Zepeng (Phoenix) Mu
@ZepengMu
Followers
252
Following
702
Media
7
Statuses
73
(牟泽鹏) Postdoc @soumya_boston @BrighamWomens @broadinstitute | PhD @yang_i_li @UChicago | BS @UCAS1978 | Twin @charlie_z_mu
Boston, MA
Joined August 2016
RT @soumya_boston: So excited to share our latest with Betty Diamond and many colleagues from the AMP RA/SLE consortium, @NIH_NIAMS , and @….
biorxiv.org
Lupus nephritis (LN), a severe manifestation of Systemic lupus erythematosus (SLE), is a heterogeneous disease driven by diverse immune and tissue cell types. Current treatments of LN are non-speci...
0
6
0
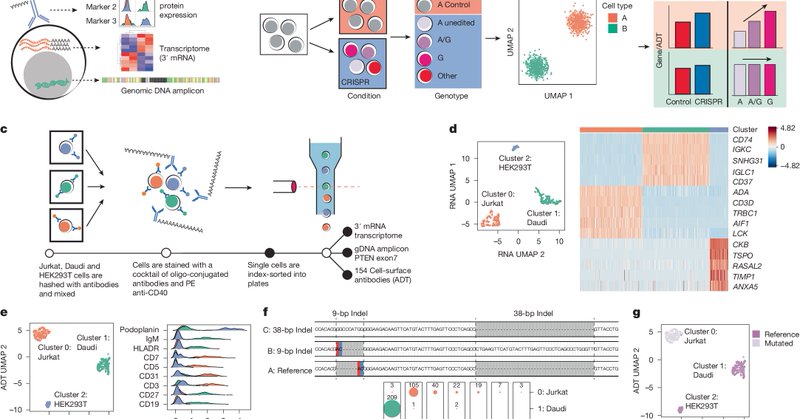
RT @YuriyBaglaenko: I am very excited to share our publication, out now in Nature, detailing our developed CRAFTseq methodology for capturi….
nature.com
Nature - A plate-based assay called CRAFTseq has been developed that uses ‘multi-omic’ single-cell RNA sequencing and direct genotyping of CRISPR edits to test the functional effects of...
0
35
0
RT @soumya_boston: Join me for this lecture to find out how we use genomics to decipher #autoimmunity and #rheumatoid arthritis.
0
6
0
RT @saorisakaue: 📣Excited to share my last postdoc paper with.@soumya_boston on eQTL mechanisms depending on where the RNA is in the cell!….
0
41
0
Huge thanks to my PhD advisor @yang_i_li and all collaborators especially @LB_Barreiro for their help and support!!! Huge thanks to all the donors! This data is a subset of a larger scRNA/scATAC data, read more about the cool discoveries in threads from @he_randolph!.
0
0
5
Excited to share the final chapter my PhD research on @medrxivpreprint! eQTLs only explain 30% of GWAS loci and 11% of trait heritability. Here we investigate if epigenetic QTL can provide new insights at the remaining GWAS loci.
medrxiv.org
Only a third of immune-associated loci from genome-wide association studies (GWAS) colocalize with expression quantitative trait loci (eQTLs). To learn about causal genes and mechanisms at the...
1
18
65
RT @XuanyaoLiu: Our work is out in AJHG! I wholeheartedly thank all the co-authors @marie_saitou @andywdahl @qbw_128 for their time and eff….
0
13
0
RT @JinghuiLi14637: Our work on trans regulation of proteome and how protein-protein interactions play a role in it🧬.
0
2
0
RT @he_randolph: check out our review discussing immune diversity through the lens of evolutionary biology! . we argue that past selection….
onlinelibrary.wiley.com
Humans exhibit considerable variability in their immune responses to the same immune challenges. Such variation is widespread and affects individual and population-level susceptibility to infectious...
0
10
0
RT @XuanyaoLiu: Our work led by @liliw_wang on identifying trans genetic effects of gene networks is out in @CellGenomics today!.
0
21
0
RT @YuriyBaglaenko: CRISPR editing is hard. Editing in primary human cells is even harder. Editing non-coding variants in human cells is d….
biorxiv.org
Genetic studies have identified thousands of individual disease-associated non-coding alleles, but identification of the causal alleles and their functions remain critical bottlenecks. Even though...
0
53
0
Very interesting and impressive work! Congrats to my friend Wenhe!.
Our paper is out in @CellGenomics! Huge thanks to all the collaborators for making it possible!.
0
0
8