Patricia Sullivan
@PatSullivann
Followers
364
Following
484
Media
26
Statuses
129
PhD from @KidsCancerInst in Computational Biology 🖥🧬 Developing tools to identify #splicing variants. @AustAmFulbright recipient 2022.
Sydney, Australia
Joined May 2019
Hard launching my new title: Dr Patricia Sullivan 🎓🥳 .After years of research, late-night writing sessions, and more splice-altering variants than I can count, I’m officially #PhDone
7
0
36
RT @FranMartinezGr: New UCSC track release!!!! #RareDisease #Genetics #UCSC. New Splicing Impact super track for hg38 and hg19. This super….
0
3
0
RT @hdashnow: I am excited to present v2!. A resource for tandem repeats associated with Mendelian disease. We have….
strchive.org
An archive of STRs associated with human diseases
0
10
0
RT @LaufferMarlen: David Cheerie and I will present at the first @N1Collaborative annual meeting tomorrow! We will discuss the newly establ….
medrxiv.org
Of the around 7,000 known rare diseases worldwide, disease-modifying treatments are available for fewer than 5%, leaving millions of individuals without specialized therapeutic strategies. In recent...
0
7
0
RT @Sumaiyalqbal: Published today @SpringerNature in @naturemethods . Genomics 2 Proteins portal @G2Pportal: a resource and discovery tool….
0
62
0
This work has been made possible by @Luminesce_All, @CancerAustralia, @MyRoomCCC, and @healthgovau. 👏. This is the 2nd publication from my splicing PhD (awarded this week!), which was generously supported by @dpetre, @FulbrightAUS, and @thekca. 🙏.
0
1
2
We’re excited to officially launch SpliceVarDB to the human #genetics community, where it will support variant curation and the development of in silico tools. 🚀. A big thank you to my coauthors @markjcowley, @MarkPinese, Alan, Julian, and the CompBio team at @KidsCancerInst.
1
1
3
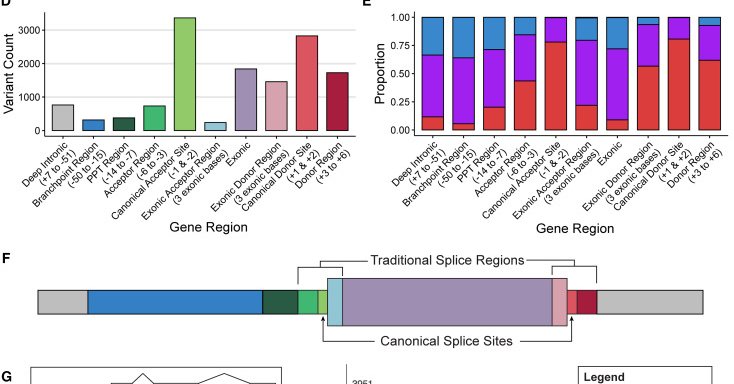
We’ve compiled 50,000 variants functionally demonstrated to affect (or not affect) splicing. These variants cover over 8,000 genes, covering 58% of OMIM genes and 68% of @ClinGenResource genes with established Gene-Disease Validity. 🧬.
1
2
1
We’re thrilled to announce the publication of SpliceVarDB: A comprehensive database of experimentally validated human splicing variants in @AJHGNews 🧬.📖 #splicing #GeneChat.
cell.com
Predicting whether a genetic variant will affect mRNA splicing is challenging. With SpliceVarDB we have consolidated and harmonized experimental splicing evidence for over 50,000 genetic variants...
1
34
126
RT @hdashnow: I am currently recruiting a computational postdoc in my lab at CU The candidate would develop and use….
0
37
0
RT @SL_Stenton: We’re happy to share our preprint on evidence yield from genome sequencing for the calibrated PP3/BP4 computational recomme….
medrxiv.org
Purpose To investigate the number of rare missense variants observed in human genome sequences by ACMG/AMP PP3/BP4 evidence strength, following the calibrated PP3/BP4 computational recommendations....
0
9
0
RT @ProgrammerDude: cat is the most misused thing by programmers new to linux. I cringe every time someone uses it wrong in a bash script.….
0
3K
0
RT @AliciaOshlack: Save the date! Have you heard the we are running a cancer bioinformatics symposium in June? Registration and abstract su….
0
24
0
RT @KidsCancerInst: EVERY child diagnosed with cancer in Australia now has access to precision medicine through our world-leading ZERO Chil….
0
2
0