GlobalPneumoSeq
@pneumowatch
Followers
817
Following
669
Media
104
Statuses
959
GPS, the Global Pneumococcal Sequencing project funded by @gatesfoundation is sequencing 50,000 pneumococci to monitor global vaccine implementation and impact.
Joined January 2014
Have you read about our #GPS_project study on how genomic surveillance can have real impact on improving #pneumococcal vaccines? If not, perhaps put it on your reading list now!
thelancet.com
Our work reveals that GPSC10 alone is a challenge for serotype-based vaccine strategy. More systematic investigation to identify lineages like GPSC10 will better inform and improve next-generation...
We are very excited to share our #GPS_project study out in @LancetMicrobe – showing how genomic surveillance can have real impact on improving #pneumococcal vaccines. https://t.co/uKtXnbaVXg 1/14
0
1
2
Performing GPSC/lineage-specific analysis? We have made assemblies and annotations for 73 common GPSCs more accessible than ever: https://t.co/z2AiUhdLv7 Annotations were generated using Bakta v1.11.4 with the v6.0 full database.
github.com
The reference genomes for common Global Pneumococcal Sequence Clusters (GPSCs) - GlobalPneumoSeq/gpsc-reference-genomes
0
1
2
This paper is out on Microbial Genomics @MicrobioSoc ! https://t.co/pxSYOsK3TW
microbiologyresearch.org
Do you need a #bioinformatic tool to detect almost all serotypes in #penumococci ? The new SeroBA v2.0 can detect 102 of 107 pneumococcal serotypes
0
1
1
It’s such a pleasure to see that #GPS_project 🩻funded by @gatesfoundation has been featured in this AMR article by @sangerinstitute , showing our global efforts on evaluating AMR after vaccine 💉 rollout. https://t.co/VaF5DoPptW
sangerinstitute.blog
Our scientists and collaborators are working together to tackle one of the biggest public health threats: antimicrobial resistance. We are using cutting-edge genomics to hunt down drug-resistant...
1
1
2
This work is the combined effort of @HarryHung @kumarnaren13 Victoria Dyster @stephlo_lo @StephenBentley5 at @sangerinstitute; Corin Yeats at @TheCGPS; Benjamin Metcalf, Yuan Li, Paulina Hawkins, Lesley McGee at @CDCgov; and funded by @gatesfoundation
0
0
1
🦠🧪🧬 Excited to share that our GPS Pipeline, the portable and scalable genomic pipeline for Streptococcus pneumoniae surveillance, is now published in @NatureComms 🔗 https://t.co/zMpJWPTMEl
#GPS_project
nature.com
Nature Communications - The GPS Pipeline enables accessible and scalable genomic surveillance of Streptococcus pneumoniae. It performs quality control and in silico typing of sequencing reads with...
1
8
8
Try it out: 📂 Code: https://t.co/2CVajlV9Sw ✅ Features: QC, serotyping, MLST, GPSC assignment, AMR prediction. 🌍 Built for global use: from powerful HPC clusters to nimble laptops.
github.com
Nextflow Pipeline for processing Streptococcus pneumoniae sequencing raw reads (FASTQ files) by the GPS Project (Global Pneumococcal Sequencing Project) - GlobalPneumoSeq/gps-pipeline
1
0
0
Why it matters: 📊 Genomic surveillance is vital for tracking serotypes, AMR, and informing vaccines. ⚡️ GPS Pipeline makes this possible in a reproducible, scalable, and portable way.
1
0
0
🦠🧪🧬 Excited to share that our GPS Pipeline, the portable and scalable genomic pipeline for Streptococcus pneumoniae surveillance, is now published in @NatureComms 🔗 https://t.co/zMpJWPTMEl
#GPS_project
nature.com
Nature Communications - The GPS Pipeline enables accessible and scalable genomic surveillance of Streptococcus pneumoniae. It performs quality control and in silico typing of sequencing reads with...
1
8
8
A brilliant study led by @Quackscience @Lucy_Weinert @BugsWormsNBats at @Cambridge_Uni and collaborated with @gemmamurray at @ucl and @stephlo_lo at @sangerinstitute and @UniofBath @MilnerCentre
0
0
2
By linking their shifts in the phenotypic map above with the amino acid changes in PBP protein, we could pinpoint particular substitutions and its impact on resistance.
1
0
0
Some pneumococcal lineages are more resistance to beta-lactams - coincides with our observations in the #GPS_project
1
0
0
AMR cartography to show the MIC values of multiple antibiotics
1
1
0
[New pre-print🚨] When a single genotype (pbp) confer resistance to multiple antibiotics (penicillin and cephalosporins), how do we make sense of genetic changes and their impact on AMR? A new concept - AMR cartography is invented by @Quackscience
https://t.co/8UWQNdHgUr
1
2
2
AMR Cartography lets us visualise how different pneumococcal lineages have distinct multivariate resistance phenotypes. @pneumowatch
0
1
1
A collaboration with National Health Research Institute in Taiwan, National Taiwan University, @MilnerCentre and @sangerinstitute
0
0
0
Our findings show how dense population, high vaccine uptake, widespread antibiotic use, and mobility shape a distinct pneumococcal landscape. Sustained genomic surveillance is key to detect emerging threats, guide vaccine strategy for the ageing population, and monitor AMR.
1
0
0
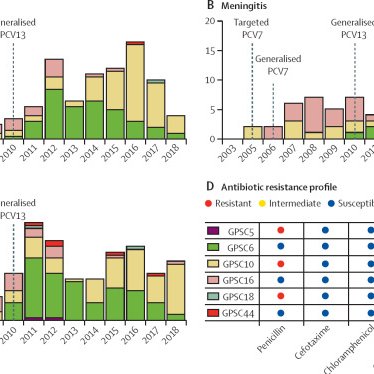
Key findings: 1) Increase in elderly IPD post-PCV13 2) Rapid post-vaccine emergence of GPSC9-serotype 15A. 2) Frequent capsular switching in GPSC6 4) GPSC1-19A are persistent after PCV13
1
0
0
[🚨 New paper] The first nationwide, longitudinal WGS analysis of Streptococcus pneumoniae in Taiwan (2006–2022). 1,343 isolates from 27 hospitals, we track serotype, AMR, and genomic lineage before and after PCV13. #GPS_project #PCV13_impact #Taiwan
https://t.co/PcRsmAmkov
microbiologyresearch.org
1
5
6