Rafael Josip Penić
@RJPenic
Followers
48
Following
805
Media
0
Statuses
38
PhD student @ Faculty of Electrical Engineering and Computing, University of Zagreb | 🇭🇷 | Machine Learning | AI in Structural Biology
Joined January 2022
Thrilled to share that our paper introducing RiNALMo is now published! Check it out 👇
Our RiNALMo RNA language model has been published in Nat Comm https://t.co/2EXzExKdzK Great work by @RJPenic and @TinVlasic, with support and patience in teaching us RNA biology from Roland G Huber and @ywan_wan RiNALMo is already an SOTA as a benchmark for RNA LLMs.
1
0
6
🚀 Excited to share that our preprint is out! Huge thanks to my amazing collaborators @DobrikG, Marin Šilić, and @PetarV_93 ! KNARsack – a neural algorithmic reasoner for the Knapsack problem. 🎒🧠
2
2
12
I am happy to introduce Campolina, a deep neural framework that replaces traditional algorithmic approaches for nanopore signal segmentation and improves segmentation quality for real-time analysis. Preprint and details in the thread👇 K. Friganovic, @BryanHooi1, @msikic 1/7
2
8
15
I am happy to share our new preprint introducing MADRe - a pipeline for Metagenomic Assembly-Driven Database Reduction, enabling accurate and computationally efficient strain-level metagenomic classification. @msikic, @r_vicedomini, @KrizanovicK 🔗 https://t.co/YUq9Xbp2Lm 1/9
biorxiv.org
Strain-level metagenomic classification is essential for understanding microbial diversity and functional potential, but remains challenging, particularly in the absence of prior knowledge about the...
4
13
12
We are organising https://t.co/bmP3bmhJHq at#ICLR2025. Topics: RNA, DNA and cell LLMs, structure, modifications, correction, variant calling... Deadline: Feb 10th 2025 Time for revision: 1 month Accepted papers - the opportunity to be invited by Nature Methods for submission
ai4na-workshop.github.io
AI4NA Workshop @ ICLR'25
0
8
19
Submit your papers to our workshop AI for Nucleic Acids workshop at ICLR 2025!
📢 Submissions are open! Join us at #AI4NA 🇸🇬 @iclr_conf to present your latest work 🧬 🚨 Updates: We've revised paper length guidelines & extended the deadline to Feb 10, 2025, AoE. 🔗 Details on our webpage (link in comments). Don’t miss out! 🌟
0
2
6
🚨 1 month to go! 🚨 The submission deadline for the AI4NA workshop at @iclr_conf is fast approaching! 🧬 ✨ Submissions on OpenReview will open soon—stay tuned! ✨ 🔗 Learn more on our web page (link below 👇) #AI4NA #ICLR2025
1
6
9
🚀Thrilled to be part of ICLR 2025! Join our workshop AI for Nucleic Acids (AI4NA) to explore cutting-edge research and connect with field leaders. Thanks to our organizers and @iclr_conf for making this possible. Find the link to the website in our bio! More info below👇
1
6
10
RNA structure prediction is still an open problem. Look at our benchmark results (including Alphafold 3!) https://t.co/i73a4AUekB w/ @im50603 Tin Vlasic Yang Li @BryanHooi1 and Zhang Yang
biorxiv.org
Several deep learning-based tools for RNA 3D structure prediction have recently emerged, including DRfold, DeepFoldRNA, RhoFold, RoseTTAFoldNA, trRosettaRNA, and AlphaFold3. In this study, we...
1
7
19
A Comparative Review of Deep Learning Methods for RNA Tertiary Structure Prediction https://t.co/vF1bm87pSh
#biorxiv_bioinfo
0
3
5
We are seeking talented PhD candidates in CS, physics or math to join us. The topics include the development of AI for cancer and ageing research. 📅 Application Deadline: 1st Dec 2024 Lab: https://t.co/8KCCBRz4w6 Scholarship: https://t.co/umyR8aeTwR Please share!
a-star.edu.sg
1
24
39
CoPRA: Bridging Cross-domain Pretrained Sequence Models with Complex Structures for Protein-RNA Binding Affinity Prediction - Curate large datasets of RNA-Protein pairs (PRI30K, 150K pairs) from BioLIP2, and an affinity dataset (PRA310, 435 complexes) from PDBBind, ProNAB, and
0
15
45
Our RNA LLM RiNALMo has received the prize for the most ambitious submission at the Machine Learning for Life and Material Science workshop at @icmlconf 2024. w/ @RJPenic Tin Vlasic @ywan_wan and Roland Huber. Workshop: https://t.co/9gg8zYrRPy Preprint: https://t.co/V1N8D1QMyP
2
7
29
Rockfish: A transformer-based model for accurate 5-methylcytosine prediction from nanopore sequencing has been published in Nat. Comm!! Great work by @domstanojevic w/ Zhe Li, @sarrabakic and @rsyf2 Paper: https://t.co/diFCOVpBwn Code: https://t.co/mEDUI4agnZ
github.com
Contribute to lbcb-sci/rockfish development by creating an account on GitHub.
0
11
61
Considering building a human pangenome? Dive into our latest preprint for insights on the sequencing technologies and the minimal coverages needed for accurate assemblies https://t.co/RNjKreOeov Joint work with Prof JJ Liu lab w/@msprasad693, @JosipaLipovac and Filip Tomas!!
biorxiv.org
Long-read (LR) technologies from Pacific Biosciences (PacBio) and Oxford Nanopore Technologies (ONT) have transformed genomics research by providing diverse data types like HiFi, Duplex, and ultra-...
1
16
45
Our new paper is out! We made a lot of progress on our GNN-based de novo genome assembly paradigm and here we present all our findings and progress 🧬🥳 Code: https://t.co/L4vDO0SSZn Paper: https://t.co/90jHKL0Jjd See details of GNNome below 👇🧵
biorxiv.org
The critical stage of every de novo genome assembler is identifying paths in assembly graphs that correspond to the reconstructed genomic sequences. The existing algorithmic methods struggle with...
1
24
97
Happy to present RiNALMO - our RNA large language model https://t.co/V1N8D1QeJh w/ @RJPenic Tin Vlasic @ywan_wan and Roland Huber. RiNALMo is the largest RNA language model to date, with 650 million parameters pre-trained on 36 million non-coding RNA sequences. 1/2
2
42
96
Happy to present an initial draft of a telomere-to-telomere diploid Indian genome. A joint effort of Jianjun Liu's and my lab spearheaded by @msprasad693 and @JosipaLipovac
https://t.co/AES2pTsQkL or
github.com
Telomere-to-Telomere diploid Indian Genome . Contribute to LHG-GG/I002C development by creating an account on GitHub.
1
24
61
Are nanopore UL reads only long reads we need? We developed Herro https://t.co/KyR3LPx5rT AI error correction model that can correct reads to accuracy above Q30 while trying to keep informative positions intact. w/ @domstanojevic @DehuiLin @sergeynurk @PaolaFlorezdeS @nanopore
github.com
HERRO is a highly-accurate, haplotype-aware, deep-learning tool for error correction of Nanopore R10.4.1 or R9.4.1 reads (read length of >= 10 kbps is recommended). - lbcb-sci/herro
6
56
143
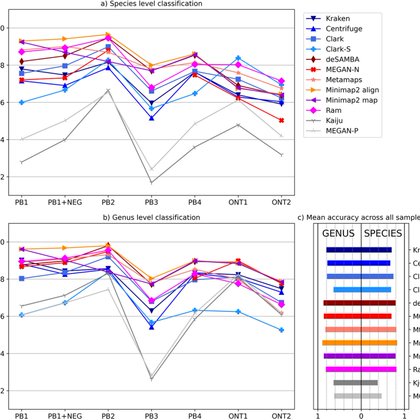
Long reads metagenome benchmark is out. Highlights. 1. In most cases Kraken, 2. minimap2/ram for slightly higher accuracy. 3. The right database is of huge importance 4. Check taxonomy files carefully. https://t.co/VqbanGKrIU w/@NiranjanTW @KrizanovicK @jmaricb @Sylvain14518009
bmcbioinformatics.biomedcentral.com
Background Long reads have gained popularity in the analysis of metagenomics data. Therefore, we comprehensively assessed metagenomics classification tools on the species taxonomic level. We analysed...
1
34
81