Liau Lab
@liaulab
Followers
719
Following
86
Media
8
Statuses
86
Academic startup at the interface of chemistry and cancer epigenetics @ Harvard CCB 🔬
Cambridge, MA
Joined June 2018
Familiarity and/or training in any aspect of genomics, sequencing data analysis, and/or CRISPR genome editing is a plus but not required. Enthusiasm to learn new areas of biology and data analysis as well as work with and mentor others are essentials.
1
0
1
We’re looking for a chemical biologist interested in pursuing postdoctoral research at the interface of chemical biology and genomics to join our collaborative team starting in Q2 2023. A strong background in chemical biology (broadly defined) and molecular biology are required.
1
2
7
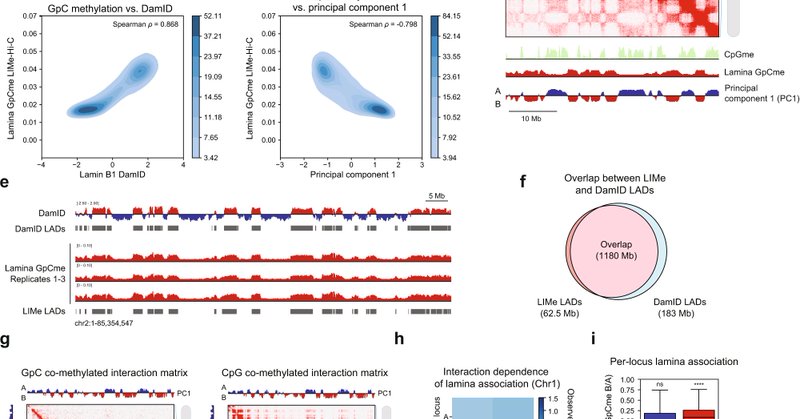
Today we share in @NatureComms our method called LIMe-Hi-C (Lamina-Inducible Methylation and Hi-C) & how we used it to uncover Polycomb-Lamina antagonism. Big thanks to Allison, Shelby, @MartinAryee, and @liaulab, & to reviewer 1 for the detailed comments.
nature.com
Nature Communications - Here the authors developed ‘Lamina-Inducible Methylation and Hi-C’ (LIMe-Hi-C) to simultaneously measure chromosome conformation, DNA methylation, and nuclear...
4
23
135
How do you get resistance mutations when a drug nearly carbon copies the substrate? 🤔 By distal, allosteric activation! See how we exploit this chemical subterfuge to systematically search for allostery in DNMT1 & UHRF1 in our latest pre-print. 1/N https://t.co/rBmjmpytVV
biorxiv.org
Allostery enables dynamic control of protein function. A paradigmatic example is the tightly orchestrated process of DNA methylation maintenance. Despite their fundamental importance, systematic...
3
9
63
We have another new preprint out! 🎉
keeping the @liaulab 2022 streak alive w/ another preprint out now tl;dr — we exploit activity-based selection and develop new tools for CRISPR scanning to tackle the difficult task of finding allosteric sites 🧵thread below (1/7): https://t.co/hZdnIneGlx
0
0
8
Congratulations to Dr. Brian Liau (@brian_b_liau) at @Harvard for being named a 2022 Camille Dreyfus Teacher-Scholar! @liaulab @HarvardCCB
1
3
24
We can map DNA-DNA contacts to chart 3D genomes with classic Hi-C, but where are these contacts positioned in the nucleus? Proud to release a new flavor of Hi-C, LIMe-Hi-C, a method that tags & maps DNA-DNA contacts relative to the nuclear periphery! (1/N)
biorxiv.org
The genome can be divided into two spatially segregated compartments, A and B,[1][1],[2][2] which broadly partition active and inactive chromatin states, respectively. Constitutive heterochromatin is...
3
28
152
DNMT3A is 1 of the most highly mutated genes in heme malignancies & clonal hematopoiesis with >250 mutations. Ever wonder what mutations in DNMT3A do to its function?🤔 (1/3)
11
59
368
Super excited to announce that we’ve just submitted the latest @liaulab ~manuscript~! Tldr: we used base editors + a methylation reporter to do a mutational scan of DNMT3A, a key cancer gene. Highlights below, but check out our preprint at @biorxiv! https://t.co/BDkaMnbAxM
biorxiv.org
DNA methylation is critical for regulating gene expression, necessitating its accurate placement by enzymes such as the DNA methyltransferase DNMT3A. Dysregulation of this process is known to cause...
8
19
87
Excited to share our latest work revealing a repressive methylation ceiling in EZH2-mutant lymphoma! We made this discovery by finding drug addiction mutations, a strategy we expect to be broadly useful for uncovering new biology across drug targets. (1/7) https://t.co/OnU8243rTS
biorxiv.org
Cancer mutations in Polycomb Repressive Complex 2 (PRC2) drive aberrant epigenetic states. Although therapies inhibiting the PRC2 enzymatic component EZH2 are FDA-approved, oncogene-specific depend...
8
30
155
Check out this awesome perspective by grad student @steph__rob on exciting work by @lilchromatinboy, @Palgosavi, @brian_b_liau, @liaulab to profile resistance-driving mutations in neosubstrates of molecular glues! Thanks @ACSCentSci for the opportunity!
pubs.acs.org
CRISPR-suppressor scanning reveals potential resistance mechanisms of cancer cells to molecular glue degraders via neosubstrate alteration.
3
8
51
#ASAP by @brian_b_liau & co-workers @HarvardCCB Profiling the landscape of drug resistance mutations in neosubstrates to molecular glue degraders: https://t.co/Soxb0jYpO5
0
13
63
Our paper employing CRISPR scanning to profile landscape of resistance mutations in neosubstrates to glue degraders is now online in @ACSCentSci Congrats! @brian_b_liau, @lilchromatinboy, @applecannon23, Cindy, @nicklue8, Jiaming, @sam_hoenig & @liaulab
pubs.acs.org
Targeted protein degradation (TPD) holds immense promise for drug discovery, but mechanisms of acquired resistance to degraders remain to be fully identified. Here, we used clustered regularly...
0
14
91
aaaaaaand we're officially out in ACS central science https://t.co/5QvtPF1X0v congrats again to @Palgosavi, @brian_b_liau, and the rest of the @liaulab team (@applecannon23, Cindy, Jiaming, @nicklue8, @sam_hoenig) now please don't ask me to look for missing commas for a week
pubs.acs.org
Targeted protein degradation (TPD) holds immense promise for drug discovery, but mechanisms of acquired resistance to degraders remain to be fully identified. Here, we used clustered regularly...
i never thought i would be in the TPD circles, but here i am, with a new preprint and two cats named pomalidomide and lenalidomide big shoutout to @Palgosavi and @brian_b_liau for bringing this to life
0
2
18
Honored and very grateful for the invaluable funding support from @DamonRunyon.
Congrats to the 2021 Damon Runyon-Rachleff Innovators! Six extraordinary researchers have received five initial grants, and four previous awardees were given “Stage 2” support. Read more about our new #braveandbold Innovators here: https://t.co/sYvH1NAUIt and below 🔽
6
2
53
(1/4) Molecular glue degraders are captivating drug discovery, but how might drug resistance emerge? Read our new preprint! We identify themes of degrader resistance involving neosubstrates. #TPD
https://t.co/5CQIKZZ0iA
4
71
376
Honored to have hosted @EsbenSv over the last few months. Hope you had a good time and see you again soon!
Many thanks to @brian_b_liau and all members of @liaulab at @HarvardCCB for hosting my exchange stay over the past ~3.5 months. So much awesome science! It has been a wonderful time in Cambridge. Now looking forward to seeing as much family back in DK🇩🇰 as covid will allow🎄
0
0
7