OMA Database
@OMABrowser
Followers
229
Following
21
Media
25
Statuses
172
Orthologs among 2000+ complete genomes across all of life. Follow us to stay informed of new releases and features. We welcome feedback and suggestions!
Zurich/Lausanne/London
Joined November 2013
FastOMA is out now in Nature Methods 🎉: https://t.co/6nNVfbCVqr A new orthology inference algorithm that scales linearly and is highly accurate. FastOMA can process all >2000 eukaryotic UniProt ref proteomes <24 hours 🚀. Try it out at https://t.co/QUPiRx33J9
1
44
100
Session 24: OMA/OMArk is starting now! To join in the session, jump over to our discord at: https://t.co/FtxlJMZQ5Y Or you can join in on the YouTube stream at: https://t.co/Y9BRNe01pE
@SinaMajidian @OMABrowser
discord.com
Discord is great for playing games and chilling with friends, or even building a worldwide community. Customize your own space to talk, play, and hang out.
0
2
3
Our OMArk paper is finally out at @NatureBiotech! I am excited to share the final results of our work with @Gloveface @cdessimoz @VictorRossier @DessimozLab. OMArk is a one-stop shop for eukaryotic genome annotation assessment. Link : https://t.co/wu11J7sOpN (🧵)
nature.com
Nature Biotechnology - A new tool checks the quality of gene annotations in genome sequences.
1
30
64
FastOMA algorithm in a nutshell! More details in the preprint
biorxiv.org
The surge in genome data, with ongoing efforts aiming to sequence 1.5M eukaryotes in a decade, could revolutionise genomics, revealing the origins, evolution, and genetic innovations of biological...
After trying many different ideas over years, we're happy to have found a scalable solution without sacrificing accuracy. It works by clustering homologs using k-mer based placement, followed by a highly parallelised taxonomy-guided processing of families using subsampling.(3/4)
0
3
16
Excited to release FastOMA, a rewrite of OMA orthology algorithm that scales linearly, while maintaining OMA’s high accuracy. FastOMA can process all >2000 eukaryotic UniProt ref proteomes within 24 hours. Try it out https://t.co/helQL3wo4c Preprint: https://t.co/vs21iN8P7o (1/4)
1
36
87
Want to use the OMA browser to get orthologs, but your species of interest isn’t there? Request to add it for the next release! (Deadline: 7 Feb 2024) Search available genomes in OMA: https://t.co/ZPbg2l1B0f Suggest a genome:
0
6
11
💡 That's a wrap on our #OMAUpdate2024 highlights! Dive into the new OMA browser to explore these features and more. Your feedback is always welcome! #Genomics #Bioinformatics
0
0
0
📈 Assess the quality of your proteomes with OMArk, a new tool that estimates completeness, detects fragments, and identifies contamination. https://t.co/ypUnqZftcI
#Proteomics #QualityAssessment #OMAUpdate2024
1
0
1
🔎 Find what you need faster with our improved search engine. Contextual understanding for more accurate results in your genomics research. #SearchOptimization #OMAUpdate2024
0
0
0
🛠️To go further, combine the OMA Browser with other tools in the OMA ecosystem: OMAmer, OMArk, OMAMO, and Read2Tree. From proteome mapping to phylogenetic tree construction, we've got your research needs covered! #BioinformaticsTools #OMAUpdate2024
2
0
0
📊 We now offer Gene Ontology annotations for Hierarchical Orthologous Groups, enabling both extant and ancestral GO enrichments. Dive into functional genomics with ease! #GeneOntology #OMAUpdate2024
1
0
0
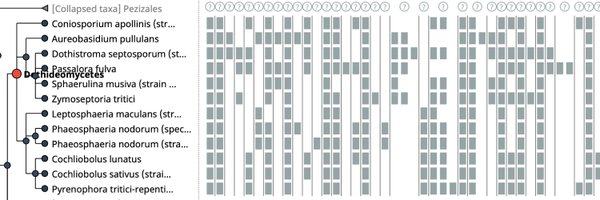
🔍 Check out our revamped Local Synteny Viewer for comparing genomic neighborhoods across extant and ancestral genomes. A powerful tool for genomic research. #Synteny #Genomics #OMAUpdate2024
1
0
0
🌳 Explore ancestral genomes like never before! Our update includes 1,133 Ancestral Genome pages, offering insights into gene content and order of ancient genomes. #AncestralGenomics #OMAUpdate2024
1
0
0
🧬 New in #OMAUpdate2024: We've significantly expanded our coverage of bacteria and archaea, including Asgardarchaeota, for a deeper understanding of microbial genome evolution. #Genomics #Microbiology
1
0
1
💡 Why choose OMA? Unique in its comprehensive orthology analysis, OMA offers unparalleled depth in prokaryotic & eukaryotic coverage, ancestral genome insights, and a suite of specialized tools. A one-stop solution for comparative genomics! #Orthology #OMAdvantage #OMAUpdate2024
1
2
4
🚀 Exciting news from the OMA team! Our 2024 update brings a host of improvements to our orthology knowledgebase. Stay tuned for a series of highlights! #OMAUpdate2024 #Genomics #Bioinformatics
https://t.co/hhTpwoj3xf
1
13
30
@SinaMajidian @OMABrowser @ISBSIB @cdessimoz @DessimozLab @Why_NeverS For those who missed it... https://t.co/n6UcQZlLec
0
3
4
A 2-hour hands-on workshop to learn about @OMABrowser and OMArk for proteome quality assessment! registration link: https://t.co/FIZ4zD9GY0 preprint: https://t.co/8a8bSWkCKT tool: https://t.co/WsKW7mOQtL
@KamilSJaron @rmwaterhouse @nuala_oleary @giulio_formenti @CibeleCaio @MahUliano @Why_NeverS @SinaMajidian @CorreardSolenne @Toby_Baril_Bio @EBPgenome @SangerVGP @SangerToL @genomeark @erga_biodiv @BioGenEurope @CatBiogenoma @DAISEA_AfricaBP New topics include: ✅ Starting a comparative genome study from CNGBdb: datasets, analysis platform, and spatial transcriptomics database ✅ Genome profiling using Genomescope ✅ OMA and OMArk for homology exploration and gene annotation quality control
0
8
12
Bringing meaning to biological #data: #knowledgegraphs meet #ChatGPT → our new Knowledge Representation team investigate how #NLP supports #lifescience researchers in exploring datasets they are not familiar with. Read more: https://t.co/LucrFfuC1J
0
12
20