 🦖
Read about her incredibly inspiring community and charity work:
She is awesome!🤩
0
3
45
Time for my favourite conference of the year
#GRD22
Super representation from the Computational Rare Disease Genomics team - 1 talk and 3 posters 🙌
(and the first time I get to write one of these team conference Tweets 🥰)
Please come and chat to us!
0
5
44
New preprint from the team 🎉
Thread from incredible postdoc
@Alextremophile
who led this work below 👇
Thank you to all the team, collaborators, clinicians, and of course the
@GenomicsEngland
participants for making this happen.
As always, feedback very welcome!
1
14
42
Despite
@jtalbotmartin
being the first member of the team back in Sept 2020, today was the first time we met in person 😱😃 (he left for a yr & now works remotely)
Refreshed team photo with Jonny, super 🌟 new postdoc
@RuebenaEDawes
and awesome DPhil student
@eloise96wells
👇❤️
2
3
43
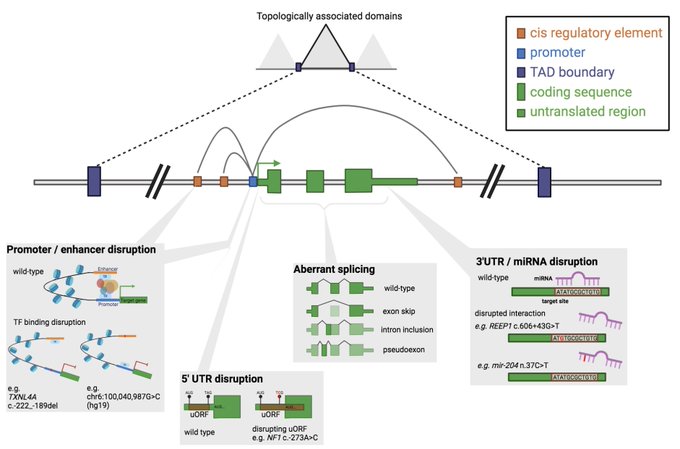
@j_ellingford
did create this awesome figure showing *some* of the non-coding region variant types we do know to cause disease 🌟 5/9
1
8
42
Updated team photo 🥰
So privileged to work with this awesome bunch💪💫
@NechamaWieder
would like me to point out the slope in the floor - we should have stood the opposite way around 🤣 I also thought we were going for stagger not line...
1
3
42
Today marks two years of the CRDG team
@HumanGeneticsOx
🎂
Thank you to all my amazing colleagues, mentors and obviously the incredible team for 2 yrs of support and awesomeness 💙
I meant to write a sum-up blog post of the year...but things got a little busy🙃
2
0
41
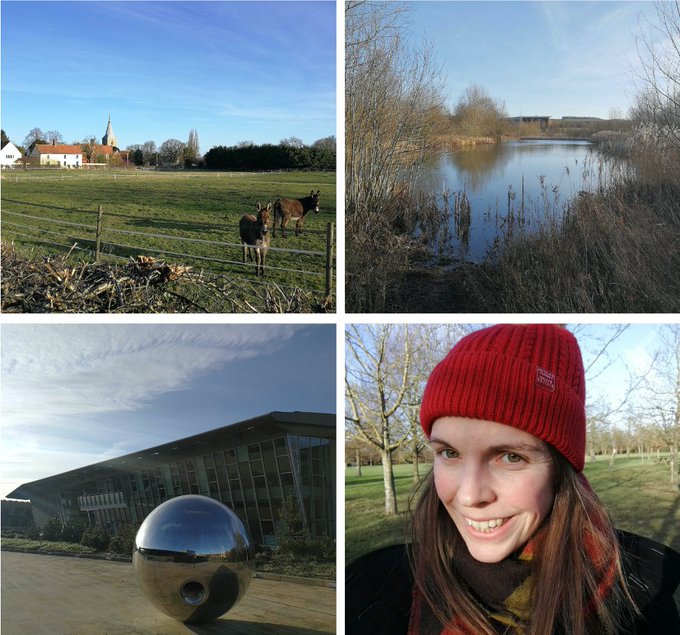
Since October I have been spending a couple of days a week at the
@sangerinstitute
hanging out with
@hilsomartin
,
@mehurles
and their awesome teams. It is mornings like this when I feel especially glad to escape London and head into the countryside 😍
3
2
41
For anyone with FOMO, the slides are all available on
Although, _huge_ caveat that these are very much a work in progress and all codes and points levels are draft and subject to change
#ESHG2023
0
8
40
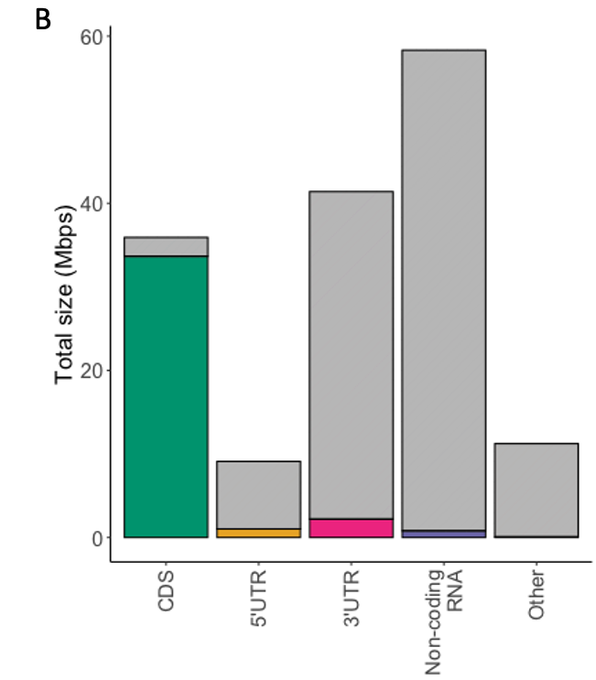
“Whole exome sequencing” is misleading and contributes to the misconception that exon = protein-coding
Only 24% of exonic bases are in capture of
@uk_biobank
exomes (89% of coding, but only 6.4% of UTR & 1.3% of ncRNA bases)
Let’s use coding exome sequencing (CES) instead!
7/8
2
6
38
Today marks three years since I moved to
@NDMOxford
@HumanGeneticsOx
and hence three years of the Computational Rare Genomics team! 🎉
A short 🧵 reflecting on some of the key changes and achievements over the last year: 1/9
3
0
39
Early start this morning. Heading into London for the
@GenomicsEngland
Research Summit. 🧬
Looking forward to catching up with many folks there!
0
1
39
It was an huge honour to introduce the simply incredible
@HeidiRehm
as the 2022
@GeneticsSociety
Curt Stern award winner 🤩🏆🍾 (if a bit cringe to watch it back).
Watch her amazing talk 👇
Congratulations again Heidi, you are truly deserving of this recognition 👏🌟
2
4
38
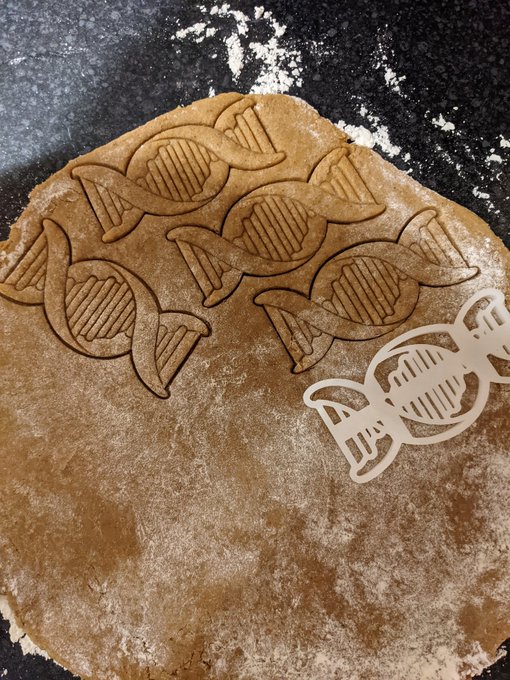
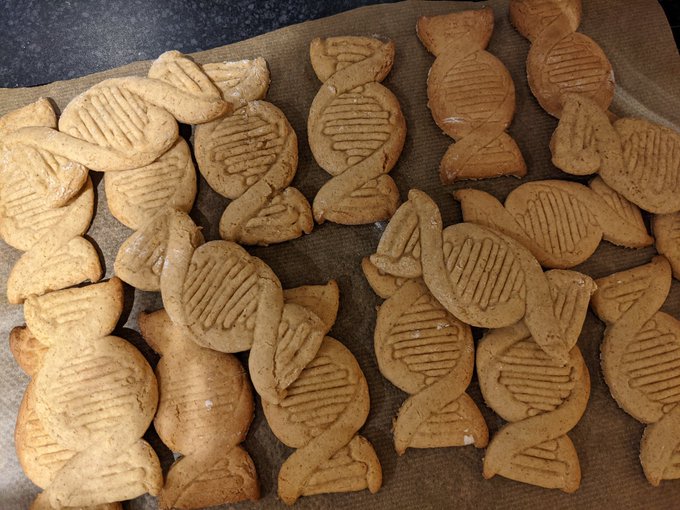
Quite pleased with how our creation for the local
#WorldBookDay2021
trail turned out 📚🤓🐲
#E17StoryWindows
3
1
38