Sandy Richardson
@RichardsonL1Lab
Followers
442
Following
2K
Media
63
Statuses
439
Group Leader—Developmental Molecular Genetics Lab and NHMRC Emerging Leadership Fellow at Mater Research Institute-UQ. Mama of 2. She/her. Views my own.
Brisbane, Queensland, AUS
Joined May 2009
🐭🧬Our paper on locus-specific mouse L1 activity and regulation, led by the indomitable @Patricia_Grds is out @genomeresearch!🧬🐭. This study evolved from a simple question: what defines the mutagenic potential of a new L1 copy in vivo?. A thread:.1/n
1
16
44
Renee will represent @materresearch, @UQmedicine, and #mobileDNA enthusiasts everywhere at the University final later this year! #scicomm #3MT.
0
0
0
Congratulations to multi-talented PhD student Renee Chu, who won the overall competition AND People’s Choice at the @UQmedicine 3MT final, using a thoughtful analogy to describe her research on jumping genes in early development 🧬🧫
1
0
12
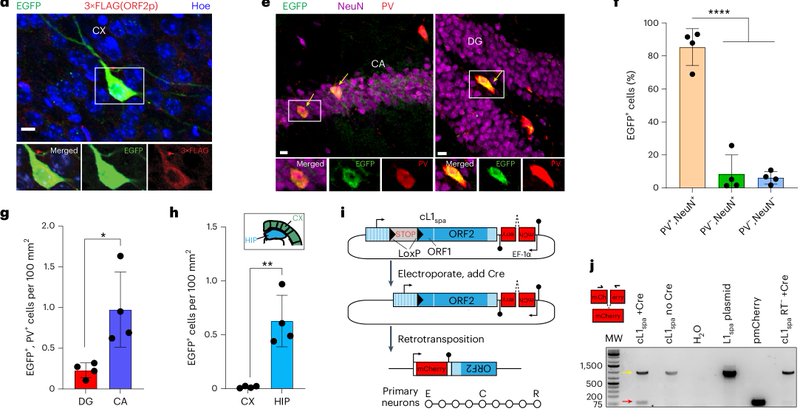
RT @gabriela_bodea: The impact of LINE-1 retrotransposons on neurobiology is not fully understood. Our new research in @NatureNeuro sheds l….
nature.com
Nature Neuroscience - LINE-1 retrotransposons are a type of mobile DNA element normally repressed in the body. Here the authors show that LINE-1 sequences can jump in mouse parvalbumin interneurons...
0
26
0
RT @FranciscoJSL2: The Andalusian Government (south Spain) has opened a call for 3-year postdoctoral positions. If you would like to join m….
0
14
0
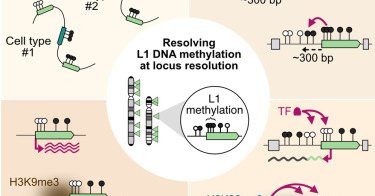
RT @CellGenomics: Locus-level L1 DNA methylation profiling reveals the epigenetic and transcriptional interplay between L1s and their integ….
cell.com
L1 retrotransposons are abundant sequences in the human genome, implicated in disease and evolution. Lanciano, Philippe et al. profile their DNA methylation patterns at single-locus resolution in...
0
6
0
RT @AEpiA: Interested in L1 retrotransposon regulation in development? . Check out AEpiA member @RichardsonL1Lab's recent paper in @genomer….
0
5
0
RT @manvendr7: Illuminating talk by @RichardsonL1Lab . Transcriptional regulation of Heritable activity of L1 activity in preimplantation e….
0
2
0
This study represents a huge amount of work by @Patricia_Grds, @Faulkner_Lab, and the most wonderful group of co-authors you could ask for: @Dory_genome, @MischaLundberg, @FranciscoJSL2, @gabriela_bodea, @_adamewing. 💗 and gratitude to all involved!. 20/20.
0
0
7
This also is likely to be true in humans, as somatic L1 insertions in the human brain frequently bear 5’ transductions consistent with upstream TSS usage. 18/n.
pmc.ncbi.nlm.nih.gov
A major unanswered question in neuroscience is whether there exists genomic variability between individual neurons of the brain, contributing to functional diversity or to an unexplained burden of...
1
2
15
The mouse L1 TSS pattern is consistent with the above study from the Kazazian lab, and also with work from John Moran’s lab demonstrating that a YY1 binding site is required for accurate human L1 transcription initiation. 16/n.
academic.oup.com
Abstract. The initial step in L ong In terspersed E lement- 1 (LINE-1) retrotransposition requires transcription from an internal promoter located within i
1
0
0
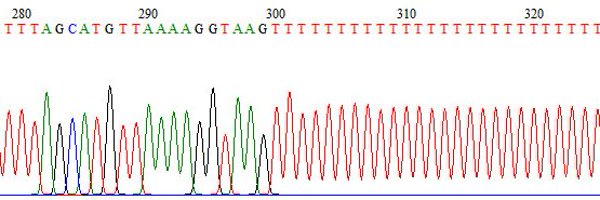
We next investigated the relationship between L1 5’UTR methylation and transcription start site (TSS) selection. We combined 5’RACE with PacBio SMRT sequencing in an approach similar to that of Prescott Deininger and @BelancioLab for human L1. 14/n.
academic.oup.com
Abstract. L1 elements represent the only currently active, autonomous retrotransposon in the human genome, and they make major contributions to human genet
1
0
0
The peaks in methylation flank binding sites for the transcription factor YY1 (red). Notably, our previous study led by @FranciscoJSL2 demonstrated that the YY1 transcription factor, or its binding site, is important for human L1 DNA methylation. 13/n
1
0
0
We therefore collaborated with @_adamewing , who developed and applied software packages TL;DR and Methylartist to examine the methylation state of human L1 insertions using ONT long-read DNA sequencing data. 9/n.
pmc.ncbi.nlm.nih.gov
Methylartist is a consolidated suite of tools for processing, visualizing and analysing nanopore-derived modified base calls. All detectable methylation types (e.g. 5mCpG, 5hmC, 6mA) are supported,...
1
0
0