Jesper V. Olsen
@jespervolsen
Followers
1K
Following
1K
Media
3
Statuses
32
Joined July 2016
From HeLa to human blood: extending single-cell proteomics to the smallest immune cells https://t.co/S8zPv9fXGU
#biorxiv_cellbio
biorxiv.org
Single-cell proteomics (SCP) holds the promise of decoding cellular heterogeneity at the functional level, yet achieving deep and reproducible proteome coverage from individual cells has remained a...
1
5
12
We are thrilled to announce that our ADIPOHealth project has been awarded funding through the prestigious ERC Synergy Grant! ADIPOHealth brings together a world-class team—Dale Abel (UCLA), Mauro Giacca (KCL), Guadalupe Sabio (CNIO), and Jesper Velgaard Olsen (UCPH) #ERCSyG
0
2
9
🚀 Check out our preprint on the new Thermo Scientific Orbitrap Astral Zoom MS 💫👇🏼 https://t.co/gwk98VoOo9
biorxiv.org
High-throughput proteomics is critical for understanding biological processes, enabling large-scale studies such as biomarker discovery and systems biology. However, current mass spectrometry...
1
4
24
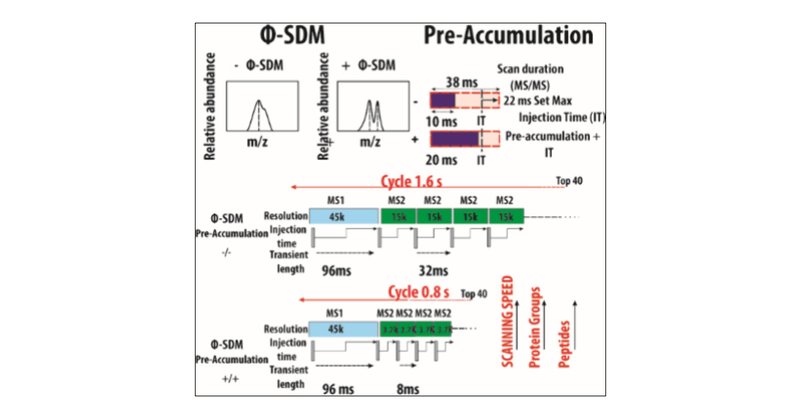
Our exploratory work on sensitivity boosting pre-accumulation, implemented into Orbitrap instruments, has been peer reviewed and published. We learned a lot here, and the method eventually became a commercial product. https://t.co/72RsvvtczH
pubs.acs.org
High-throughput mass spectrometry-based proteomics has gained increasing interest for both academic and industrial applications. As implementation of faster gradients has facilitated higher sample...
1
5
29
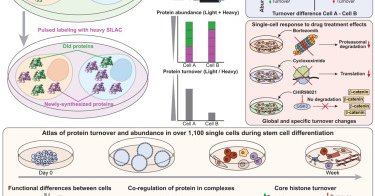
🚀 Excited to share our latest article in #singlecell proteomics published in Cell! SC-pSILAC allows to simultaneously measure protein turnover and abundance in single cells, unlocking the first large-scale, 2D proteomic insights at single-cell resolution! https://t.co/hFLrJzunjm
cell.com
The SC-pSILAC method enables single-cell measurement of both protein abundance and turnover, providing notable advances in the depth and versatility of proteomic technologies.
3
25
86
We are excited to share our new Nature Methods paper describing the Chip-Tip workflow for single-cell proteomics identifying >5,000 proteins in single cells, enabling PTM analysis without enrichment and throughput of up to 120 single cell samples per day: https://t.co/ZI19WxwzlK
1
38
164
Global analysis of protein turnover dynamics in single cells https://t.co/BsDBLrTziL
#biorxiv_sysbio
0
2
8
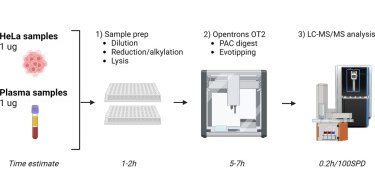
Fully automated workflow for integrated sample digestion and Evotip loading enabling high-throughput clinical proteomics - Molecular & Cellular Proteomics
mcponline.org
In BriefThe throughput of shotgun proteomics has been increased significantly and opened up for analysis of large clinical cohorts. Automation of the sample preparation step is highly desirable for...
0
6
45
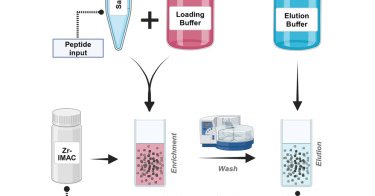
Our manuscript is online: Systematic optimization of automated phosphopeptide enrichment for high-sensitivity phosphoproteomics - Molecular & Cellular Proteomics
mcponline.org
In BriefOptimization of experimental parameters for automated on-bead phosphoproteomics sample preparation for low-input samples was conducted. Key factors assessed included glycolic acid concentra...
2
7
65
Excited to share that our manuscript exploring the analytical merits of the Orbitrap Astral mass spectrometer is out in Nature Biotechnology. Using nDIA, we profile >100 full yeast proteomes per day, or 48 human proteomes per day of ~10,000 proteins. https://t.co/nGdquJWbDt
1
23
119
Hey #Proteomics enthusiasts, come and join us next summer in Brixen. @BayBioMS part of orga team of this great event. More info will follow soon… #TeamMassSpec #Bioinformatics
The Brixen Summer School is back and is, as usual, building up a strong speaker list. Thank you @UKBspr and @EuPAProteomics for additional support. @EU_YPIC @JesperOlsenLab @lilley_ks @RubBritta @BayBioMS
https://t.co/pOqP98vMR2
0
2
9
Fully automated workflow for integrated sample digestion and Evotip loading enabling high-throughput clinical proteomics https://t.co/lmTKHx9LkO
#biorxiv_molbio
0
3
10
Did you ever wonder about how every parameter during phospho-enrichment affect your results? In our latest preprint we systematically analyzed almost every parameter, from beads amount to sequential enrichment. We even have data on the Astral MS!
Systematic optimization of automated phosphopeptide enrichment for high-sensitivity phosphoproteomics https://t.co/mxT9u5ToIx
#bioRxiv
2
14
54
🔬 Exciting new research on EGFR from @JesperOlsenLab ! Akihiro Eguchi, @kbemdal and colleagues used high-throughput proteomics to uncover the molecular mechanisms behind EGFR's response to its 6 highest-affinity ligands. Check out the preprint:
biorxiv.org
The epidermal growth factor receptor (EGFR) induces different signaling outputs depending on ligand identity and biological context. This phenomenon is known as functional selectivity, but the...
0
13
31
Less is more with the One-Tip method! From single mouse zygotes to HeLa cells, discover unmatched proteome depth & efficiency in our new preprint from @zilu_ye @PierreSabatier8 , coupling @EvosepBio and #Orbitrap #Astral #TeamMassSpec. https://t.co/FS44JljRXX
3
15
70
Our work at @JesperOlsenLab on describing and benchmarking hybrid-DIA methodology for simultaneous targeted and discovery phosphoproteomics is now out in @NatureComms: https://t.co/dVmUh9TTmn
0
19
54
Exciting developments in #proteomics with the new ultra-fast scanning DIA strategy in Orbitrap Astral MS. This innovation is set to reshape proteome analysis, blending speed, precision, and accuracy. Dive into the details at bioRxiv:
biorxiv.org
Mass spectrometry (MS)-based proteomics aims to characterize comprehensive proteomes in a fast and reproducible manner. Here, we present an ultra-fast scanning data-independent acquisition (DIA)...
1
37
120
Boost your analysis with our new hybrid-DIA method, developed by our superstar postdoc @mdval_ana, that allows to combine targeted and discovery #proteomics in just one run. We have used it to boost drug screening phospho-research on spheroids. Check it at https://t.co/fqnTMJS1Db
Targeted or DIA? No-one likes to choose when you can have it all. We present hybrid-DIA! A new method to combine targeted and discovery proteomics in one run. Don't miss again the key targets in a pathway in your DIA analysis. Check our last preprint https://t.co/z3z71QiM27
1
8
39
Proud to be named a Highly Cited Researcher 2022 by @ClarivateAG. See the full list https://t.co/akwYr0ZzwC
#HighlyCited2022
clarivate.com
The Highly Cited Researchers 2025 list identifies and celebrates individuals who have demonstrated significant and broad influence in their fields of research. Through rigorous selection criteria and...
3
2
62