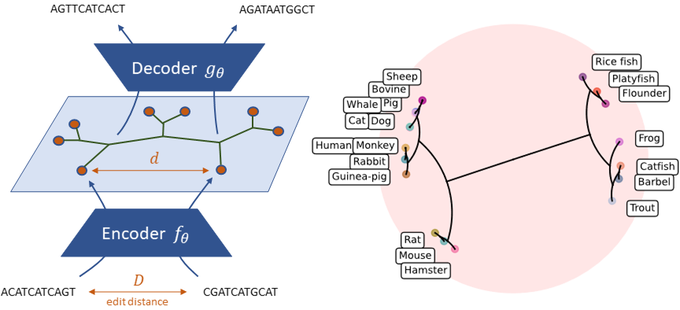
Can ML help us obtain precise approximations of fundamental bioinformatics problems? We present NeuroSEED a framework to embed biological sequences, its effectiveness in the hyperbolic space and how it can be used for hierarchical clustering and MSA
5
26
133
Replies
We show the improvement provided by data-dependent embedding methods in preserving the evolutionary distance between sequences. In particular, the hyperbolic space is able to capture the hierarchical relationship between the sequences, significantly reducing the distortion.
1
2
7
Finally, we propose a series of ways of adapting the framework to perform the combinatorial intractable tasks of hierarchical clustering and multiple sequence alignment, all of which show significant runtime improvements over baseline methods.
1
1
7
You can find the paper at and the code at . Work done under the amazing supervision of
@RexYing0923
@Mpmisko
@PetarV_93
@jure
and
@pl219_Cambridge
0
3
13
@GabriCorso
@PetarV_93
@jure
@Mpmisko
@RexYing0923
@pl219_Cambridge
Indeed great work. I enjoy to read it a lot.
Just my concern is the claim that edit distance is the best way to measure evolutionary distance between biological sequence. I guess even small edit distance might change function a lot, sometimes vice versa.
1
0
1
@GabriCorso
@PetarV_93
@jure
@Mpmisko
@RexYing0923
@pl219_Cambridge
@ixxmael_freitas
mais uma aplicação em ciências biológicas do meu tema de estudo ó que legal
0
0
1
@GabriCorso
@PetarV_93
@jure
@Mpmisko
@RexYing0923
@pl219_Cambridge
Really enjoyed this paper - great observations regarding performance of hyperbolic vs. standard distances. However, I gotta say that labeling these performance tables as "Figures" is rather... generous :)
0
0
1