Josh Welch
@LabWelch
Followers
1K
Following
2K
Media
65
Statuses
554
Computational Biologist, Associate Professor at U. Michigan
Ann Arbor, MI
Joined July 2019
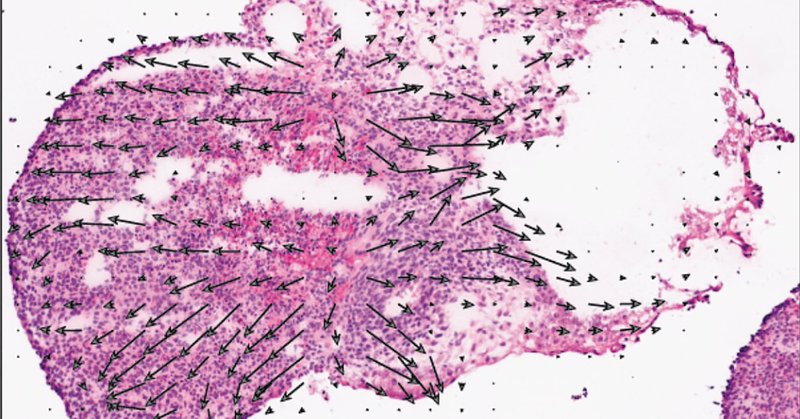
Interested in studying cell differentiation at the cellular level but don't trust your UMAP plots? Try visualizing your cell differentiation in space with our TopoVelo tool!
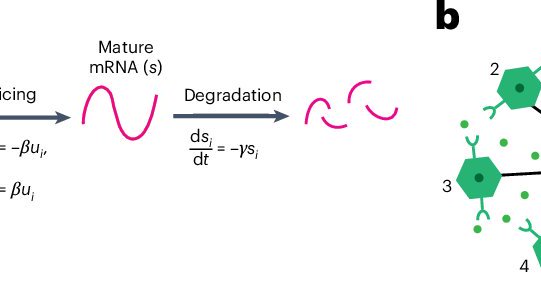
Our paper is now in @NatureBiotech! Topological velocity inference from spatial transcriptomic data. TopoVelo infers the direction of differentiation/migration; quantifies spatial cell influence; and identifies morphology changes during differentiation. 🧵
4
30
157
Our code, models, data, and tutorials are available on GitHub and HuggingFace:.
huggingface.co
2
0
14
My department issued a press release about our paper:
medschool.umich.edu
How do cells come together to make tissues? To answer this question, using AI, the Welch laboratory has developed a new computational tool that incorporates space and time into models of cell fate...
Our paper is now in @NatureBiotech! Topological velocity inference from spatial transcriptomic data. TopoVelo infers the direction of differentiation/migration; quantifies spatial cell influence; and identifies morphology changes during differentiation. 🧵
0
0
8
@JialinLiuSKP @therealjklee Also check out the research briefing for our article here:
nature.com
Nature Biotechnology - Cells in a tissue influence each other through physical and chemical cues as they differentiate to their final fates and migrate through space. A new technique integrates...
0
0
5
I'm really proud of my trainees Yichen Gu, Jialin Liu (@JialinLiuSKP) and Justin Lee (@therealjklee) who led the project. This paper represents an important milestone for our group: it's the first in-house data we have published since opening our own wet lab!.
1
0
5
Check out our Python package, with tutorials and notebooks to reproduce the paper results!.
github.com
Contribute to welch-lab/TopoVelo development by creating an account on GitHub.
2
0
5