Xuehua Zhong
@zhonglab
Followers
697
Following
2K
Media
11
Statuses
388
Biology Professor @WUSTL studying Epigenetic regulation of plant traits and environmental adaptation
Washington University St Louis
Joined June 2019
🌱 Wrapping up in Fort Collins, CO at the Plant Epigenetics & Epigenome Engineering conference! 👏 Congrats to our Future of Science Fund Scholars - Vanda Adamkova, Carl Simmons & Kehui Zhu! 🔗 https://t.co/NnCFFG1D8X
#KeystoneSymposia #KSPlantEpi26 @Doris_Wagner_ @zhonglab
0
2
4
Check out our latest review article on DNA methylation dynamics: patterns, regulation, and function
0
6
11
Please join us!
🌱 Poster abstracts due Sept 18! Early-career scientists, submit your work for Keystone Symposia: Plant Epigenetics & Epigenome Engineering. Showcase your research, get feedback & connect with leaders. 🔗 https://t.co/LvdOFyXN3G
#KSPlantEpi26 @Doris_Wagner_ @zhonglab
0
2
14
🌱 Poster abstracts due Sept 18! Early-career scientists, submit your work for Keystone Symposia: Plant Epigenetics & Epigenome Engineering. Showcase your research, get feedback & connect with leaders. 🔗 https://t.co/LvdOFyXN3G
#KSPlantEpi26 @Doris_Wagner_ @zhonglab
0
7
18
Save $200! Early reg for Plant Epigenetics & Epigenome Engineering ends Aug 14, 11:59 PM MDT. Join top researchers incl. @Doris_Wagner_ & @zhonglab exploring plant epigenome regulation & engineering. 🎥 https://t.co/Uh0clJslYx
https://t.co/rdYedhTF8u
#KSPlantEpi26
keystonesymposia.org
Join us at the Keystone Symposia on Plant Epigenetics and Epigenome Engineering, October 2025, in Fort Collins, with field leaders!
1
6
10
Please join us at this Keystone Symposium on Plant Epigenetics and Epigenome Engineering.
keystonesymposia.org
Join us at the Keystone Symposia on Plant Epigenetics and Epigenome Engineering, October 2025, in Fort Collins, with field leaders!
0
12
23
Why is it that gene promoters are so good at avoiding DNA methylation? Work from Ming Wang, Yan He et al., shows that H3K4me3, enriched at promoters, acts as an anti-DNA methylation mark by recruiting DNA demethylase enzymes! Nice work guys!
2
36
92
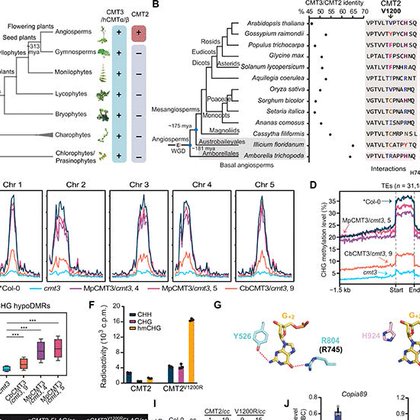
Key findings include: 1. CMT2 is originated from a more ancient CMT3 during plant’s whole genome duplication. 2. CMT2 lost its CHG activity due to the lacking of a key arginine residue.
0
0
3
5. CMT2 N-terminus is highly plastic and can be tolerant to various mutations observed from natural accessions in a wide variety of environment conditions worldwide.
0
0
0
3. An engineered mutation empowers CMT2 a “one stone two birds” like activity in restoring both CHG and CHH methylation. 4. Unlike CMT3, CMT2 has a long, flexible N-terminal that controls its own protein stability.
1
0
0
Our work on evolution and functional divergence of DNA methyltransferases is out @ScienceAdvances. Kudos to @jiang_jianjun. We revealed how two closely related CMTs evolve and coordinately shape the non-CG methylation landscapes in plants.
science.org
Functional diversification of substrate-specific DNA methyltransferases accounts for plant genome complexity during evolution.
3
25
78
With much anticipation and excitement, @ThePlantCell announces that Pablo Manavella @manavellalab has been appointed its next Editor-in-Chief! 🙌 Read the full announcement: https://t.co/ihMwRk4XwK
@CSIC @IHSM_CSIC_UMA @ASPB #plantscience
39
56
255
Totally agree with Keiko. Great experience @AnnualReviews editorial board meting with excellent discussions and tons of fun!!!
Excellent plant scientists, colleagues and friends; stimulating & insightful scientific discussions; fun & exciting conversations; and fantastic cuisine at the restaurant called “Botanist”. What would you need more? Thanks @AnnualReviews 🌹🌻🌱🌲🌾🍋🍅🌽🥰
0
0
9
Wash U Biology is hiring. Please repost and encourage your trainees to apply. https://t.co/N3kICLahkv
jobs.sciencecareers.org
Faculty positions at the rank of assistant or associate professor for a cluster hire in the departments of Biology, Chemistry, & Physics at WashU.
0
7
10
Our deep dive into cis-regulatory diversity using single-cell genomics across maize germplasm reveals how genetic variants reshape chromatin accessibility, transcription factor binding, chromatin interactions and much more. Led by @Marand_Lab! https://t.co/keTimFGCJt
biorxiv.org
Gene expression and complex phenotypes are determined by the activity of cis -regulatory elements. However, an understanding of how extant genetic variants affect cis -regulatory activity remains...
0
60
136
Our latest preprint on the mechanistic origin and evolution of CHH DNA methyltransferase CMT2 and how it diverged from CMT3 to achieve substrate specificity.
Substrate specificity and protein stability drive the divergence of plant-specific DNA methyltransferases https://t.co/xoQcpmOU2S
#biorxiv_plants
0
10
31
Substrate specificity and protein stability drive the divergence of plant-specific DNA methyltransferases https://t.co/xoQcpmOU2S
#biorxiv_plants
0
4
10
Abstract deadline update is July 18th for MIDWEST SDB 2024 Registration deadline is August 6th @SridharanLab @midwestSDB
0
7
6
Registration is now open!
Registration is open for the Midwest SDB @SocDevBio meeting August 11th-13th in beautiful Madison, Wisconsin. Highlights include @ItaiYanai @zhonglab @PetersenNeuro Follow us @midwestSDB Thanks to @SridharanLab & Jaime Gingerich @UWEauClaire for organizing
0
6
7
I'm thrilled to announce the complete program for #ICAR2024SanDiego! 👉🏽Please check out the amazing line of up over 300 speakers in nearly 50 sessions! I hope to see you soon in San Diego.. https://t.co/AFa9qcvHfo
0
23
44