Luca Denti
@l_denti
Followers
50
Following
59
Media
6
Statuses
30
Postdoc @unimib Computer Scientist with a passion for Bioinformatics
Joined July 2014
Morning! Let's start the day! SVDSS: structural variation discovery in hard-to-call genomic regions using sample-specific strings from accurate long reads by Luca Denti. #RECOMB2023
0
3
10
SVDSS combines multiple strategies to improve detection of structural variation using low-error long-read sequencing data. @l_denti @ParsoaKhorsand @DavisCompGen @BonizzoniPaola @PangaiaProject @RayanChikhi
https://t.co/EY2znczRBb
nature.com
Nature Methods - SVDSS combines multiple strategies to improve detection of structural variation using low-error long-read sequencing data.
0
10
21
Code is freely available at
github.com
Improved structural variant discovery in accurate long reads using sample-specific strings (SFS) - Parsoa/SVDSS
0
0
4
For now SVDSS can be used to discover insertions and deletions. We have plans to extend its functionalities to other SVs, such as inversions and translocations. Other extensions include somatic SVs discovery and SVs genotyping.
1
0
2
Analysis of SVs inside medically-relevant genes shows that SVDSS outperforms state-of-the-art tools in calling heterozygous SVs.
1
0
4
Even on CHM13 HiFi reads (provided by the T2T consortium, https://t.co/DGUyFwMYqZ), SVDSS achieves superior accuracy. But the homozygous nature of CHM13 makes SV calling relatively easier.
1
0
2
SVDSS achieves overall superior SV discovery performance outside Tier 1 regions of the genome and even at lower coverage (10x).
1
0
3
While most approaches focus on HG002 GIAB Tier 1 callset, we built our own truthsets using dipcall ( https://t.co/LLXhMBOF2T) and starting from HG002/HG007 assemblies. Truthsets are publicly available at
1
0
3
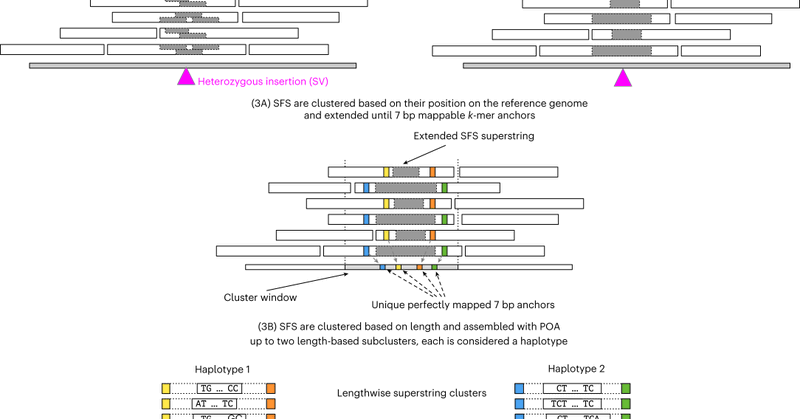
SVDSS achieves significant improvements in calling SVs in repetitive and hard-to-call regions of the genome.
1
0
3
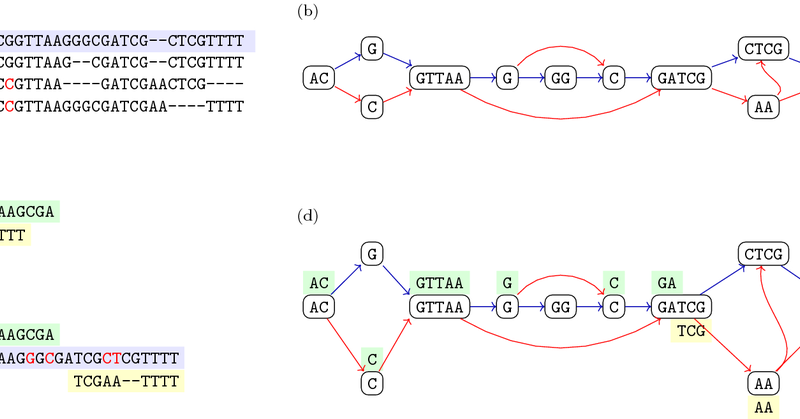
SVDSS finds potential SV sites using SFS and performs local POA of clusters of SFS to produce accurate SV predictions.
1
0
2
SVDSS is based on our recent notion of sample-specific strings (SFS, from https://t.co/faGaEMJon8) and effectively leverages alignment-free, mapping-based, and assembly-based methodologies.
academic.oup.com
AbstractMotivation. Comparative genome analysis of two or more whole-genome sequenced (WGS) samples is at the core of most applications in genomics. These
1
1
4
Our work on Structural Variations discovery from accurate long reads (@PacBio HiFi) is out ( https://t.co/EmfM2xunJo). Joint work with @l_denti @ParsoaKhorsand @BonizzoniPaola @RayanChikhi @DavisCompGen
1
21
55
Pangenome Graph Construction from Genome Alignment with Minigraph-Cactus https://t.co/SIukrRJpTT
#biorxiv_bioinfo
0
18
44
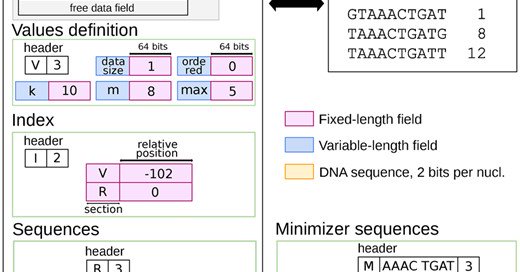
In this new Bioinformatics Application Note, we introduce a file format named KFF for efficiently storing kmers on disk. https://t.co/20gTzasFuD
academic.oup.com
AbstractSummary. Bioinformatics applications increasingly rely on ad hoc disk storage of k-mer sets, e.g. for de Bruijn graphs or alignment indexes. Here,
3
48
163
First steps in #pangenomics? Check out our new tutorial! https://t.co/VlOzRfuZzl Thanks @jasmijnbaaijens Paola Bonizzoni Christina Boucher @giandellavedova Raffaella Rizzi @jltsiren @PangaiaProject
link.springer.com
Natural Computing - Computational pangenomics is an emerging research field that is changing the way computer scientists are facing challenges in biological sequence analysis. In past decades,...
2
24
70
Code is freely available at
github.com
Comparative genome analysis using substring-free sample-specific strings (SFS) - Parsoa/PingPong
1
0
1
SFS strings are more expensive to compute than k-mers. In the paper we introduce an heuristic algorithm that finds non-overlapping SFS strings efficiently.
1
0
0
There are intrinsic similarities between SFS strings and other stringology concepts such as maximal exact matches, however we have not yet fully elucidated them.
1
0
0
This makes sense because SFS strings can better capture variations in repeated regions where k-mers would be confused by many identical instances.
1
0
0