Ying Ge Research Lab
@Ge_Lab_UW
Followers
2K
Following
1K
Media
144
Statuses
682
Mass spectrometry-based proteomics group studying cardiovascular disease @ UW-Madison. Account handled by the students🫀🧀 https://t.co/ZCH80vOKe7
Madison, WI
Joined March 2019
If you are interested in utilizing our photocleavable surfactant Azo for your protein applications (bottom-up MS, top-down MS, protein extraction), it’s available for purchase below from Sigma!⚡️⬇️.
1
3
17
In our newest publication, we reveal the structural heterogeneity of AMPK -a heterotrimeric complex that acts as a master regulator of cellular metabolism- using integrated native and denatured top-down mass spectrometry (TDMS)! Check it out below 👇.
pubs.acs.org
Adenosine monophosphate-activated protein kinase (AMPK) is a heterotrimeric complex (αβγ) that serves as a master regulator of cellular metabolism, making it a prominent drug target for various...
0
0
6
Congratulations to Dr. Kalina Rossler on a successful thesis defense! Your years of service & friendship to the lab will forever be cherished. You did it, Doctor! 🫀🎓🎉 #ProudMoment
0
0
14
Huge congratulations to Emily Reasoner on a successful PhD defense! All those years of hard work, research, and perseverance have paid off — Dr. Reasoner is official! 🧠🎓👏 #Lucky18
1
0
19
Check our our recent collab in @NatureCVR ! Congratulations & many thanks to Ieng Lam Lei & @PaulTangMD for the opportunity to be part of such a great project! 🫀💪.
Research | Lei et al. show that cold preservation of heart transplants triggers mineralocorticoid receptor clustering in cardiomyocyte nuclei, and blocking this improves heart transplant function.
0
1
2
RT @PaulTangMD: Heartfelt thanks to Ienglam Lei & all collaborators on this very exciting trip down the rabbit hole of organ preservation b….
0
2
0
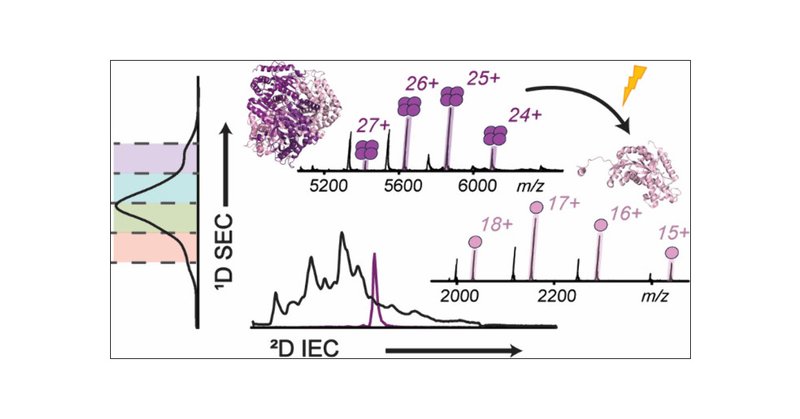
🚨Paper Alert🚨.We've developed the first nondenaturing online 2D-LC method for native top-down MS, enabling rapid structural analysis of endogenous protein complexes. 🧬⚡ #MassSpec #Proteomics #nTDP .
pubs.acs.org
Protein complexes are essential for virtually all biological processes, yet their structural characterization remains a major challenge due to their heterogeneous, dynamic nature and the complexity...
0
0
6
#ASMS2025 The Ge Lab wrapped up Wednesday & Thursday with some *popular posters* from Dr. Gao, Emily, Liam, and Anna Grace!
0
0
11
RT @FemalesInMS: Thank you all for attending the #FeMS+ networking session at #ASMS2025! Special thank you to @livia__se for delivering an….
0
4
0
#ASMS2025 We had some tasty seafood at our Ge Lab reunion dinner last night! So happy to see familiar faces at our favorite conference 🦀🦪🍰
0
0
9
#ASMS2025 The Monday poster session was a hit for the Ge Group! Thanks to everyone who stopped by to see Rob, Mallory, & our alumnus Jake Melby (now at Astrazeneca)! 🦀
0
1
13
RT @TopDownProteome: Join us for #Proteoform Thursday June 19 as Juan Vizcaíno presents "The Implementation of Open Science Practices Can E….
0
3
0
#ASMS2025 Congrats to our fearless leader @yingge2121 for a great presentation during Agilent's HRAM User Group Meeting yesterday!
0
0
11
The Ge Group is enjoying some weekend fun before #ASMS2025 kicks it into high gear tomorrow! #CrabbyForMassSpec
0
0
8
The Ge Research Group is gearing up for #ASMS2025 in Baltimore, MD! ✨. You can check out our presentation and poster schedule below. Come find us and chat science (and maybe have a crab cake or two 😉🦀)!. #CrabbyForMassSpec #ScienceOnTheBay
0
0
2
RT @WeiWei_Stanford: My lab is recruiting a postdoc fellow to characterize new biochemical pathways that control taurine and glucose metabo….
0
125
0
RT @TopDownProteome: Please join us May 15 for #Proteoform Thursday - David Roberts presents "Guiding the Rational Design of Protein Assemb….
0
4
0
Big news - our group now has a LinkedIn page! Please give us a follow to get even more updates on the Ge lab's shenanigans! 😉
linkedin.com
Hello, LinkedIn! We are a mass spectrometry-based proteomics group studying cardiovascular disease at the University of Wisconsin-Madison under the direction of Dr. Ying Ge. Here's a little more...
0
1
11