Stephane Peyregne
@StephPeyregne
Followers
322
Following
180
Media
0
Statuses
123
Evolutionary geneticist at the Max Planck Institute for Evolutionary Anthropology
Allemagne
Joined August 2012
RT @benmpeter: Today is a very big day for our research group, with two of my students, @arevsumer and @IasiLeonardo publishing papers on….
science.org
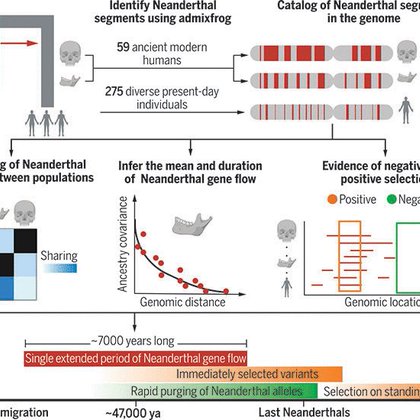
Gene flow from Neanderthals has shaped genetic and phenotypic variation in modern humans. We generated a catalog of Neanderthal ancestry segments in more than 300 genomes spanning the past 50,000...
0
40
0
RT @IasiLeonardo: Out today in @sciencemagazine, we've journeyed into our shared history with Neandertals by analyzing over 300 present-day….
science.org
Gene flow from Neanderthals has shaped genetic and phenotypic variation in modern humans. We generated a catalog of Neanderthal ancestry segments in more than 300 genomes spanning the past 50,000...
0
41
0
RT @ElenaIreneZ: Interested in sedaDNA, aDNA, or forensic genetics? . Join me in Copenhagen! . I'm hiring a PhD and a Postdoc to start in 2….
0
49
0
RT @PopliRatan: PCA and F-statistics are routinely used in population genetic studies. We provide a statistical framework to combine them i….
0
11
0
RT @iosif_lazaridis: Congratulations to Ali Akbari.@aliakbari23.on his amazing new work on selection in Western Eurasia that is finally rel….
biorxiv.org
We present a method for detecting evidence of natural selection in ancient DNA time-series data that leverages an opportunity not utilized in previous scans: testing for a consistent trend in allele...
0
52
0
RT @FossilHistory: The first Neanderthal DNA sequence was published #OnThisDay in 1997. It suggested Neanderthals did not interbreed with m….
0
51
0
RT @evolutionscribe: This is a big deal--a second high quality Denisovan genome has been sequenced, and it's old!.The most ancient human ge….
science.org
200,000-year-old DNA from Siberian cave shows our elusive, extinct cousins mated repeatedly with Neanderthals
0
301
0
RT @FSanchezQuinto: Casi 20 generaciones de la @lcgunam presentes en #SMBE2024! Un honor ser parte de esta gran iniciativa que transformado….
0
22
0
Many thanks to @jfkelso for giving me the opportunity to co-advise Yaniv on this work, and congratulations to Yaniv for this very thorough analysis for the first chapter of his PhD thesis! 7/7.
0
0
7
Yaniv has made all his code for the analyses available (, which we hope will be useful. 6/7.
github.com
Contribute to yanivsw/y_chr_reference_bias development by creating an account on GitHub.
1
1
3
RT @_JBallesterosV: Come to my poster on exploring genetic diversity in the Americas through ancient whole genomes. I’m eager to discuss an….
0
1
0
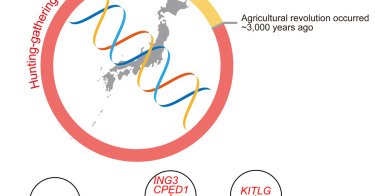
RT @NiallPC: Very happy that the new paper on imputation and selection in ancient Japan, from myself, @shige_nakagome and our collaborator….
cell.com
Genomics; Anthropology
0
16
0
RT @oliveira_srs: Do you use Local Ancestry Inference (LAI)? Wanna know how much error to expect given source divergence or time since admi….
0
13
0