amelie_schreiber
@amelie_iska
Followers
1,020
Following
343
Media
27
Statuses
568
I ❤️ proteins! Researching protein language models, equivariant transformers, LoRA, QLoRA, DDPMs, flow matching, etc. intersex=awesome😎✡️🏳️🌈🏳️⚧️💻🧬❤️🇮🇱
California
Joined May 2023
Don't wanna be here?
Send us removal request.
Explore trending content on Musk Viewer
Ayuso
• 184593 Tweets
No Riley
• 142264 Tweets
Sean
• 94056 Tweets
#NEDFRA
• 83451 Tweets
Jacob Juma
• 82697 Tweets
Austria
• 78694 Tweets
Francia
• 65112 Tweets
#LondonTSTheErastour
• 60223 Tweets
Olise
• 54646 Tweets
Bayern
• 49058 Tweets
Griezmann
• 39453 Tweets
Kanté
• 37588 Tweets
Rabiot
• 37243 Tweets
オランダ
• 30546 Tweets
فرنسا
• 30288 Tweets
Thuram
• 29193 Tweets
Holanda
• 26050 Tweets
Dembele
• 22724 Tweets
Clancy
• 22528 Tweets
Deschamps
• 21079 Tweets
The Dutch
• 18313 Tweets
DIAN
• 15037 Tweets
Barcola
• 14815 Tweets
THE BLACK DOG
• 13954 Tweets
HITS DIFFERENT
• 13886 Tweets
#PBSFRA
• 13291 Tweets
Depay
• 13145 Tweets
كانتي
• 13067 Tweets
Xavi Simons
• 10298 Tweets
Maroon
• 10282 Tweets
Last Seen Profiles
Found out yesterday some of my
@huggingface
blogs inspired some undergrads to start studying AI applied to proteins and someone applied to and received an internship based on their interest in replicating and extending some of them. 😎 Feeling very inspired and grateful now. ❤️
4
8
130
Here’s a new method for sampling the equilibrium Boltzmann distribution for proteins using GFlowNets:
If you aren’t familiar with GFlowNets, head over to
@edwardjhu
’s twitter and watch his video. I’ll also post a link to a related lecture soon.
3
4
40
Not specifically for proteins or other molecules, but this is a nice intro to flow matching. Thanks for the video
@ykilcher
any chance you’d ever do something on this applied to proteins?
0
8
38
Whenever an open source version of
#AlphaFold
3 is being created, be sure to try swapping out the diffusion module for a flow matching module. It’ll probably turn out better that way 😉
2
4
29
Let’s go!
“CRISPR-GPT leverages the reasoning ability of LLMs to facilitate the process of selecting CRISPR systems, designing guide RNAs, recommending cellular delivery methods, drafting protocols, and designing validation experiments to confirm editing outcomes.”
2
2
18
Apparently you can in fact do flow matching on discrete data, for those interested in diffusion applied to discrete data like language and NLP, this is a good reference for how to do it with the more general flow matching models:
0
1
17
Another E(3)-equivariant model that should be SE(3)-equivariant. E(3) doesn’t preserve chirality of molecules.
GitHub:
0
1
14
Love child from Distributional Graphormer (
#DiG
) and
#alphafold3
when? C’mon
@Microsoft
and
@GoogleDeepMind
. If
@OpenAI
and
@Apple
can team up to deliver
#Her
we can also have a new model that does dynamics for complexes of biomolecules. You’re almost there 🔥🔥🔥you got this.
3
2
13
Thank you for inviting me and for the wonderful conversation.
The AI Revolution in Biology is here - it's just not evenly distributed, even among biologists
@amelie_iska
previews biology as an experimental information science, on the latest Cognitive Revolution – out now!
Listen to catch up!
(link in thread)
2
4
22
2
2
11
RFDiffusion could’ve been used and it’s suggested this would improve outcomes. Why wasn’t it used for this problem? I’m curious. This would establish more use cases for RFDiffusion and similar methods (like flow matching with FoldFlow-2 for example).
Now online: We developed novel oligomers and turned them into FGFR agonists via binder induced receptor clustering.
#denovo_proteins
3
29
109
1
1
11
Seems like an interesting method. I find it very interesting that it works better (SOTA?) if you give it conformational ensembles to work with. Could be very interesting to see how conformational sampling, Distributional Graphormer, or AlphaFlow might yield better results.
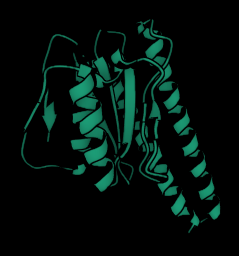
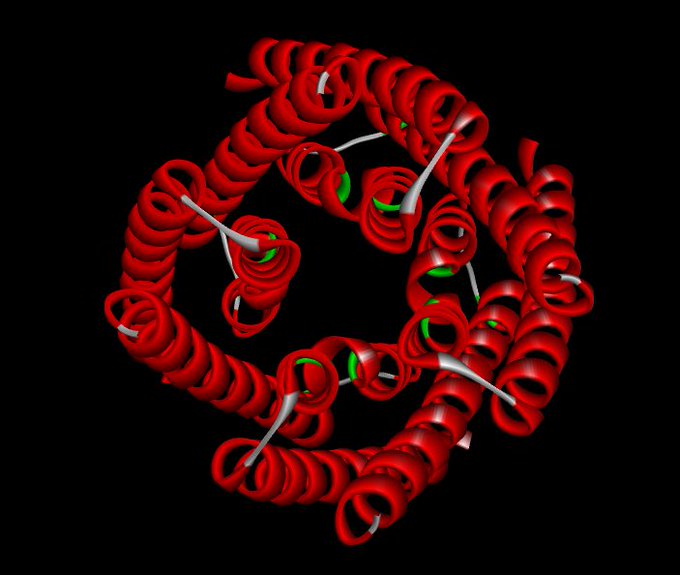
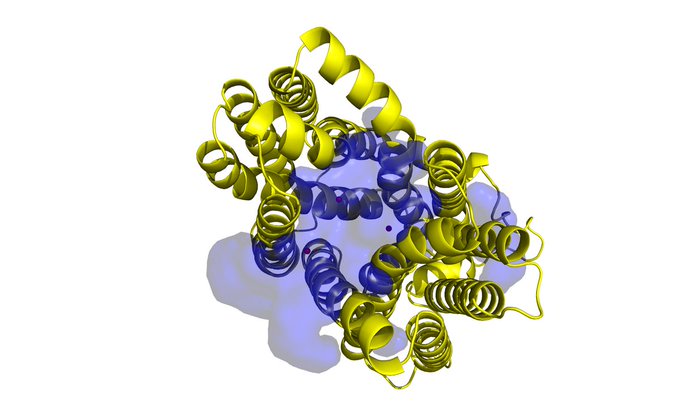
Having a lot of fun visualising the ligand binding site predictions of
#IFSitePred
with
#PyMol
! A new ligand binding site prediction method that uses
#ESMIF1
learnt representations to predict where ligands bind! Check it out here:
#Q96BI1
0
2
20
2
1
10
@TonyTheLion2500
I highly recommend this reference along with his “smooth manifolds” book: Introduction to Riemannian Manifolds (Graduate Texts in Mathematics)
0
0
10
@SimonDBarnett
Code is linked to in the Nature paper. This AI model actually samples the Boltzmann distribution, giving all the metastable states (low energy conformations) as well as the transition pathways between them. It’s a “generative diffusion model”:
0
1
8
And it has a BSD-3 license! Not bad.
0
2
7
@alexrives
I have a method for detecting AI generated proteins that I would like to open source at some point if people are interested. It seems to work on proteins generated by most models out right now, although there are a couple models it does not work for, hesitant to say which ones.
2
0
7
@samswoora
You should also check out flow matching models. Flow matching generalizes diffusion (diffusion is a special case of flow matching). They're doing a lot with proteins and flow matching, but there's less buzz about it in vision and language domains.
2
0
5
@maurice_weiler
@erikverlinde
@wellingmax
Could someone recommend a similar resource for other architectures like equivariant transformers or equivariance in geometric GNN models? Just curious what the go to resources are for people for other architectures.
2
0
4
@pratyusha_PS
This is awesome. When will the code be available? I would love to try this with a protein language model like ESM-2 and see if it improves performance.
2
0
5
@310ai__
It might also be good to look into computing the LIS score based on the PAE output of RoseTTAFold All Atom similar to what was done with AlphaFold-Multimeter here. This is a new approach for protein-small molecule complexes.
0
1
3
@HannesStaerk
@chaitjo
@SimMat20
@ADuvalinho
Just found this and it seems to address some of the concerns over computational cost of equivariant architectures:
0
0
3
Pretty neat. How does it compare to a contrastive model like ProteinDT or ProteinCLIP? And could we use it for annotating in order to train a new ProteinDT or ProteinCLIP? Is there an exceptional text-guided diffusion model coming soon for proteins?
Excited to share new work from
@GoogleDeepMind
: “ProtEx: A Retrieval-Augmented Approach for Protein Function Prediction”
3
41
153
1
0
3
@andrewwhite01
You can also learn equivariance. I think equivariance is an overrated mathematical concept tbh. It's fancy and neat from a mathematical perspective, but otherwise I think you could have your network learn it and get just as far if not further.
0
0
4
This is such a cute animation! 😎 Now do it for 3 modalities like ProTrek (text, protein sequence, protein structure)! 🧬
0
1
5
@befcorreia
@karla_mcastro
@_JosephWatson
@jueseph
@UWproteindesign
Curious to know why RFDiffusion motif scaffolding wasn’t tried here instead of or in addition to the RoseTTAFold constrained hallucination.
1
1
4
I think actually training this model could be done on Lambda Labs for around $150K (20 GPU days on 256 A100s) no? There is a difference between training and inference too that should be made clear here. Inference (using the model for predictions) is much cheaper than training.
1
2
4
@HannesStaerk
Still REALLY want to see this done with AlphaFold-Multimer. Maybe there’s a dynamic model of PAE and LIS that comes out of this that helps determine how strong or transient a PPI is.
1
2
4
@biorxiv_bioinfo
Cool idea, but how was the dataset split into train, test, and validation? Was sequence similarity/homology used to split the protein dataset? If not, this paper's results are unreliable. You have to split your data based on sequence similarity; 30% similarity is pretty standard
0
0
3
@KKapusniak1
@PPotaptchik
@TeoReu
@leoeleoleo1
@AlexanderTong7
@mmbronstein
@bose_joey
@Francesco_dgv
@FrankNoeBerlin
this maybe useful for transition pathways between conformations.
0
0
4
I love this. Thank you! Gotta go watch now! p(shittakes | Michael Levin) << p(shittakes | Amelie Schreiber) 😂 Also,
@drmichaellevin
…feel free to DM anytime with project ideas 🤓
1
0
4
@iScienceLuvr
SVD initialization would’ve helped a lot.
0
0
3
@ZymoSuperMan
RFDiffusion works with structures, not sequences. For designing sequences that fold into the backbones that RFDiffusion generates you’ll need something like LigandMPNN which does allow for things like biasing particular residues in various ways to constrain the sequences designed
1
0
3
After that, team up with
@BakerLaboratory
and make the best “RFDiffusion” and “LigandMPNN” anyone’s ever seen, but this time use continuous and discrete flow matching resp. and make it for all the biomolecules. 4 essential “foundation models” and we’ll be all set +/-ε 🎉😎🧬
0
0
3
@lpachter
For academic uses that don’t compete with Isomorphic Labs’ research… that part is subtle, but important. It means if you want to develop a new drug and have any hope of taking it to market, or if you’re not in academia, you’re out of luck. And no reproducing because patents! 😒
0
0
3
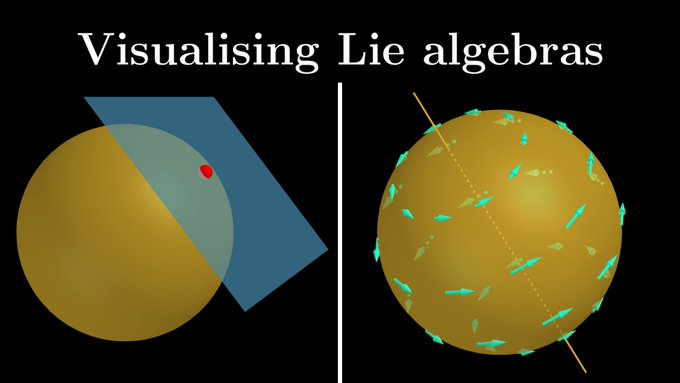
Really cool channel. Maybe we’ll get a video on SE(3)-equivariant neural networks one day🤞This would be great for folks trying to understand new SOTA models for proteins and small molecules. I would totally be down to collaborate
@mathemaniacyt
🧬
0
0
3