Marta Farré
@marta_f_b
Followers
404
Following
1K
Media
33
Statuses
652
Senior Lecturer @biokent working on evolutionary and functional genomics
Canterbury, England
Joined March 2010
New paper from the lab in Animal Genetics! We developed a new method to detect chromosome rearrangements (in this case reciprocal translocations) in pigs using Hi-C. Great job by @FrancesBurden3 , in collaboration with PIC. #chromosomerearrangements
onlinelibrary.wiley.com
A chromosomal rearrangement such as a reciprocal translocation (RT) in a breeding boar can produce unbalanced gametes during meiosis, leading to a decreased litter size with detrimental economic...
0
6
18
RT @BrianHie: We trained a genomic language model on all observed evolution, which we are calling Evo 2. The model achieves an unprecedent….
0
327
0
RT @coreykirkland21: So happy to share that I passed my PhD viva on Friday!! 🤓📖. A huge thank you to my supervisor @marta_f_b for all her s….
0
1
0
RT @molecology: New ME Special Issue!.𝘼 𝙂𝙚𝙣𝙤𝙢𝙞𝙘 𝙐𝙥𝙙𝙖𝙩𝙚 𝙤𝙣 𝙩𝙝𝙚 𝙀𝙫𝙤𝙡𝙪𝙩𝙞𝙤𝙣𝙖𝙧𝙮 𝙄𝙢𝙥𝙖𝙘𝙩 𝙤𝙛 𝘾𝙝𝙧𝙤𝙢𝙤𝙨𝙤𝙢𝙖𝙡 𝙍𝙚𝙖𝙧𝙧𝙖𝙣𝙜𝙚𝙢𝙚𝙣𝙩𝙨, edited by @HAugustijnen, @….
0
13
0
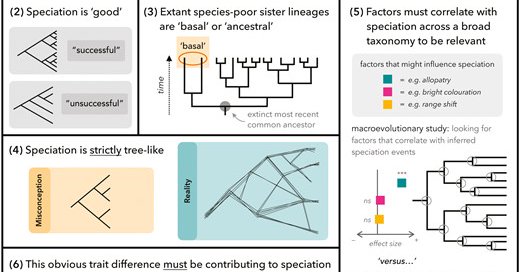
RT @joana_meier: My entire team wrote an opinion paper about Common misconceptions of speciation research. Each section is written by 1-2 t….
academic.oup.com
Abstract. Speciation is a complex process that can unfold in many different ways. Speciation researchers sometimes simplify core principles in their writin
0
236
0
RT @NaturePortfolio: Two papers in @Nature present a new genome editing technique that enables the insertion, inversion and deletion of lon….
0
292
0
At the Annual Postgraduate Conference @UniKent @GRCKent I was the runner up for Best Research Supervisor Award of the Uni😊 THANKS to my research group @Squigley1998 @ccanedoribeiro @coreykirkland21 and @FrancesBurden3 for nominating me! 🥹 🥹#EmotionalAndSlightlyEmbarrassed
1
2
11
RT @JesperBoman: MEIOTIC DRIVE AND THE EVOLUTION OF CHROMOSOMES. In 2001, Pardo-Manuel de Villena and Sapienza observed that positions of c….
0
23
0
RT @jkpritch: Two new chapters from my open-access textbook in human population genetics are now online:.- Population structure: I. Ancestr….
0
230
0
RT @biokent: Join us on Tuesday 12 March 1pm in SLT1 where @marta_f_b will present her lab's work on 'Genome evolution in mammals: from mar….
0
5
0
RT @AdamEyreWalker: PhD position available to work to work with me, @FrancesPearl2 and @marta_f_b on Patterns of Mutation in the Human Geno….
0
4
0
@Squigley1998 took over @joana_damas and my messy scripts to create a super cool R package - find the manual and core here
0
1
2
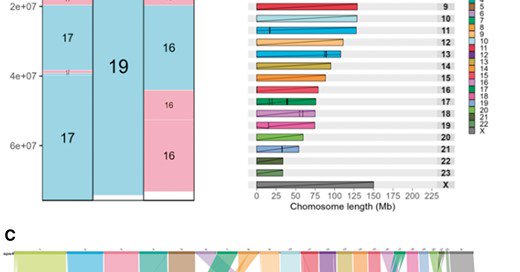
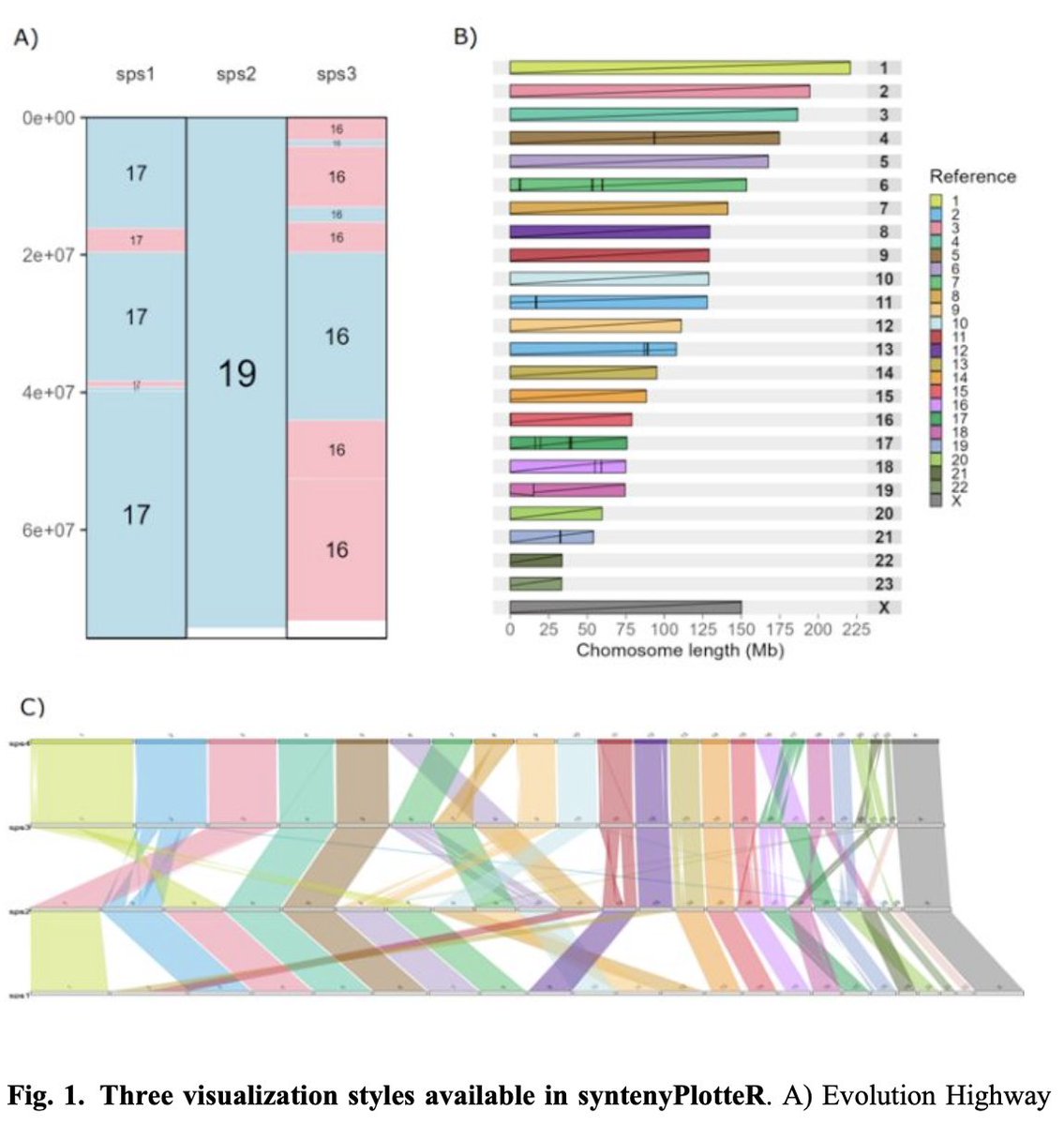
🚨 And another paper from the lab! We developed a new R package to plot genome synteny - now published in @BioinfoAdv and deposited in CRAN
academic.oup.com
AbstractMotivation. The rapid increase in the number of chromosome-scale genome assemblies has renewed interest in chromosome evolution studies. The visual
syntenyPlotteR: a user-friendly R package to visualize genome synteny, ideal for both experienced and novice bioinformaticians by Sarah Quigley, @joana_damas, Denis Larkin, and @marta_f_b in
3
35
119
Our results characterize at fine detail the location of chromosome rearrangements in ruminant evolution and provide new insights into the formation of these changes! #proudPI.
0
0
0
@Squigley1998 then used all single copy orthologous genes in 4 species (cow, sheep, pig and human) and used transcriptome data for 9 tissues (brain, colon, heart, kidney, liver, lung, skeletal muscle, spleen and testis) to identify housekeeping genes
1
0
0
🚨New paper from the lab! "Patterns of chromosome evolution in ruminants" @CrisAriasSarda defined 5 ancestral karyotypes of ruminants using 26 genomes! She identified all the #chromosomeRearrangements between them.
1
12
27