Martin Mayta
@MartinMayta2
Followers
214
Following
5K
Media
17
Statuses
3K
BSc Biotechnology - PhD in Biological Sciences. Teaching assistant professor at @UNRoficial & @UAPArgentina (Argentina)🇦🇷.
Joined October 2020
Nice tool!!.
Exciting to see our protein binder design pipeline BindCraft published in its final form in @Nature ! This has been an amazing collaborative effort with Lennart, @csche11h, @sokrypton, @befcorreia and many other amazing lab members and collaborators.
0
0
1
wow is this the next Nobel prize??Only time will tell. meanwhile amazing discovery!! Serendipities never end in science.
FROM CELL DEATH TO REGENERATION: What if dying cells -come back to life? Now online @embojournal I , we aim to answer this question. Here is the journey. If you🩷science, please dont 🛑till the end of 🧵, I promise you will not regret. 1/n
1
0
7
these proteins never dissapoint!.
A new paper in Cell shows that it is possible (simple, really) to convert DNA-editing CRISPR proteins, such as Cas9, into RNA editors. All you have to do is delete a chunk of them. Deleting 55 amino acids from a DNA editor called IscB (the ancestor of Cas9) turned it into a
0
0
4
Say what??.
Is it time to speak bout a "sixth sense" arising in the gut?. Scientists have identified a "neurobiotic sense" that enables the host to adjust its behaviour in response to a molecular pattern from its resident microorganisms. @nature.
0
0
0
Cool!.
Out Now! Aggresomes protect mRNA under stress in Escherichia coli #mRNA #EscherichiaColi #Aggresomes
0
0
0
RT @BiologyAIDaily: DynaRepo: The repository of macromolecular conformational dynamics. 1. DynaRepo is a novel repository that addresses th….
0
33
0
Very cool!.
Where are all the de novo designed therapeutics? In a guest post by @frdreyer, we break down what the actual bottlenecks, challenges, and opportunities are in building and deploying purely in silico drug discovery engines, with a focus on antibody structure. Link below 👇
0
0
1
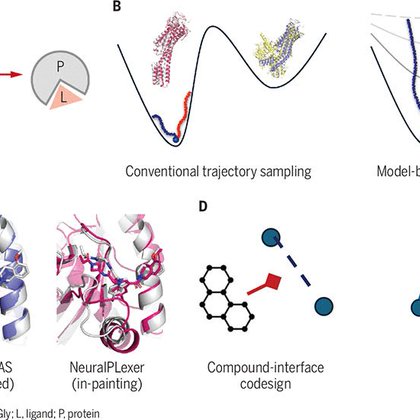
Modelling the landscape of protein-ligand binding with generative diffusion.
science.org
Structural foundation models help to elucidate and reprogram molecular biology
0
0
0
Cool! In silico affinity maturation of mini binders.
Massively Parallel Free Energy Calculations for In Silico Affinity Maturation of Designed Miniproteins @JCIM_JCTC . 1. Researchers from Temple University have developed a method for in silico affinity maturation of designed miniproteins using massively parallel free energy
0
0
0
Wow so cool!!.
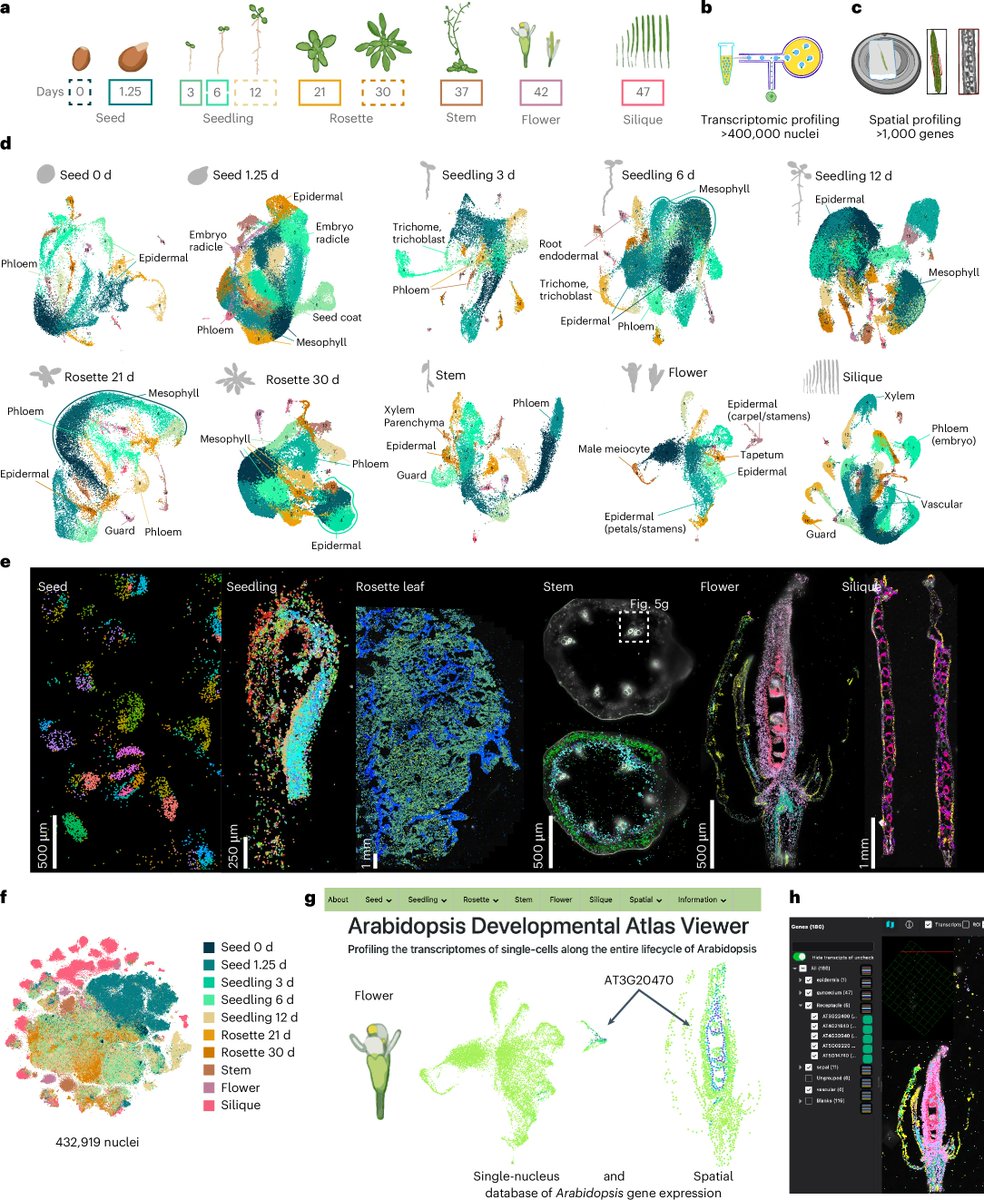
🌹Meet #ROSIE, our new AI that predicts spatial multi-protein expression straight from routine H&E images. Built on a large co‑staining dataset (>1k samples, 16M+ cells). It enables cell phenotyping & tissue structure insights w/o extra assays.
0
0
0