DiMicheleLAB
@DiMicheleLAB1
Followers
2K
Following
771
Media
61
Statuses
446
Lorenzo Di Michele's research group @cebcambridge @MemBiophysIC @fabriCELL_UK #Nanotechnology #SyntheticCells #Biophysics
Cambridge, United Kingdom
Joined May 2018
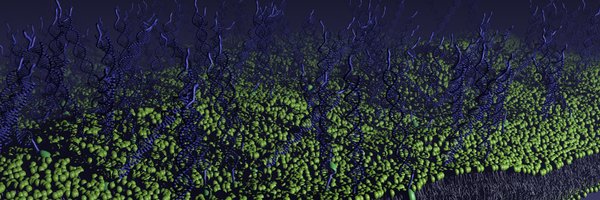
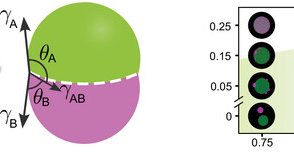
Thrilled to share this new Adv. Sci. paper by @dianatanasedt with Dino Osmanovic @rogerrubio Layla Malouf and Elisa Franco. We demonstrate a modular approach to program internal phase separation in DNA condensates 1/n .
advanced.onlinelibrary.wiley.com
The modular, programmable system of DNA nanostructures developed provides programmatic control over multiphase condensate behavior, enabling mapping onto a predictive Flory-Huggins model. This...
2
7
33
Thanks also @rogerrubio Layla Malouf, @FabriniGiacomo Sabrina Pia Nuccio @cebcambridge and funding from @ERC_Research @BBSRC @royalsociety n/n.
0
0
4
Thrilled to share our latest preprint on expressing synthetic organelles made from RNA nanostructures in bacterial cells. Great work from @BrianNgSH, Catherine Fan, @milandordevicc, Adam Knirsch, coll. Graham Christie @TakinoueLab @CicutaGroup 1/n.
biorxiv.org
Designing synthetic biomolecular condensates, or membrane-less organelles, offers insights on the functions of their natural counterparts, and is equally valuable for cellular and metabolic enginee...
1
5
24
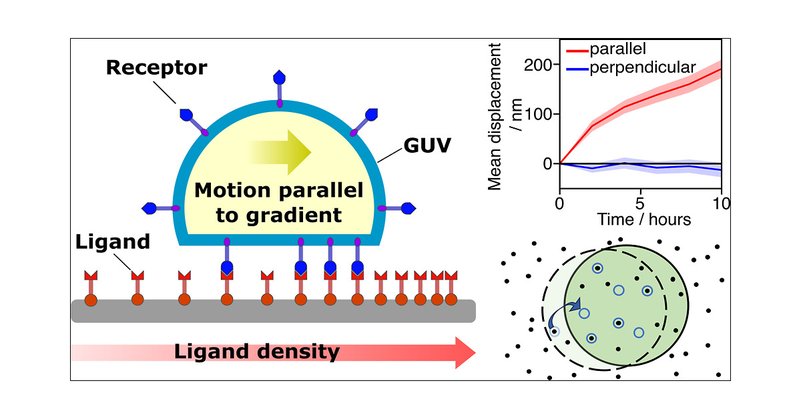
RT @YuvalElani: Check out our PhD student Hannah Sleath’s paper on vesicle-based synthetic cells that can crawl up surface concentration gr….
pubs.acs.org
Multivalent adhesion between cell-membrane receptors and surface- or particle-anchored ligands underpins a range of active cellular processes, such as cell crawling and pathogen invasion. In these...
0
7
0
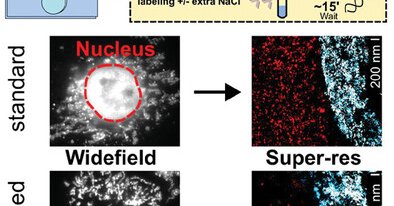
RT @A_Clowsley: Check out @evelucinskaite's recently published work reducing non-specific interactions of DNA-PAINT secondary antibodies. Y….
onlinelibrary.wiley.com
Conventional immunohistochemistry labeling with oligonucleotide conjugated secondary antibodies can lead to significant non-specific uptake across biological cells and is especially prominent within...
0
2
0
The candidate will use Synthetic Cell models and DNA nanotechnology to study interactions between lipid membranes and condensates. The project involves exciting training opportunities across Europe and placements with @SchwilleLab in Munich and @ContiniClau in London.
1
1
4
Happy new year everyone, and some great news!.We have an opening for a PhD candidate to join our lab @cebcambridge @Cambridge_Uni in October 2024 as part of the new exciting @MSCActions Doctoral Network @ComeInCell .
1
9
18
RT @EngBioIRC: Great talks from from Juliette @BucciJuliette @DiMicheleLAB1 and Honghao Su @HonghaoS @plantsci at our EngBio ECR Meet today….
0
4
0