Vasilis Ntranos
@vntranos
Followers
791
Following
218
Media
12
Statuses
118
Assistant Professor @UCSF; Computational Biology and Information Theory
San Francisco, CA
Joined October 2012
We are hiring! Looking for a #postdoc to lead a new and exciting research direction we are taking in the lab https://t.co/ogpfq9h4RB
@UCSF, in collaboration with the #DataScience team at Maze Therapeutics ( https://t.co/prQckDZC2X) See here for more: https://t.co/OhBGCHWEAU 1/
1
17
16
For the first (and probably last) time in my life I understand the technical details of both the physics and chemistry Nobel prizes.
BREAKING NEWS The Royal Swedish Academy of Sciences has decided to award the 2024 #NobelPrize in Chemistry with one half to David Baker “for computational protein design” and the other half jointly to Demis Hassabis and John M. Jumper “for protein structure prediction.”
12
64
1K
🎉Dr. James Gardner has received the prestigious W.M. Keck Foundation Award for 1.3M for his research on extrathymic Aire-expressing cells, potentially impacting a range of conditions from type 1 diabetes to cancer immunotherapy to organ transplantation👉 https://t.co/Ud1oKFlt5V🌟
2
6
48
We have trained ESM3 and we're excited to introduce EvolutionaryScale. ESM3 is a generative language model for programming biology. In experiments, we found ESM3 can simulate 500M years of evolution to generate new fluorescent proteins. Read more: https://t.co/iAC3lkj0iV
139
809
3K
"Where do I think the next amazing revolution is going to come? And this is going to be flat out one of the biggest ones ever. There's no question that digital biology is going to be it." Jensen Huang, founder & CEO of NVIDIA. https://t.co/7j4WrHKPok
30
177
883
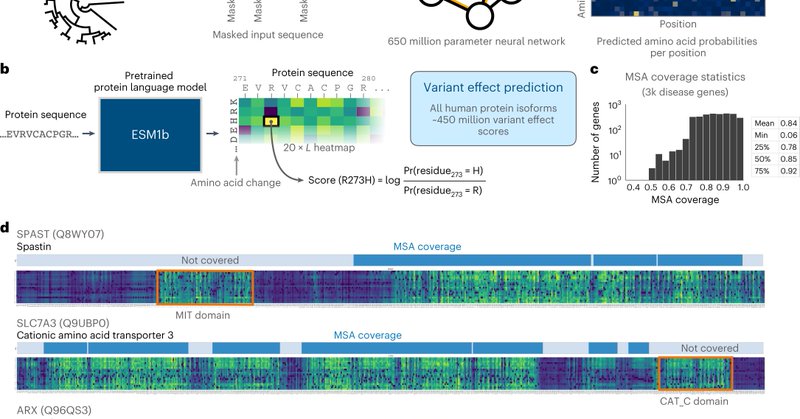
A protein language model predicts the effects of 450 million missense variants across >40,000 protein isoforms! Very cool work from @BrandesNadav @vntranos @yimmieg in @NatureGenet
7
73
337
@yimmieg @BrandesNadav @alexrives @MetaAI @Meta Finally, as a new PI, I want to say that I am very impressed with the high quality peer review process at @NatureGenet. The editors and reviewers provided amazing feedback and the paper almost doubled in content since our first submission a year ago: https://t.co/HdQ0EmiuMC 5/n
Our new preprint on genome-wide prediction of disease variants with protein language models: https://t.co/qK4OdodQu6
1
0
6
@yimmieg @BrandesNadav @alexrives @MetaAI Although I was disappointed with the recent news that @Meta decided to disband their protein team, it was great to hear from @AlexRives that the ESM team is trying to find a way to continue this line of research https://t.co/RyIRtWG5Qj 4/n
@KevinKaichuang Thanks Kevin. A group of us from the ESM team are committed to this research and are taking steps to make sure it will continue. It has been incredible to see all the ways the community has been using ESM and we think continuing to develop it is important.
1
1
9
@yimmieg @BrandesNadav Our approach uses ESM1b, a powerful protein language model developed by @AlexRives @MetaAI and among others we provide a complete online catalog of all ~450M possible missense variant effects across all human protein isoforms at: https://t.co/DRDP74ZyoO 3/n
huggingface.co
1
1
8
This has been an amazing collaboration with @yimmieg and superstar postdoc @BrandesNadav -- who by the way is on the market this year! See his thread for more details on our work: https://t.co/ydeYraBu7X 2/n
Protein language models can predict the clinical consequences of mutations, and they apply for any possible coding mutation. Even though they aren’t trained to do that. We show that in our latest Nature Genetics paper: https://t.co/aVUfoc8XjG 🧵👇
1
0
8
Excited that our paper is now published in Nature Genetics: https://t.co/C5dvCXoKeP Overall we show that protein language models are an effective, accurate and general approach to predicting variant effects. 1/n
nature.com
Nature Genetics - A modified framework leveraging a protein language model (ESM1b) is used to predict all possible 450 million missense variant effects in the human genome and shows potential for...
5
53
233
New version for those who don't have access to #GPT4. Just leave the api-key field blank in settings and try it out. We'll try to keep this going for a few days.
github.com
Contribute to ntranoslab/celltypewriter development by creating an account on GitHub.
0
0
3
👉UPDATE: You can now use #CellTypeWriter without a #GPT4 api-key. We've set up a #free version so you can try it out on your #data. ✨This will be active for a few days or until funds run out : ) Have fun! Github link:
github.com
Contribute to ntranoslab/celltypewriter development by creating an account on GitHub.
Introducing #CellTypeWriter, a user-friendly tool to explore and analyze #scRNAseq data using an interactive chat interface powered by #GPT4. Check it out on github: https://t.co/6grqsez26H
0
1
9
I've been exploring #GPT models' capabilities in #singlecell analysis for a while now, and they've all struggled with #scanpy. Although #ChatGPT-3.5 showed promise, it fell short in grasping the complexities of single cell RNA-seq data. Enter #GPT4. 👇
Introducing #CellTypeWriter, a user-friendly tool to explore and analyze #scRNAseq data using an interactive chat interface powered by #GPT4. Check it out on github: https://t.co/6grqsez26H
0
3
8
Follow the instructions on #github to install it locally and try it out! Let us know what you think.
github.com
Contribute to ntranoslab/celltypewriter development by creating an account on GitHub.
0
0
5
Note: CellTypeWriter is an experimental *proof-of-principle* project employing GPT-4 for single cell analysis. AI-generated code requires caution & due diligence for research purposes. Use responsibly.
1
0
2
We all make mistakes. #CellTypeWriter will try to execute the #GPT4 generated code and self-correct any errors it encounters. In the end, it will provide an explanation of the issues encountered and how they were resolved.
1
0
2
#CellTypeWriter is designed to recall its own output and maintain context throughout the entire conversation thread. This allows for a more coherent and efficient exploration of #singlecell data.
1
1
2
Introducing #CellTypeWriter, a user-friendly tool to explore and analyze #scRNAseq data using an interactive chat interface powered by #GPT4. Check it out on github: https://t.co/6grqsez26H
3
14
102