Frank Noe
@FrankNoeBerlin
Followers
9,714
Following
578
Media
266
Statuses
4,502
Scientist, #MachineLearning and #AI for the Sciences (esp. Physics/Chemistry). Scuba Diver and Traveler.
Berlin, Germany
Joined June 2017
Don't wanna be here?
Send us removal request.
Explore trending content on Musk Viewer
Northern Lights
• 240636 Tweets
オーロラ
• 202844 Tweets
DeNA
• 62036 Tweets
#withMUSIC
• 44840 Tweets
Fulham
• 32294 Tweets
BQUIK X BKPP
• 24972 Tweets
太陽フレアのせい
• 24350 Tweets
NUNEW AT BITEC
• 23656 Tweets
#العين_يوكوهاما
• 21491 Tweets
ベイスターズ
• 18910 Tweets
バチコン
• 18854 Tweets
ハマスタ
• 17752 Tweets
FOURTH SINGING TIME
• 15899 Tweets
マリノス
• 12705 Tweets
自分の魚
• 12465 Tweets
#マルハニチロ
• 12413 Tweets
GenG
• 12299 Tweets
#誕生魚診断
• 11706 Tweets
わたしの誕生魚
• 11570 Tweets
Buju
• 11454 Tweets
Last Seen Profiles
Tweeps - news! I'll join
@MSFTResearch
as research manager starting Oct 1st, opening a new AI4Science Lab in beautiful Berlin! We'll focus on fundamental challenges between
#MachineLearning
&
#Physics
/
#Chemistry
. We'll start at Alexanderplatz, and we'll hire at all levels! 🧵
56
76
867
I made my introduction to
#DeepLearning
lectures (lots of references to shallow
#MachineLearning
and
#Physics
) public.
Hopefully useful for newbies who want to get an overview of methods before specializing on state-of-the-art stuff. Feel free to use!
7
145
541
Wohoo - PauliNet, our deep
#NeuralNetwork
solution of the electronic Schrödinger equation is out in
@NatureChemistry
. Huge respect for
@jhrmnn
and Zeno Schätzle for championing this. Deep Quantum Monte Carlo is the future!
15
109
473
German future prize for Sahin, Türeci, Kariko and Huber for developing the first approved Covid-19 vaccine and bringing the m-RNA technology to scale with
@BioNTech_Group
. I couldn't think of anything more appropriate. Huge congrats and thank you!
#zukunftspreis
@kkariko
10
46
399
Excited to join
@MSFTResearch
on this journey. Big shoutout to Chris Bishop,
@sebnowozin
,
@wellingmax
,
@vdbergrianne
,
@mmjb86
,
@MaziarzKris
,
@marwinsegler
, Becky Frost and the whole team for being super welcoming and amazing.
Microsoft Research welcomes Visiting Researcher, Prof Frank Noé (
@FrankNoeBerlin
), a leading expert in machine learning for the natural sciences.
Learn more about opportunities across our global labs.
3
16
114
21
19
306
It's finally here: the DeepTime software library!!!
Massive work led by Moritz Hoffmann, with Martin Scherer,
@tmhmpl
,
@andreasmardt
, Brian de Silva,
@brookehus
, Stefan Klus, Hao Wu, Nathan Kutz and
@eigensteve
.
6
68
289
Let's celebrate this seminal paper by Katalin Karikó
@kkariko
. Describes how mRNA can be modified so that it doesn't cause an immune reaction. This is the basis for mRNA to be used in the Covid-19 vaccines of
@BioNTech_Group
,
@moderna_tx
and
@CureVacRNA
.
4
70
270
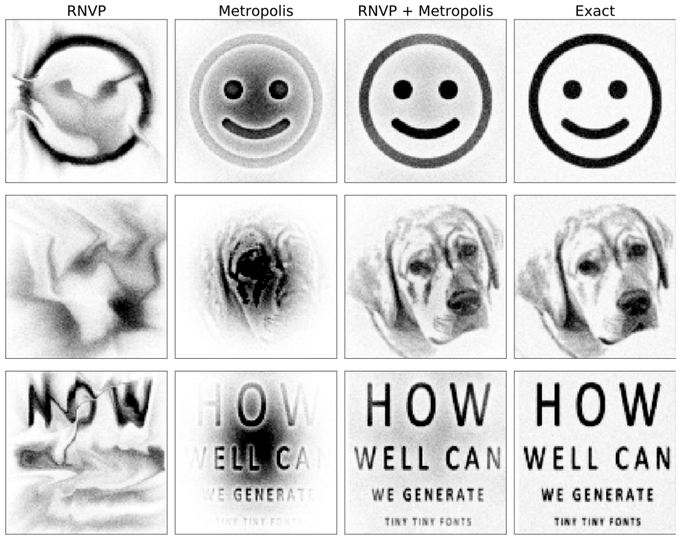
Stochastic Normalizing Flows: We can train normalizing flows by combining invertible networks and stochastic dynamics (MCMC, Langevin) in any order. Better expressivity and sampling than either one.
With
@jonkhler
& Hao Wu
#MachineLearning
#DeepLearning
7
51
261
Unbelievable to be elected to the Berlin-Brandenburg academy of sciences with these outstanding folks. Founded w/ Leibniz in 1700, Einstein revealed general relativity here in 1915. Now in good company with
@GroupParrinello
@c_drosten
@e__charpentier
...🤯
@bbaw_de
@FU_Berlin
Wir begrüßen unsere neuen
#Akademiemitglied
|er: Antje Boetius
@AWI_de
, Christian Drosten
@ChariteBerlin
, Eva Geulen
@zflkomm
, Volker Markl
@TUBerlin
/
@DFKI
, Frank Noé
@FU_Berlin
, Dagmar Schäfer
@MPIWG
, Matthias Warstat
@FU_Berlin
#Einsteintag
0
2
39
38
6
226
15 PhD positions available on
#MachineLearning
for
#drugdiscovery
in EU grad school. Apply to project ESR15 to work with me at
@FU_Berlin
/ Germany and Ola Engkvist
@AstraZeneca
/ Sweden on deep learning for protein simulation.
4
104
219
Who wants to come to Berlin and work with
@jhrmnn
and myself on the next generation of deep learning systems for Quantum Chemistry, especially deep QMC, exploiting ideas from Stat Mech, generative deep learning, equivariant graph nets etc? Send Email or DM.
9
64
184
Learning coarse-grained molecular dynamics with graph neural networks. Led by
@brookehus
, Nick Charron and
@CecClementi
2
36
182
Just read this. Beautiful and elegant method for computing chemical potentials by
@ChengBingqing
. This is the first time that a grand-canonical "simulation method" makes sense to me - by simply avoiding to do the simulation grand-canonical at all.
2
34
179
Proud of our 2019
#MachineLearning
work. Highlights:
#BoltzmannGenerators
: Unbiased sampling of target distributions
PauliNet: Solving electronic Schrödinger equation
CGnet: Learn coarse-grained MD
1
38
161
2020 won't be less exciting. My personal/private highlights:
-
@CecClementi
will move to Berlin as an Einstein professor for physics
- We are buying an apartment in Schöneberg
- We are getting married!!
20
0
149
Antisymmetry is what makes electronic structure calculations hard. We need more
#MachineLearning
work to make progress with this fundamental problem. Stefan Klus takes a stab at it by developing antisymmetric kernels:
5
24
143
We propose a new+data-efficient way for
#MachineLearning
coarse-grained molecular force fields from simulation data (e.g. all-atom). With
@jonkhler
,
@__kraemer__
, Yaoyi Chen,
@CecClementi
.
@bifoldberlin
@MSFTResearch
@ERC_Research
@FU_Berlin
6
32
138
Online workshop on protein dynamics coming up April 26-28. Awesome speakers including
@RommieAmaro
@LindorffLarsen
@gdefabritiis
@CarloCamilloni
@CecClementi
and many folks not on twitter ;-) Registration required.
3
53
136
A bright moment in dire times.
@CecClementi
being appointed Einstein professor for physics in Berlin by FU president Günter Ziegler (with safety distance).
@Einstein_Berlin
,
@FU_Berlin
.
15
2
133
Internship opportunity at
@MSFTResearch
#AI4Science
on generative protein models. Location Berlin, Amsterdam or Cambridge.
Apply here asap:
4
45
128
First official release of the DeepQMC code for Quantum Monte Carlo with
#NeuralNetworks
wave functions, including architectures like PauliNet, FermiNet and DeepErwin. By Zeno Schätzle,
@PBerntSzab1
, Matej Mezera and
@jhrmnn
1
36
126
#MachineLearning
coarse-grained molecular dynamics with flow-matching is now out in JCTC
@JCIM_JCTC
!
@jonkhler
Yaoyi Chen (not on Twitter?)
@__kraemer__
@CecClementi
.
1
14
124
Tweeps at
#NeurIPS2022
#ML4PS2022
@lrml_bio
@AI_for_Science
looking for exciting jobs in deep learning and molecular sciences?
@MSFTResearch
AI4Science is hiring in Berlin and other sites. DM me or approach my colleagues for more info.
0
38
121
LinkedIn keeps suggesting I should apply for our jobs at
@MSFTResearch
AI4Science Berlin, and I am tempted to…
4
1
119
2006 PhD
2012 Scuba diver
2013 Professor
2021 Married love of my life
2022 First real job
Seems like a weird order I know
3
2
115
Microsoft research announces AI4Science initiative focusing on next generation
#MachineLearning
for natural sciences. Super exciting!
1
17
113
I have a proposal. How about removing any occurrences of
#MachineLearning
and
#ArtificialIntelligence
from paper and workshop names as it’s trivially an ingredient in pretty much everything now?
8
5
112
Tweeps - apply here for senior researcher and engineering positions in
@MSFTResearch
AI4Science Berlin! We'll keep these open for a few more weeks.
If you have already applied please give us some time to respond.
Tweeps - news! I'll join
@MSFTResearch
as research manager starting Oct 1st, opening a new AI4Science Lab in beautiful Berlin! We'll focus on fundamental challenges between
#MachineLearning
&
#Physics
/
#Chemistry
. We'll start at Alexanderplatz, and we'll hire at all levels! 🧵
56
76
867
6
42
106
Awesome
@CecClementi
giving a honorary speech in the Italian embassy Berlin on the occasion of the international women's day. 💪👩🔬❤💥
6
4
106
I am so happy to see this. A big round of applause to the team for this heroic effort!!!
🚨ANNOUNCEMENT🚨
The
@covid_moonshot
is now published in
@ScienceMagazine
! The paper reflects the major contributions of the FAH community and a global network of scientists collaborating to make drug discovery open, globally accessible, and affordable
1
88
237
1
11
106
Exciting: excited PauliNet - computing excited states of electronic Schrödinger equation w. deep neural nets + Quantum MC. By M. Entwistle, Z. Schätzle, P. Erdman and
@jhrmnn
#MachineLearning
#quantum
@FU_Berlin
@bifoldberlin
@MSFTResearch
@ERC_Research
4
15
105
We are setting up an
#AI
/
#MachineLearning
-driven protein design platform. Looking for a highly motivated PostDoc with experience in protein purification and/or RNA biology. Ideal: experience with experimental protein design. Collab with Daumke (MDC) and Schacherl (Charite).
3
33
102
Cool paper on transition-path sampling with Boltzmann-Generator moves by Michael Plainer,
@guennemann
@_bunnech
@HannesStaerk
:
1
18
99
Woohoo, Stochastic Normalizing Flows coming up as a
#NeurIPS2020
spotlight talk. Thanks 10^6 to Hao Wu and
@jonkhler
.
P.S. Arxiv version is identical in methods and results. That version was rejected from ICML. Except a few improvements for camera ready!
Stochastic Normalizing Flows: We can train normalizing flows by combining invertible networks and stochastic dynamics (MCMC, Langevin) in any order. Better expressivity and sampling than either one.
With
@jonkhler
& Hao Wu
#MachineLearning
#DeepLearning
7
51
261
3
12
100
Just out in
@NatureComms
: Excited states calculation with deep quantum Monte Carlo by Mike Entwistle, Zeno Schätzle,
@PBerntSzab1
,
@jhrmnn
.
#MachineLearning
@MSFTResearch
@FU_Berlin
@ERC_Research
@dfg_public
@MATHplusBerlin
@bifoldberlin
1
21
99
Excited: first joint paper with Klaus-Robert Müller, Alex Tkatchenko and
@CecClementi
, coming up as an Annual Review in Physical Chemistry.
First strike in our efforts to unite Machine Learning for Quantum and Statistical Mechanics
1
26
97
Roadmap on
#MachineLearning
in Electronic Structure: Perspectives on current + future challenges on predicting material properties, learning force-fields, solution of the many-body problem etc.
0
23
94
Pls Share: Wanna do a
#MachineLearning
PhD in Europe? Any core ML discipline or ML + Phys / Chem / Bio?
Apply to
@ELLISforEurope
here by Dec 1st. Work with two top Labs/PIs of your choice.
My group has openings, please contact for more info.
3
78
96
Our deep learning approach to identify independent Markov domains in molecules is out in
@NatureComms
.
@andreasmardt
@tmhmpl
@CecClementi
1
19
96
Working towards computing kinetics for large biomolecules with independent Markov decomposition.
@tmhmpl
championed this work, big shoutout to the team Mauricio del Razo,
@CTLeeRes
@RommieAmaro
@biobryn
.
So happy to have a paper with you Rommie!
7
23
93
Join us for on a wonderful postdoc position at
@MSFTResearch
AI4Science Berlin on
#MachineLearning
for Molecules!
0
31
93
This one was very tough, and it's finally out! In-depth analysis of ubiquitin-SH3 protein-protein binding mechanism combining extensive MD, Markov modeling and NMR with
@smnlssn
, Kalyan Chakrabarti, Christian Griesinger, Thomas Weikl and many colleagues
3
13
89
Welcoming postdoc openings for
#MachineLearning
in Statistical Physics and/or
#quantum
Physics / Chemistry. Pls Email frank.noe
@fu
-berlin.de or DM with CV+publications+1-2 refs.
#postdocposition
#AcademicTwitter
@jhrmnn
@CecClementi
@bifoldberlin
@FU_Berlin
2
37
88
Code for PauliNet - deep neural network solution of the electronic Schrödinger equation in open-source deepqmc👇
Paper: , Talk:
#MachineLearning
@ELLISforEurope
@jhrmnn
#QuantumMechanics
@MATHplusBerlin
0
25
84
As someone loving
#MachineLearning
and Physics/Chemistry, I am overwhelmed by
#NeurIPS2020
having 5 exciting workshops on this interface. We are not writing papers that fast...
1
4
84
Opening for PhD positions in
#MachineLearning
#ArtificialInteligence
at ELIZA/
@ELLISforEurope
Berlin. Various research and application areas are possible :-) Apply by Dec 8.
3
37
82
@andreasmardt
&
@tmhmpl
have developed the
#MachineLearning
approach iVAMPnet: decompose macromolecules into dynamically independent subunits & describe each by a Markov State Model. With
@CecClementi
. 🧵⬇️
1
18
82
This
@ipam_ucla
workshop on learning and emergence in molecular systems was really spectacular and a lot of fun!
0
6
82
New work led by Yaoyi Chen:
#MachineLearning
implicit solvation for molecular dynamics using the coarse-graining principle and CGnet.
2
21
81
Fantastic work by my
@MSFTResearch
AI4Science colleagues.
#GenerativeAI
for Materials is going to be a gamechanger, and this is only the beginning.
[1/N] Generative AI has revolutionized how we create text and images. How about designing novel materials? We at
@MSFTResearch
#AI4Science
are thrilled to announce MatterGen: our generative model that enables broad property-guided materials design.
👇
27
225
902
1
10
81
Wonderful work led by
@MohsenSadeghi
on the coarse-grained simulation of biomembranes with incorporating hydrodynamics, realistic kinetics and long timesteps. This is meant to be used in particle-based reaction-diffusion simulations of cells:
2
22
78
Excited to share our approach for
#machinelearning
path-integral molecular dynamics to account for quantum nuclear effects in spectra and other observables. Fantastic team led by
@venkatkapil24
@CecClementi
@FelixMusil
.
An attempt towards
#pathintegral
quantum dynamics via classical molecular dynamics with
#machinelearning
and coase-graining alla
@FelixMusil
@CecClementi
@FrankNoeBerlin
. Feedback welcome. :) Thanks
@snsf_ch
@ChemCambridge
@ChurchillCol
1
14
90
1
13
79
Interested in a PhD on the interface of
#machinelearning
and biology/biophysics/comp chem? This is a good opportunity and allows us to admit both students with Bachelor and Master degrees on separate tracks:
0
29
77
@leonklein26
introduces flow matching with equivariance. This allows us for the first time to get flows working for iid sampling of molecular structures in Cartesian coordinates in such a way that we can get reasonable Monte Carlo acceptance rates.
0
16
78
1/ IMHO
@nature
's guided OA pilot is misguided. While it's optional now, a scenario in which researchers need to pay upfront to get reviews is unethical. Don't go there
@NaturePhysics
@NatureComms
@NatureChemistry
1
19
77
@adad8m
On the risk of getting yelled at, but I would do this in <5 min with PowerPoint. Takes some practice but is super efficient for these things when you know how.
3
0
75
Our paper on discovery of a new state across the bromodomain superfamily is out in
@PNASNews
work led by Lluis Raich and
@smnlssn
, super grateful to the fantastic team and support of
@ClaraDChrist
@Bayer
@MATHplusBerlin
@FU_Berlin
1
7
75
Happy to share our MLST paper on symmetric and antisymmetric kernels for
#MachineLearning
problems in physics and chemistry. Just out
@bifoldberlin
@MATHplusBerlin
@FU_Berlin
@ERC_Research
'Symmetric and antisymmetric kernels for
#machinelearning
problems in
#quantum
#physics
and
#chemistry
' by
@FrankNoeBerlin
S Klus, P Gelß and F Nüske
@FU_Berlin
@UniOfSurrey
@unipb
@RiceUniversity
hits 500 downloads already!
#compchem
#opendata
#compphys
1
3
21
1
10
73
Immense pleasure to having worked with this team and finally see this out. Using
#MachineLearning
, the first CG forcefield that predicts free energy landscapes similar to all-atom for unseen proteins, and it's much faster than all-atom. Tour-de-force by
@CecClementi
+team
0
3
72
Very happy and grateful that Chemistry at
@RiceUniversity
has renewed my adjunct appointment. Looking forward to come back and visit when the Covid situation allows us.
0
0
71
Ho ho ho, now I a have a machine ... actually it's a camera :-) Thanks to
@CecClementi
I have a wonderful new scuba toy - can't wait to try it out 🐠🐟🐡🐬🦈📸
10
0
70
Welcome new lab mate
@SherryLixueC
joining today as a researcher in
@MSFTResearch
AI4Science. Looking forward to working with you on the frontier of AI for quantum Chemistry.
2
2
68
Most downloaded articles in Annual Reviews 2020 includes our review on ML for molecular simulation with
@CecClementi
, Aleks Tkatchenko and Klaus Müller. It's free for download temporarily here (also on arXiv):
@AnnualReviews
@bifoldberlin
2
9
68
Marvelous day with the
@CecClementi
and Noe groups and we managed to sway the weather into admitting a BBQ. A few people were missing but now we have a nearly complete group shot that is photoshoppable to perfection.
0
4
68
Excited to speak at the AI4Science kickoff workshop of our
@ELLISforEurope
colleagues in Amsterdam tomorrow. Event starts at 9am CET:
@wellingmax
,
@BerndEnsing
,
@marcelworring
,
@KyleCranmer
,
@gdefabritiis
,
@cosmo_shirley
,
#MachineLearning
2
20
68
Opening at
@FU_Berlin
for a three-year postdoc position for
#MachineLearning
in the molecular sciences. Candidates with strong theory and method development knowledge in Quantum Chemistry or Statistical Mechanics are particularly encouraged to apply.
0
26
65
We are building a sklearn-style
#MachineLearning
framework for dynamical models from time series data: MSMs, (E)DMD, VAMP, Sindy, kernel&deep versions of those. Led by M. Hoffmann w.
@eigensteve
,
@brookehus
,N. Kutz,S. Klus & everyone invited!! What's your favorite package name?
sktime / scikit-time
80
skdyn / scikit-dyn
125
dynamo (dynamical model)
95
cyberdyne
49
17
13
65
New version of excited PauliNet -- a deep Quantum Monte Carlo approach to compute ground and excited states of the electronic Schrödinger equation is online. Now including conical intersections!
Exciting: excited PauliNet - computing excited states of electronic Schrödinger equation w. deep neural nets + Quantum MC. By M. Entwistle, Z. Schätzle, P. Erdman and
@jhrmnn
#MachineLearning
#quantum
@FU_Berlin
@bifoldberlin
@MSFTResearch
@ERC_Research
4
15
105
1
9
63