2. In February, Dylan Cable followed up his popular RCTD method with C-SIDE, enabling differential expression for spatial transcriptomics (w/
@rafalab
)
Our new method, C-SIDE, for cell type-specific DE in spatial tx:
Detects DE in diverse contexts such as space, disease, cellular interactions. Controls for cell type effects and mixing. Led by Dylan Cable and colab w/
@rafalab
@macosko
@Vignesh_Shan
0
6
37
1
0
1
Replies
Happy New Year from the Chen lab!
As we all get back to our experiments, we wanted to look back and share some of our lab members' amazing work from 2022:
2
6
71
1. In January, we introduced RADARS, a new programmable way to target cells using RNA sensing
With
@idmjky
,
@jgkoob
,
@dawnchenx
,
@RohanKrajeski
, Yifan Zhang,
@jgooten
, and
@omarabudayyeh
Today we report in
@biorxiv
programmable RNA sensors that express any protein gated on specific RNAs in a cell. Much like gene sequencing necessitated gene editing tools, we answer “What is the Cas9 of single cells?” 🧵👇🏼w
@jgooten
&
@insitubiology
1/17
9
162
647
1
1
4
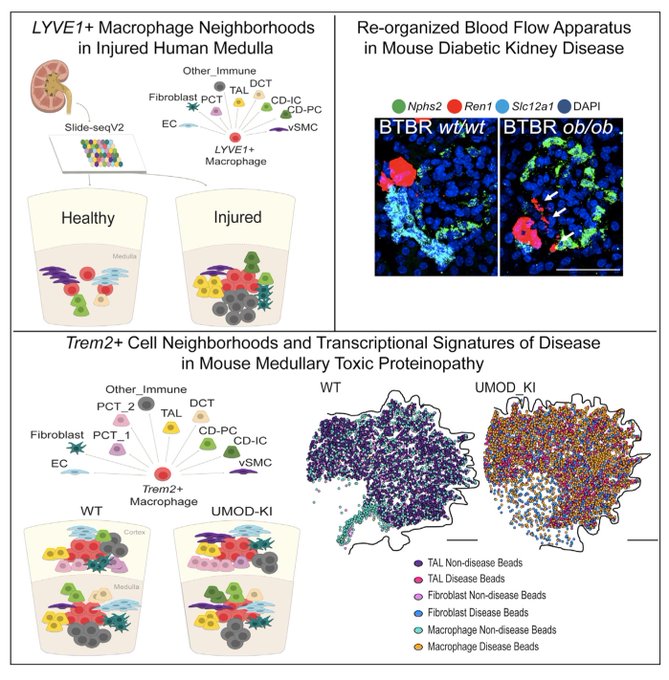
3. March saw the publication of
@JamieLMars
's work on the spatial organization of kidney disease
With Teia Noel,
@qbw_128
,
@haiqi_chen
, and
@AnnaGreka
🎉 Greka lab paper alert! Our team is excited to share a new story published in
@iScience_CP
led by
@JamieLMars
describing the use of Slide-seqV2 to identify disease-specific cell neighborhoods in human and mouse kidneys.
5
17
82
1
0
3
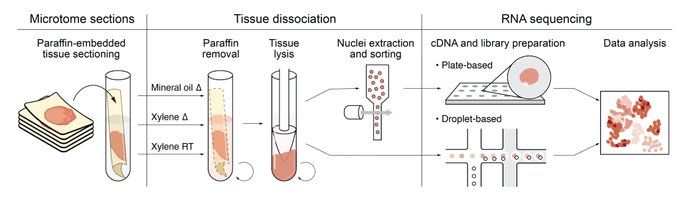
4. Jumping to August,
@hattaca
shared snFFPE-seq, paving the way for single nucleus RNA-seq of archival clinical samples
With Alexandre Melnikov, Cristin McCabe, and Aviv Regev
1
0
2
5. In September,
@s_mangiameli
introduced photoselective sequencing (PSS), which uses fluorescence imaging to make precise spatial genomic measurements (w/
@JD_Buenrostro
)
Sharing Photoselective Sequencing (PSS), a new method for genomic and epigenomic measurements within spatial regions of interest.
Thanks to the whole team
@haiqi_chen
,
@H3K4me1
,
@juliedobkin_
,
@DLesman
,
@JD_Buenrostro
, and Fei Chen.
(1/9).
1
43
113
1
1
2
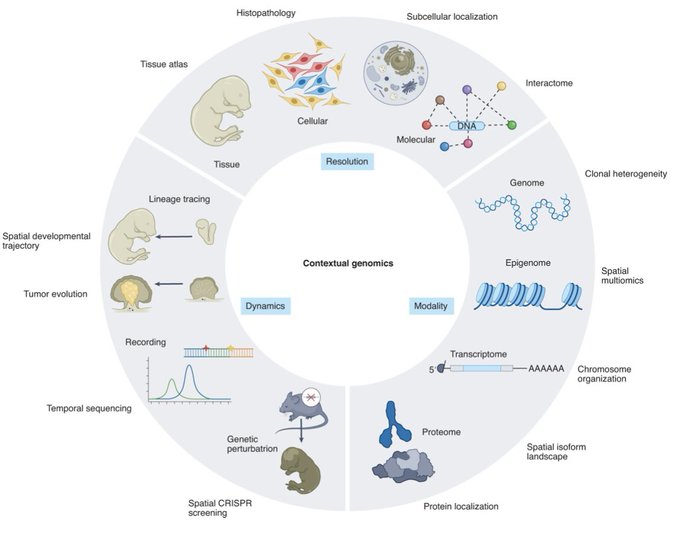
6. In October,
@Luyi_T
outlined a vision for the future of tissues genomics in a comprehensive technology review (w/
@macosko
)
Luyi is starting his own group at Guangzhou Lab, be on the lookout for openings!
Our review on Spatial Transcriptomics is out
@NatureBiotech
! With
@insitubiology
and
@macosko
we reviewed the technology advance in sequencing and imaging-based ST and envisioned the future of tissue genomic where dynamics and perturbation measured at spatial levels.
3
229
785
1
0
6
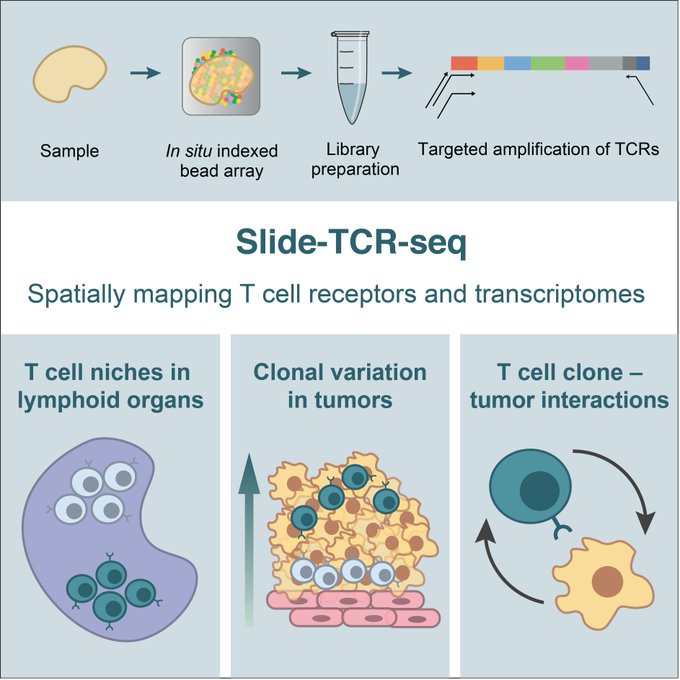
7. A week later,
@soph_liu
brought together spatial tech and immunology with Slide-TCR-seq, a method for spatially mapping T cell receptors and transcriptomes
With Bryan Iorgulescu, Shuqiang Li, and Catherine Wu
Presenting Slide-TCR-seq: our method for measuring the transcriptomes of cells & T cell receptor (TCR) sequences in situ!
This collaboration w/ Bryan Iorgulescu, Shuqiang Li, Catherine Wu, and
@insitubiology
led to some cool findings...
1/8
19
205
805
1
0
4
8. Lastly, in 2022 we started to really see the results of our work to make spatial tech more broadly accessible -- a true team effort led by Evan Murray and
@Irving_Barrera1
1
0
5
2022 brought so much new energy with our group moving into a new space and welcoming
@HuChenlei
,
@jacksonweir4
,
@sundakao
,
@SandeepKambham2
,
@benno_orr
,
@WollensakDavid
, &
@RuthRaichur
We also celebrated
@juliedobkin_
&
@Luyi_T
taking their next steps!
Started at the bottom (
@broadinstitute
10th floor), now we're here (Broad 11th floor) 🚀
Celebrating the opening of our new lab space with a DIY ribbon cutting ceremony
9
7
401
1
0
5
Here's hoping for another great year of science and lab fun in 2023!
Still in a reminiscing mood? Check out our 2021 year in review:
0
0
1