Zijie Zhao
@ZijieZhao1996
Followers
146
Following
100
Media
0
Statuses
12
PhD student in biomedical data science @UWMadison. Interested in statistical genetics, genetic risk prediction, polygenic risk score.
madison, WI
Joined January 2017
@Q_StatGen, @yy_stat @ZijieZhao1996 and I show that sibling relative rank shapes associations between PGS and educational attainment in @PNASNexus Implications for within-family inequalities and interpreting PGS Paper: https://t.co/sKKDNMZQ7X
@uwcdha @UWLaFollette @backwugs
2
6
17
You'll want to check out @Jiacheng_Miao's new paper if your work involves gene-environment interaction. This is our best GxE work so far. We introduce a statistical framework named PIGEON to reimagine how GxE studies could and should unfold.
biorxiv.org
In this study, we introduce PIGEON—a novel statistical framework for quantifying and estimating polygenic gene-environment interaction (GxE) using a variance component analytical approach. Based on...
2
28
78
Optimizing and benchmarking polygenic risk scores with GWAS summary statistics https://t.co/hSom844HO9
0
8
43
Very excited to share our recent work! PUMA-CUBS can fine-tune, benchmark, and do ensemble learning for various PRS methods using just one GWAS sumstats. Come by poster 3346 today #ASHG22 to see more details about our method!
Preprinted in time for #ASHG2022! Happy to share @ZijieZhao1996's new work. We present PUMA-CUBS which advances PRS benchmarking & optimization. We can now fine-tune any PRS models and even do ensemble learning to combine models using GWAS sumstats alone!
0
2
11
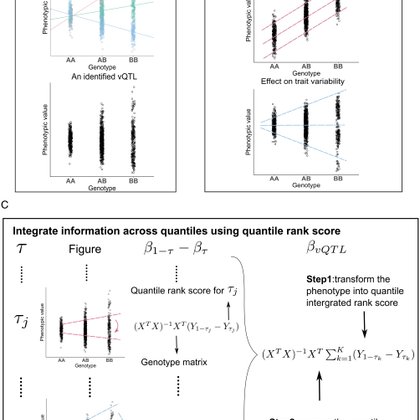
Extremely excited to see my (first first-author) paper now out @PNASNews! We develop a novel framework named QUAIL that uses quantile regression to identify SNPs associated with phenotypic variability (vQTL). Work is supervised by @Q_StatGen. https://t.co/y4vkTerFq5. 1/n
pnas.org
Detecting genetic variants associated with the variance of complex traits, that is, variance quantitative trait loci (vQTLs), can provide crucial i...
2
8
36
Excited to finally throw our hat in the ring. Here is our new method (named X-Wing) for cross-ancestry PRS. A key takeaway is that we now do EVERYTHING with sumstats alone which is a major methodological advance. Work is led by @Jiacheng_Miao & Hanmin.
biorxiv.org
Polygenic risk scores (PRS) calculated from genome-wide association studies (GWAS) of Europeans are known to have substantially reduced predictive accuracy in non-European populations, limiting its...
3
29
80
New preprint by MS student @JiawenChennn (now doing her PhD at UNC) 🍾 We built a tool (named ORIGAMI) to simulate offspring genome-wide data from parental genomes. It accounts for technical issues such as genetic distance, sex difference, etc. #ASHG2020
biorxiv.org
Polygenic risk scores (PRS) derived from summary statistics of genome-wide association studies (GWAS) have enjoyed great popularity in human genetics research. Applied to population cohorts, PRS can...
2
12
41
Super excited to share our preprint on decomposing the direct and indirect genetic effects using only GWAS sumstats. A very fun project! Started < 1 year ago; really thankful to get it out during a pandemic! Miss the lunch over which Q @Q_StatGen peached the idea to me.
New work preprinted! This one is about estimating genetic nurture with GWAS sumstats. Most approaches for estimating genetic nurture requires family-based, individual-level data. We show that all you need is offspring phenotypes on existing GWAS cohorts
1
1
12
🍾New preprint from us! We found it puzzling that rare and common variants do not affect the same genes in autism. In this project, we generalize TWAS to trio data and highlight a key TF POU3F2 that regulates many ASD genes affected by rare mutations
biorxiv.org
Recent advances in consortium-scale genome-wide association studies (GWAS) have highlighted the involvement of common genetic variants in autism spectrum disorder (ASD), but our understanding of...
1
19
49
Celebrate Halloween with new preprint!🎃We introduce a new method to identify precise genome regions with local genetic correlations between traits. This was a fun collaboration with Hanmin Guo and Lin Hou from @Tsinghua_Uni (all those late-night calls🤣)
biorxiv.org
Genetic correlation analysis has quickly gained popularity in the past few years and provided insights into the genetic etiology of numerous complex diseases. However, existing approaches oversimpl...
2
10
31
Very excited to share our recent work in my VERY FIRST tweet! Lots of thanks to @Q_StatGen, there were tons of advising behind all the work. Try the package on your GWAS! Any feedback is welcome and we'll add more features soon.
Very, very pleased to share this new work from my lab with all of you. Parameter-tuning in polygenic scores has always been challenging in our field. We had this crazy idea about a year ago that maybe we can perform cross-validation on GWAS sumstats. Indeed we can! #ASHG19
0
5
11