Martin Vögele
@Blendenfleck
Followers
4,547
Following
3,023
Media
495
Statuses
3,191
Computational biophysicist at Schrödinger, prev. @Stanford & @MPIBP | amateur photographer | history+language nerd | ö=oe | 🇩🇪 🇪🇺 🏳️🌈 | all views my own
New York, NY
Joined July 2014
Don't wanna be here?
Send us removal request.
Explore trending content on Musk Viewer
The Place
• 352326 Tweets
Cataluña
• 115090 Tweets
#FatalTrouble
• 112862 Tweets
#MEMORABILIA_OutNow
• 89510 Tweets
चुनाव आयोग
• 84326 Tweets
つばさの党
• 76768 Tweets
Illa
• 71460 Tweets
Teeth
• 63956 Tweets
Lucifer
• 60115 Tweets
Puigdemont
• 59717 Tweets
NuNew 3rd Showcase
• 40797 Tweets
We Love Daddy Tay🩵
• 35780 Tweets
#ネプリーグ
• 30548 Tweets
Kamuda Tasarruf
• 26407 Tweets
Happy New Week
• 23762 Tweets
作者の顔
• 16579 Tweets
Mehmet Şimşek
• 16557 Tweets
STOP MISTREATING NEWJEANS
• 12711 Tweets
Square Enix
• 12013 Tweets
Solar POWER
• 11745 Tweets
Wacom
• 11348 Tweets
Last Seen Profiles
Pinned Tweet
Now online in
@JCIM_JCTC
:
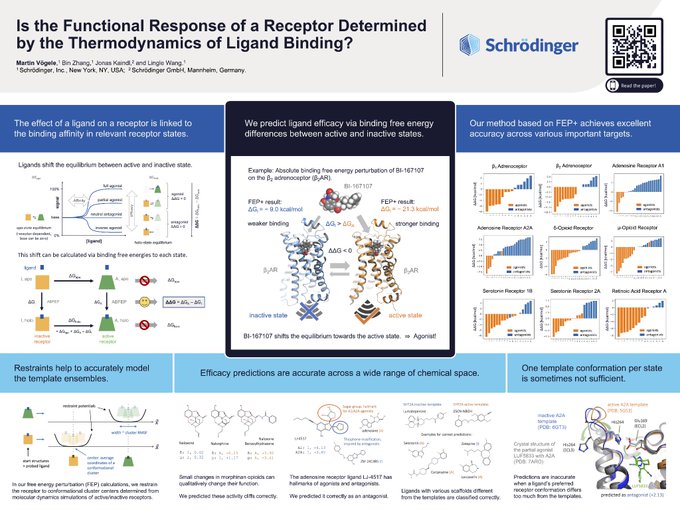
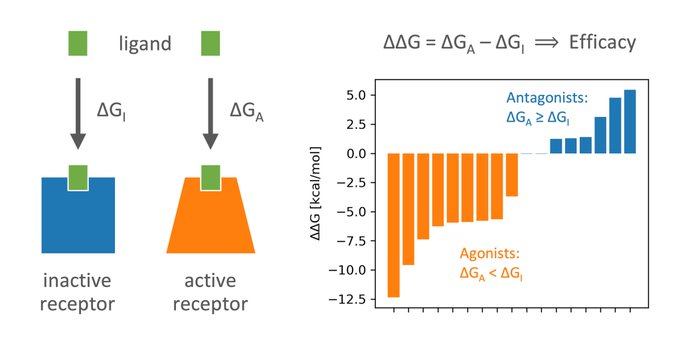
“Is the Functional Response of a Receptor Determined by the Thermodynamics of Ligand Binding?”
Our answer: Yes, it is!
And we can use it to predict ligand efficacy.
6
33
147
Exciting times ahead:
In two weeks, I am going to move to New York City to join
@Schrodinger
as a senior scientist in the life science software team.
👨💻
This is a dream coming true and I am deeply grateful for everyone who has been supporting me on the way there.
🙏
53
0
380
Pro tip for junior academics: If a professor has given two students or postdocs the same research project to see who does it faster, run and stay away from them. Competition is tough enough between labs; you don't need it within.
#AcademicTwitter
#AcademicChatter
#ScienceTwitter
4
7
109
This was definitely the most pleasant surprise of the year: Walter Greiner Award for best physics PhD thesis 2019/20 at
@goetheuni
. 🥳
(+ Corona-delayed remote ceremony)
Many thanks to the
@MPIbp
Theor. Biophysics group for the opportunity and the support - and the amazing time!
21
0
109
Arrived in Philadelphia for the
@BiophysicalSoc
Annual Meeting, my first in-person conference after almost four years.
#BPS2024
Come visit me at my poster (2643-Pos) on Wednesday 10:30 am in Exhibit Hall AB, B35.
3
4
90
Sign at the railway station:
"Trespassers will be cited."
Me, an academic:
*runs back and forth across the tracks*
#AcademicTwitter
1
12
88
Best paper award for ATOM3D!
🎉 ⚛ 🍾
Many thanks and congratulations to
@raphaeljlt
,
@PatriciaSuriana
,
@awfderry
and everyone who contributed! It has been a blast to be part of this project and I am looking forward to
#NeurIPS2021
.
9
7
86
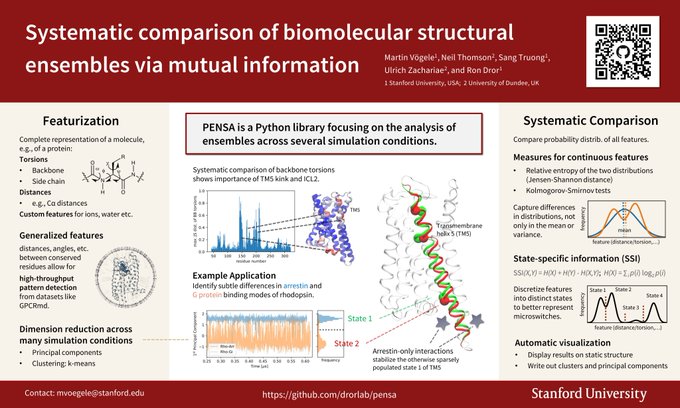
New PENSA release: Main addition in version 0.2.8 is functionality to estimate uncertainty via block analysis. Thanks to
@njthomson_
for taking the lead on implementing this!
Update it via: pip install pensa==0.2.8
Feedback is always welcome!
0
16
86
I contributed to
@500QueerSci
, a visibility campaign for LGBTQ+ people and their allies working in STEM and STEM-supporting jobs. 🏳️🌈
2
10
86
#autumn
#sunset
behind the
#library
of the Max Planck Institute of Biophysics in
#frankfurt
.
@maxplanckpress
@Stadt_FFM
1
7
80
Life update: My appointment at
@Stanford
has been extended beyond the end of my fellowship and I found a new room. 🥳
Being cautiously optimistic, I'm going to move closer to campus – to beautiful Menlo Park. 🙂
5
0
81
I am happy to join the pool of early-career reviewers for
@eLife
and looking forward to be part of the
@eLifeCommunity
in "structural biology and molecular biophysics."
3
3
80
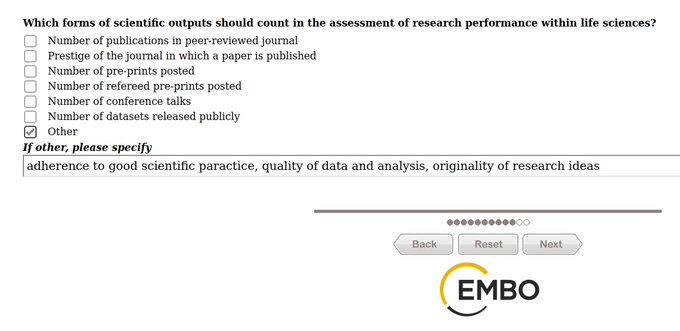
Trying to switch to a Central European sleep rhythm so I can virtually attend
#EMBOBioSim
.
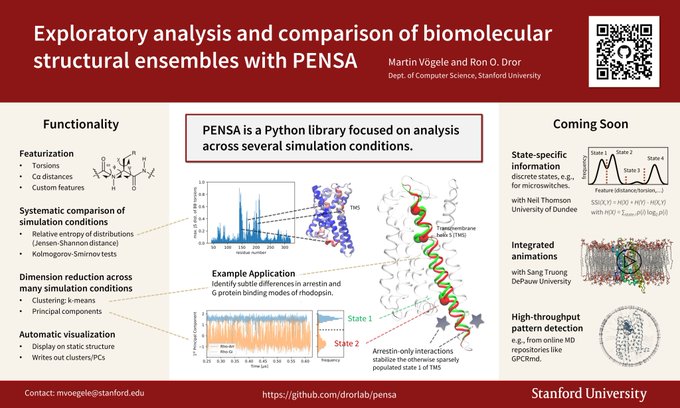
I have a poster on PENSA tomorrow (P87).
3
5
72
Just submitted my poster for the
@BiophysicalSoc
annual meeting.
Meet me there on Wednesday, February 24, 2021 to ask whether we've already managed to release some of the "coming soon" functionality. 🤞
#BPS2021
3
4
72
Me: "Make sure we cite all relevant literature."
Raphael: "Everything?"
Me: "Yes, everything."
Raphael:
Unhappy your work hasn't been cited? 😡 Finding it tricky to get credit attribution right? 🧐 We present a definitive solution: simply cite everything!
11M citations, 200K pages, 1.8 GB PDF.
Enabled by the great
@opencitations
resource.
27
224
1K
4
5
65
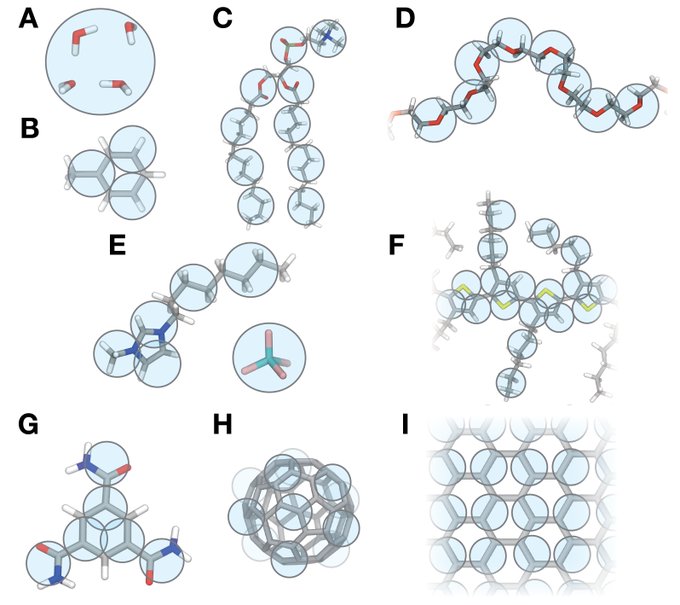
Had a great read catching up with coarse-grained simulations in materials science. A lot has happened in the last few years:
Fascinating review by
@ricalessandri
, Fabian Grünewald and
@CG_Martini
.
1
11
61
So excited to release this project into the wild. 🥳
ATOM3D started as a coffee break discussion with
@raphaeljlt
and escalated into a huge data curation and benchmark project with
@PatriciaSuriana
,
@awfderry
and many more joining in. Big thanks to everyone in this amazing team!
1
4
62
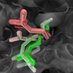
Rotational diffusion in membrane protein simulations depends on the box size!
Luckily, these effects turned out to be easier to estimate than for translational diffusion.
We
@MPIBP
even found an easy rule for the size of the simulation box to avoid them.
2
16
58
Last week I recorded a talk in which I claim that protein structure prediction is a hard and yet unsolved problem. When the talk will stream next week, this
#CASP14
result will make me look like an clown. 😅
CASP14
#s
just came out and they’re astounding—DeepMind looks to have solved protein structure prediction. Median GDT_TS went from 68.5 (CASP13) to 92.4!!!! Cf. their 2nd best CASP13 struct scored 92.8 (out of 100). Median RMSD is 2.1Å. I think it's over
35
566
2K
1
0
56
“The Unknown Postdoc Weeping Over the Rejected Manuscript”
— Marble sculpture, 1908
#PostdocAppreciationWeek
#NPAW
0
2
54
Good news for molecular simulations:
@CompBioPhys
@VytasGapsys
confirm that the simulation box size only minimally affects thermodynamics and kinetics of typical biomolecular systems:
... in contrast to diffusive dynamics. 😉
2
7
50
Happy
#LGBTQSTEMDay
!
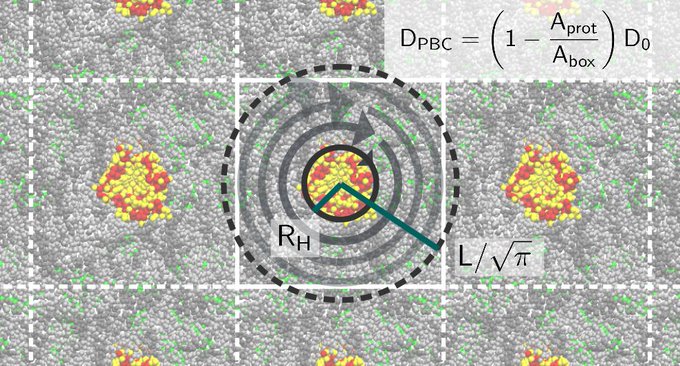
Even some of the G protein-coupled receptor complexes I'm currently simulating are in pride mood.
0
0
48
Cool paper on the mechanism of a blood-brain-barrier lipid transporter – with MD simulations by my amazing colleague
@DenizAydinPhD
.
#lipids
#biophys
#compchem
2
4
48
Our ATOM3D paper was selected for an oral presentation at
#NeurIPS2021
in the first ever Datasets and Benchmarks Track.
🎉🍾🎉
Big thanks to the reviewers who "note the importance of the problem, the clarity of the presentation, and the scope of the work."
0
2
49
Pro tip for all junior scientists:
Create a Google Scholar profile for yourself!
This is little effort but helps many people to find your papers - especially if they only remember your name or pieces of the title.
#AcademicTwitter
#ScienceTwitter
3
4
49
The
@lindaunobel
meeting is over - a fantastic week with amazing people. 😍
I learnt a lot about great discoveries in physics and visions for its future. We also discussed a lot about our roles in science and how to make it more open, welcoming, and beneficial to society.
#LINO19
1
5
45