Jesse Dixon
@Jesse_R_Dixon
Followers
1K
Following
360
Media
5
Statuses
142
Scientist | Assistant Professor at the Salk Institute for Biological Studies | M.D./Ph.D. from UC San Diego | Princeton Alum
San Diego, CA
Joined December 2017
It’s a bit belated, but I am happy to share the latest work from our group, led by a very talented post doc in the lab, Tessa Popay @tessapopay with great support from the 4D Nucleome consortium, Rita Allen, and the Pew Biomedical Scholars.
biorxiv.org
The organization of the genome in three-dimensional space is highly dynamic, yet how these dynamics are regulated and the role they play in genome function is poorly understood. Here, we utilized...
5
14
65
The federal government’s demand of $1 billion from @UCLA would devastate the nation’s best public university, cut off life-saving care, halt tech and economic growth, and reduce educational access. Stand with UC to protect our students, staff, faculty and our vital mission:
universityofcalifornia.edu
The federal government’s demand of $1 billion from UCLA puts the entire University of California system at risk. Stand with us to protect UC.
4
14
52
1/10 Excited to share our latest - the first whole-body map of both DNA methylation and 3D genome at single-cell resolution.
10
147
576
Introducing RAEFISH, our lab's new flagship image-based spatial transcriptomics technology that simultaneously enables single-molecule spatial resolution and whole-genome level coverage of long and short, endogenous and engineered RNA species in cell cultures and intact tissues.
1
55
276
Lastly, the genes that fail to be established during mitotic exit are less affected when NIPBL is degraded in asynchronous cells, suggesting that loss of cohesin may be more impactful for establishing expression during mitotic exit rather than maintenance of expression.
0
0
2
During mitotic exit, loss of NIPBL affects the expression of ~500 genes, with a strong enrichment for genes linked to distal super enhancers. With prolonged NIPBL depletion the cells undergo obvious changes in their morphology and changes of epithelial-vs-mesenchymal identity.
1
0
4
What distinguishes these stable chromatin loops? The clearest determinant we can identify so far is chromatin state. One interesting class is associated with repressive H3K27me3 sites in the genome, and these can persist for hours in the absence of NIPBL.
1
0
2
While we are limited in terms of temporal resolution, we observe both chromatin loops that rapidly lost and loops that persist in the absence of NIPBL. The loops are cohesin dependent as they are lost if we degrade RAD21 and require NIPBL for their formation during mitotic exit.
1
0
2
Inspired by recent reports from Anders Hansen and Luca Giorgetti using imaging to show that loops persist for 5-20 minutes in vivo, we wondered whether all chromatin loops were equally as transient or if there were populations that could “persist” on chromatin without NIPBL.
1
0
1
In this study, we were interested in the role of NIPBL in regulating the dynamic formation of chromatin loops. NIPBL is essential for loop extrusion to occur in vitro, seemingly promoting the formation of chromatin loops before Cohesin is evicted from chromatin by WAPL.
1
0
1
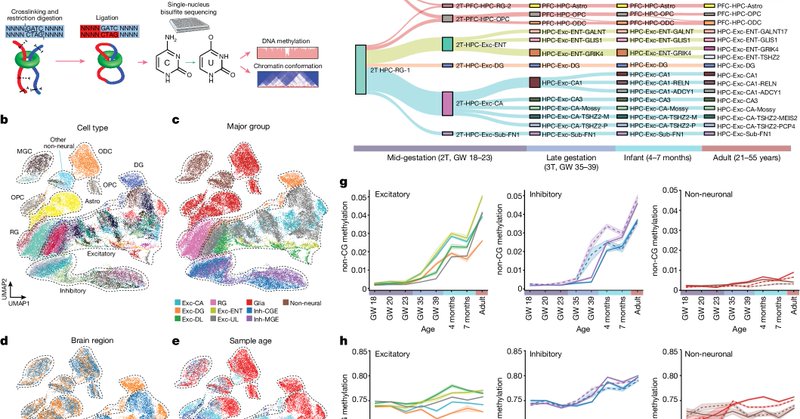
Glad to share the peer-reviewed version of the @mgheffel manuscript now in @Nature. In collaboration with @lab_paredes , we studied the chromatin conformation and DNA methylation changes during human hippocampus and frontal cortex development.
nature.com
Nature - Using a single-nucleus multi-omics approach, a study jointly profiles the reorganization of the epigenome and the three-dimensional chromatin conformation during the development of the...
14
46
194
Very excited and honored to be named a Pew Biomedical Scholar
🎉 Congrats to the 37 scientists joining Pew’s class of scholars and fellows! From engineering cutting-edge cancer treatments to unraveling the secrets of the genome, this outstanding group of researchers will help advance human health and medicine. https://t.co/voXzE8czkp
12
4
62
📢 Calling all recent PhD or MD graduates 📢 Explore immunology frontiers from neuro-immunology to infectious disease, while launching your independent career! Benefit from a 5-year appointment, mentorship, and funding support @salkinstitute. Learn more: https://t.co/WnjwEOHydr
0
31
82
Thrilled to announce the launch of our pilot run for the inaugural DISCOVER Salk Postdoc Symposium, where we invite potential postdocs to visit @salkinstitute, meet faculty, take lab tours, join career development workshops and seed new collaborative opportunities that would be
7
129
366
In our first Fellow search, we hired @zhou_jingtian, a computational biologist at the forefront of studying the gene regulatory landscape of the mammalian brain. Come join Jingtian, @li_lingyin, @LukeGilbertSF, @SKonermann, @genophoria, and @BrianHie + our growing Arc community
1
2
16
It takes too long to start your own lab—the average R01 recipient is 42 y/o 😵. Want to start your independent research group out of grad school? Come to my @ArcInstitute Fellows Program info session next Wed (Nov 15, 12pm PT). You'll get a $250K/year research budget + salary.
At Arc, we give early career scientists a springboard for launching their independent careers through our Science Fellow program. Learn more at our virtual information session next Wednesday, Nov. 15, from 12-1pm PST, hosted by Arc co-founder & Core Investigator @pdhsu. 👇
13
106
512
Happy to report that this study is now out in Nature Communications https://t.co/r1NI2S41vC
nature.com
Nature Communications - Cell lineage specification involves substantial chromatin conformation reorganization. Here, the authors integrate in-situ Hi-C and PLAC-seq to map the dynamic changes in 3D...
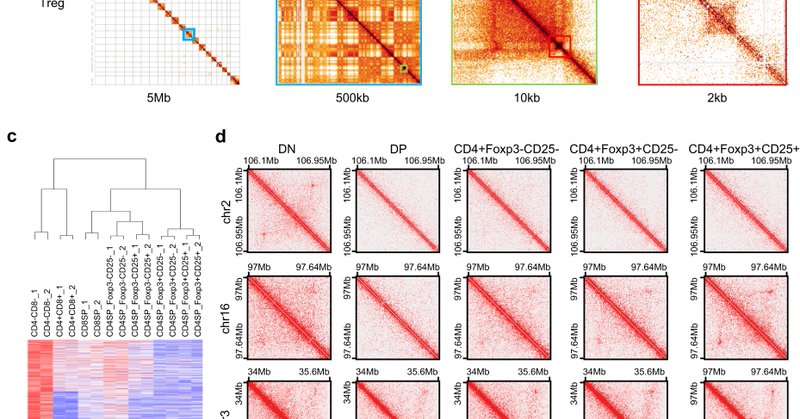
Excited to share the latest work from our lab, part of a collaboration with Ye Zheng's lab at Salk. We studying 3D genome organization in Tregs and investigated the role for Foxp3 in establishing Treg specific 3D genome interactions. https://t.co/FHGYZg4b00
1
6
38
.@Jesse_R_Dixon, Ye Zheng, Dong-Sung Lee & Zhi Liu discovered Foxp3 as the protein that determines regulatory #Tcells genome structure in mice–and that Foxp3 may be a good therapeutic target for regulating immunosuppression to treat #cancer @NatureComms
https://t.co/3f4mTE9FN4
salk.edu
LA JOLLA—Regulatory T cells are specialized immune cells that suppress the immune response and prevent the body from attacking its own cells. Understanding how these cells work is key to determining...
0
3
7
Attention Post-Docs!! We would love to invite you to The Salk Institute to give a talk about your research, meet Salk faculty, and create career-changing networks! The first ever Salk Rising Stars Symposium will be on April 10th. Apply here! Please RT! https://t.co/WX35cITcFp.
6
145
291
@zhou_jingtian joins as our first Science Fellow. Jingtian is a recent PhD graduate in Bioinformatics & Systems Biology from @UCSD and a former Salk Institute scientist. His pioneering work has developed new algorithms and insights into the epigenetic basis of brain development
2
4
31